Overview
- Irreducible complexity, the claim that certain biological systems could not have evolved because removing any single part destroys function, rests on a logical error: it confuses the inability to remove a part from a finished system with the inability to build that system incrementally through exaptation, gene duplication, and scaffolding.
- Each of the three systems originally cited as irreducibly complex — the bacterial flagellum, the blood clotting cascade, and the adaptive immune system — has been explained through well-supported evolutionary mechanisms documented in dozens of peer-reviewed studies.
- The scientific rejection of irreducible complexity was formalized in the 2005 Kitzmiller v. Dover federal court ruling, in which the judge found that the concept had been refuted in peer-reviewed literature and that it does not qualify as science.
Irreducible complexity is a concept introduced by biochemist Michael Behe in his 1996 book Darwin's Black Box, defined as "a single system composed of several well-matched, interacting parts that contribute to the basic function, wherein the removal of any one of the parts causes the system to effectively cease functioning."1 Behe argued that such systems cannot be produced by natural selection acting on successive small modifications, because any precursor lacking even one component would be nonfunctional and therefore invisible to selection. He cited the bacterial flagellum, the blood clotting cascade, and the adaptive immune system as principal examples.1 The concept became a central argument of the intelligent design movement, which holds that certain biological features are best explained by an intelligent cause rather than by undirected evolutionary processes.
The scientific community has rejected irreducible complexity as both logically flawed and empirically refuted.17 The core error in the argument is the conflation of two distinct claims: that a system cannot function if a part is removed from its current configuration, and that the system could never have been assembled incrementally. Evolutionary biology provides well-documented mechanisms — including exaptation, gene duplication, and scaffolding — by which complex systems are built step by step from components that originally served different functions.2, 18 For each of Behe's three showcase systems, peer-reviewed research has identified plausible and well-supported evolutionary pathways, and the concept was explicitly rejected in the 2005 federal court decision Kitzmiller v. Dover Area School District.15
The logical flaw in the argument
The argument from irreducible complexity proceeds from a valid observation — that certain biological systems fail if a component is removed — to an invalid conclusion — that such systems therefore could not have evolved. The reasoning assumes that the only way to build a system is by adding parts one at a time to the current function, but evolutionary biology has long recognized at least three additional pathways that circumvent this assumption.2, 17
The first is exaptation, a term coined by Stephen Jay Gould and Elisabeth Vrba in 1982 to describe a trait that evolved for one function and was later co-opted for a different one.2 Feathers, for example, evolved originally for thermoregulation or display and were subsequently recruited for flight. A component that appears essential to a system's current function may originally have performed a different role entirely, meaning it was already present and available for incorporation into the evolving complex without having to arise de novo for its current purpose.2
The second mechanism is gene duplication. When a gene is duplicated, one copy continues to perform the original function while the other is freed from selective constraint and may acquire new capabilities through mutation. This process, first articulated comprehensively by Susumu Ohno in 1970, is the principal source of new genes and new biochemical functions throughout evolutionary history.18 Many of the components of supposedly irreducibly complex systems, including the serine proteases of the blood clotting cascade, arose through serial gene duplication and subsequent divergence.21
The third mechanism is scaffolding, sometimes called the scaffolding model or removal of scaffolding. In this scenario, a structure is first assembled with the help of additional components that later become dispensable. Once the core system is functional, natural selection can eliminate the scaffolding, leaving a structure that appears irreducibly complex because its evolutionary intermediates no longer exist. An archway made of stones is a useful analogy: the arch cannot stand without every stone in place, yet it was originally built atop a wooden scaffold that was removed after the keystone was set.4, 16
Darwin himself anticipated the basic objection in On the Origin of Species, writing that "if it could be demonstrated that any complex organ existed, which could not possibly have been formed by numerous, successive, slight modifications, my theory would absolutely break down. But I can find out no such case."19 More than a century and a half later, the cases proposed as counterexamples have been resolved through precisely the kinds of successive modifications Darwin envisioned.
The bacterial flagellum
The bacterial flagellum is a rotary molecular motor composed of approximately 30 to 40 different protein types, enabling bacteria such as Escherichia coli and Salmonella enterica to swim through liquid environments. Behe described it as a system in which the removal of any component — the basal body, the hook, the filament, or the associated regulatory proteins — would eliminate motility, and argued that no evolutionary precursor with fewer parts could have been functional.1 The flagellum became the signature example of the intelligent design movement.
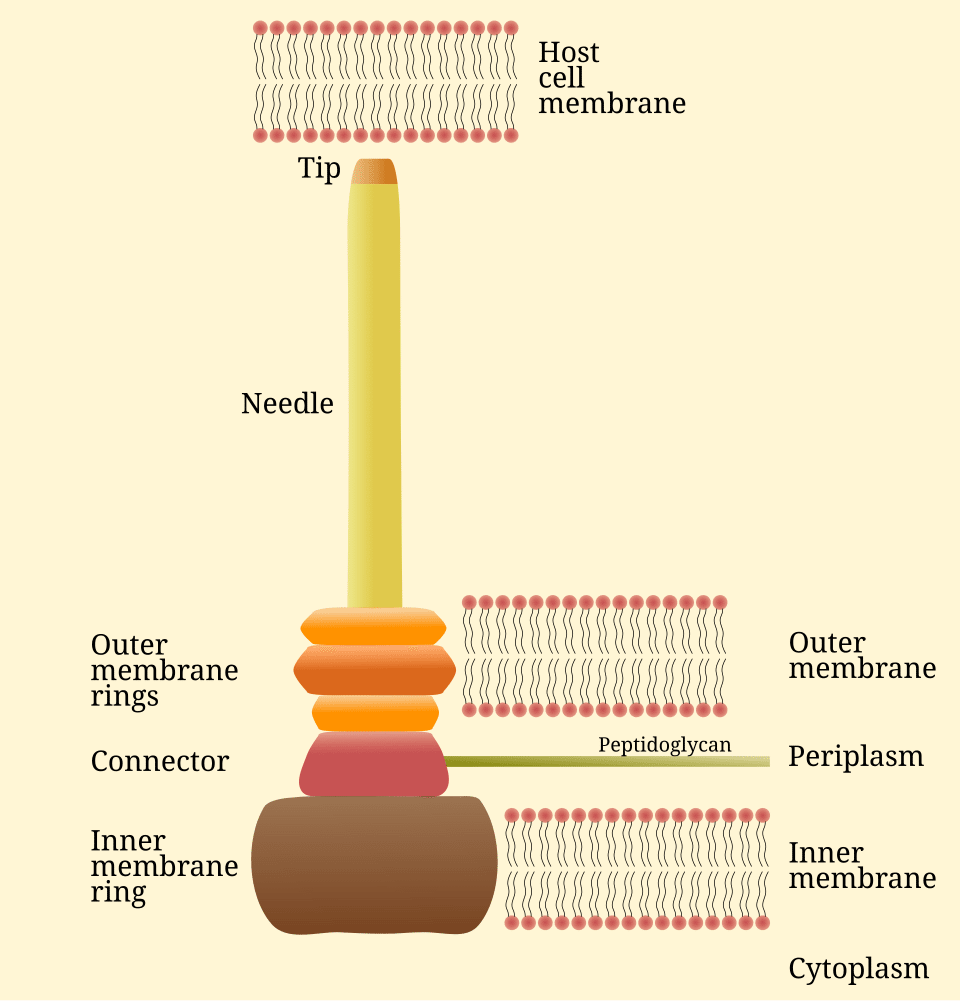
The evolutionary explanation centres on the type III secretion system (T3SS), a needle-like apparatus used by many pathogenic bacteria to inject proteins directly into host cells. The T3SS shares extensive structural and sequence homology with the basal body of the flagellum: approximately ten proteins are clearly homologous between the two systems, and multiple additional flagellar proteins have identifiable homologs in other non-flagellar systems.3, 24 This demonstrates that a functional subset of flagellar components already existed and served a non-motility purpose — precisely the kind of exaptation that undermines the irreducible complexity claim.
Mark Pallen and Nicholas Matzke, in a 2006 review in Nature Reviews Microbiology, demonstrated that nearly all of the core flagellar proteins have identifiable homologs with non-flagellar proteins, indicating that the flagellum was assembled through the successive recruitment and modification of pre-existing components rather than being constructed de novo.3 Pallen, Bailey, and Beatson further showed that the flagellar protein FliH shares homology with components of the F0F1 ATPase, a rotary enzyme found in all domains of life, suggesting that the ancestral rotary motor predates the flagellum itself.22
In 2007, Renyi Liu and Howard Ochman published a phylogenetic analysis in PNAS tracing the evolutionary history of flagellar proteins across 41 flagellated bacterial species from 11 phyla. They identified a core set of 24 structural genes present in the common ancestor of all bacteria and showed that many of these genes are related to one another by sequence similarity, indicating that they arose through serial gene duplication from a small number of ancestral genes — perhaps even a single precursor.5 Their results demonstrated a plausible order in which the structural components of the flagellum arose, from a simple secretion apparatus to the full rotary motor.
A comprehensive phylogenetic study by Sophie Abby and Eduardo Rocha in 2012, published in PLoS Genetics, further clarified the evolutionary relationship between the flagellum and the non-flagellar T3SS. Their analysis of 84 percent of phylogenetic trees supported a flagellum-first scenario, in which the non-flagellar T3SS evolved from the flagellum through a series of gene deletions, innovations, and recruitments, rather than the reverse.6 This finding reinforces the broader point: the flagellum and the T3SS are related systems that share a common ancestry, and neither is irreducibly complex in the sense that Behe intended.
Kenneth Miller summarized the state of the evidence in a 2004 chapter titled "The Flagellum Unspun," noting that the existence of the T3SS as a functional subset of flagellar components directly contradicts the claim that any subsystem of the flagellum would be nonfunctional. A system does not need to perform the same function at every stage of its evolution; it need only perform some selectively advantageous function.4
The blood clotting cascade
The mammalian blood clotting cascade is a multi-step enzymatic pathway in which the sequential activation of serine protease zymogens ultimately converts soluble fibrinogen into insoluble fibrin, forming a clot that stops bleeding. In mammals, the cascade involves more than two dozen genetically encoded factors arranged in a complex network with multiple feedback loops and regulatory checkpoints. Behe argued that the cascade is irreducibly complex because the removal of any single factor would prevent clotting, and that the entire pathway could not have arisen through gradual evolutionary modification.1
The evolutionary history of the clotting cascade is in fact well understood, primarily through the comparative genomics work of Russell Doolittle and colleagues spanning more than three decades. Doolittle and Feng demonstrated as early as 1987 that the serine proteases of the clotting cascade are related to one another by sequence similarity, indicating that they arose through a series of gene duplications from a common ancestral protease.21 The clotting factors, cofactors, and regulatory proteins were assembled stepwise through gene duplication, exon shuffling, and the accretion of new regulatory domains over hundreds of millions of years.9
Comparative genomics has provided direct evidence that simpler, functional clotting systems exist in organisms with fewer cascade components. In 2003, Jiang and Doolittle reconstructed the clotting scheme of the pufferfish (Fugu rubripes) from its genome sequence and found that 21 orthologs of mammalian clotting factor genes were present, but that genes for the contact pathway factors (XI, XII, and prekallikrein) were absent.7 Despite lacking these components, pufferfish clot their blood effectively. No authentic orthologs of any clotting factor were found in the sea squirt (Ciona intestinalis) genome, indicating that the cascade arose after the divergence of vertebrates from invertebrate chordates.7
The clotting system of the sea lamprey (Petromyzon marinus), a jawless vertebrate that diverged from the jawed vertebrate lineage approximately 500 million years ago, proved even more informative. Doolittle and colleagues showed in 2008 that lampreys possess a substantially simpler clotting scheme in which factors V and VIII appear to be represented by a single unduplicated gene, and factor IX is absent entirely.8 Lampreys nonetheless achieve effective hemostasis, demonstrating that a clotting system with fewer components is functional — precisely the kind of intermediate that Behe claimed could not exist.
Davidson and colleagues independently confirmed these findings using degenerate RT-PCR and the Fugu genome, concluding that the blood coagulation network is present in all jawed vertebrates and likely benefited from two rounds of whole-genome duplication early in vertebrate evolution.10 Doolittle's 2009 synthesis demonstrated that the full complement of clotting factors found in humans does not appear until pouched marsupials, with at least one key factor still absent in egg-laying mammals such as the platypus.9
Distribution of clotting factors across vertebrate lineages7, 8, 9
| Organism | Divergence time | Clotting factors present | Notable absences | Functional clotting? |
|---|---|---|---|---|
| Sea squirt (Ciona) | ~550 Ma | 0 orthologs | All factors | No cascade |
| Lamprey (Petromyzon) | ~500 Ma | ~15 | Factor IX; V/VIII unduplicated | Yes |
| Pufferfish (Fugu) | ~430 Ma | 21 | Factors XI, XII, prekallikrein | Yes |
| Chicken | ~310 Ma | ~23 | Contact pathway reduced | Yes |
| Platypus | ~170 Ma | ~25 | Factor XII | Yes |
| Opossum | ~160 Ma | ~26 | None significant | Yes |
| Human | — | ~27 | None | Yes |
This comparative evidence directly contradicts the irreducible complexity claim. The clotting cascade was not assembled all at once but accumulated new components progressively over vertebrate evolution, with each intermediate stage producing a functional hemostatic system. Species that lack components Behe deemed essential still clot their blood, demonstrating that the cascade is reducible and that simpler versions are fully operational.8, 9
The adaptive immune system
The adaptive immune system of jawed vertebrates generates an enormous diversity of antigen receptors — antibodies and T-cell receptors — through a process called V(D)J recombination, in which gene segments are cut and reassembled in different combinations by the RAG1 and RAG2 recombinase enzymes. Behe argued that this system, with its interdependent molecular components, could not have evolved gradually.1
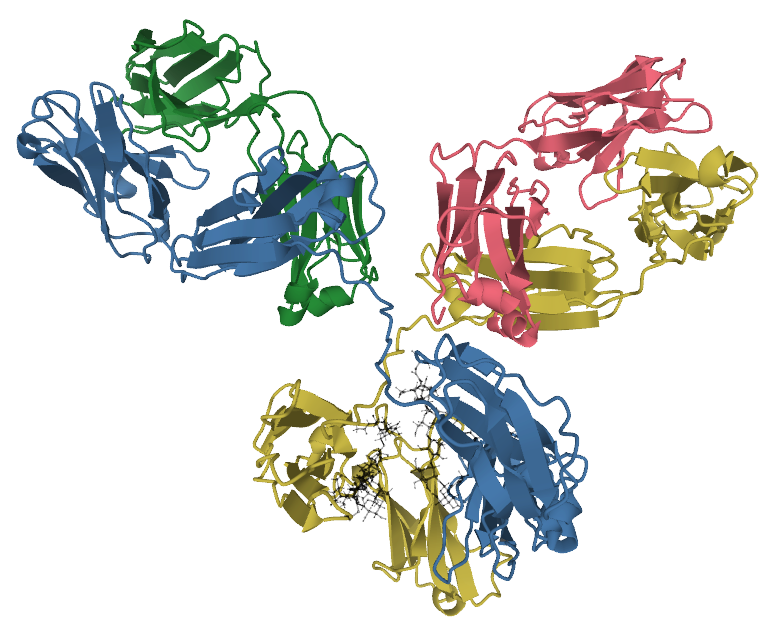
The evolutionary origin of V(D)J recombination has been traced to a mobile genetic element — a DNA transposon. In 1998, Agrawal, Eastman, and Schatz demonstrated that RAG1 and RAG2 together possess transposase activity: they can excise a DNA segment flanked by recombination signal sequences (RSSs) from a donor molecule and insert it into a target molecule, producing the short target-site duplications characteristic of all known transposition reactions.11 This experiment established that the RAG recombinase is, at its core, a domesticated transposase.
In 2005, Kapitonov and Jurka provided the first direct phylogenetic link between RAG1 and a specific transposon superfamily. They showed that the approximately 600-amino-acid catalytic core of RAG1 is significantly similar to the transposase encoded by Transib transposons, a superfamily of DNA transposons found in insects, sea urchins, and other invertebrates. They further demonstrated that the recombination signal sequences recognized by RAG are derived from the terminal inverted repeats of an ancestral Transib element.12
The discovery of ProtoRAG in the lancelet (Branchiostoma belcheri), the most basal extant chordate, provided a direct window into the ancestral state. Huang and colleagues reported in 2016 that ProtoRAG is an active DNA transposon flanked by terminal inverted repeats resembling RSSs, containing RAG1-like and RAG2-like genes oriented tail to tail. The ProtoRAG proteins function as an active endonuclease and transposase both in vitro and in living cells, with striking mechanistic similarities to vertebrate RAG.13 ProtoRAG demonstrates that a RAG-like transposon existed in the chordate lineage before the origin of jawed vertebrates and the adaptive immune system.
Structural studies using cryo-electron microscopy by Zhang and colleagues in 2019 compared the ProtoRAG transposase with the vertebrate RAG recombinase and identified specific molecular changes — including arginine 848 in RAG1 and an acidic region in RAG2 — that suppress transposition activity more than 1,000-fold in jawed vertebrates.14 These modifications represent the molecular domestication of a selfish genetic element into a precisely regulated recombinase essential for immunity. Carmona, Fugmann, and Schatz further demonstrated that RAG2 was recruited to collaborate with RAG1-like proteins during the evolutionary transition from transposon to recombinase, adding another layer of stepwise complexity.23
The evolutionary narrative is now well established: a Transib-family transposon inserted into or near an ancestral antigen receptor gene in the germline of an early jawed vertebrate, splitting the gene into segments. The transposase was subsequently domesticated through mutations that suppressed its transposition activity while preserving its ability to cut and rejoin DNA at the signal sequences, giving rise to the V(D)J recombination machinery that generates antibody and T-cell receptor diversity.11, 12, 13, 14
Scaffolding and molecular exploitation
Beyond the three specific systems Behe cited, experimental molecular biology has demonstrated general mechanisms by which apparently irreducibly complex molecular interactions can evolve. One of the most compelling demonstrations comes from the work of Joseph Thornton and colleagues on the evolution of steroid hormone receptors.
Using ancestral gene resurrection — the phylogenetic inference, synthesis, and experimental characterization of extinct proteins — Thornton's laboratory has reconstructed the evolutionary history of the mineralocorticoid receptor (MR) and the glucocorticoid receptor (GR).20 In 2006, Bridgham, Carroll, and Thornton showed that the specific functional partnership between the hormone aldosterone and the MR evolved through molecular exploitation: the ancestral receptor, which predated the hormone, already possessed structural affinity for aldosterone as a by-product of its binding to older, chemically similar ligands. A gene duplication then produced the MR and GR, and subsequent mutations in GR abolished its sensitivity to aldosterone while maintaining cortisol responsiveness.16
This study demonstrates the scaffolding model in action at the molecular level. The ancestral receptor's promiscuous ligand-binding ability served as a scaffold that permitted the new hormone-receptor partnership to evolve, after which further mutations refined and restricted the specificity of the system. The result is a modern hormone-receptor pair that appears to require both components simultaneously, but which was in fact built through a stepwise process in which each intermediate was functional.16
The broader principle is that complex molecular interactions routinely evolve through the recruitment of pre-existing molecules into new functional relationships — a process that Thornton has termed molecular exploitation.20 This principle applies generally to the evolution of biological complexity and provides a well-supported mechanistic alternative to the claim that multi-component systems require simultaneous assembly of all parts.
The Kitzmiller v. Dover ruling
The scientific rejection of irreducible complexity received its most prominent public airing in the 2005 federal trial Kitzmiller v. Dover Area School District, in which parents challenged the Dover, Pennsylvania school board's policy requiring that intelligent design be presented as an alternative to evolution in ninth-grade biology classes. The trial, presided over by Judge John E. Jones III of the United States District Court for the Middle District of Pennsylvania, included extensive expert testimony on irreducible complexity from both proponents and critics.15
Behe, testifying as the lead expert witness for the defence, presented his arguments about the bacterial flagellum, the blood clotting cascade, and the immune system. On cross-examination, he was confronted with fifty-eight peer-reviewed publications, nine books, and several immunology textbook chapters documenting the evolution of the immune system. Behe acknowledged the existence of this literature but maintained that it was insufficient to explain the origin of the adaptive immune system.15
In his 139-page decision, Judge Jones ruled that intelligent design is not science and that the school board's policy violated the Establishment Clause of the First Amendment. On the specific question of irreducible complexity, the court found that "Professor Behe's claim for irreducible complexity has been refuted in peer-reviewed research papers and has been rejected by the scientific community at large." The court further found that the argument employed "the same flawed and illogical contrived dualism" as earlier creationist arguments — that is, it treated the failure to explain a system by evolution as automatic evidence for design, rather than recognizing that evolutionary explanations may simply not yet be complete.15
The court also noted that Behe's definition of "theory," as applied to intelligent design, was so expansive that it would encompass astrology, and that there were no peer-reviewed articles by any author advocating for intelligent design that were supported by pertinent experiments or calculations providing detailed, rigorous accounts of how intelligent design of any biological system occurred.15
Scientific consensus
The scientific community's rejection of irreducible complexity is longstanding and unequivocal. The National Academy of Sciences and the Institute of Medicine, in their 2008 publication Science, Evolution, and Creationism, stated that the failure to identify a specific evolutionary mechanism for a complex structure does not validate intelligent design, and that the study of evolution remains "one of the most active, robust, and useful fields in all of biology."17
The central problem with irreducible complexity, as the accumulated research demonstrates, is not that the systems Behe described are simple — they are genuinely complex and sophisticated. The problem is that the inference from present-day complexity to the impossibility of evolutionary assembly does not follow. The bacterial flagellum shares components with the type III secretion system.3 The blood clotting cascade was built through serial gene duplication, and organisms with simpler versions clot effectively.7, 8, 9 The adaptive immune system was assembled from a domesticated transposon.11, 12, 13 In every case, the components had prior functions, the system was built incrementally, and intermediate stages were operational. Irreducible complexity, as a scientific argument, has been refuted by the evidence it claimed would never be found.15, 17
References
The non-flagellar type III secretion system evolved from the bacterial flagellum and diversified into host-cell adapted systems
The evolution of vertebrate blood coagulation as viewed from a comparison of puffer fish and sea squirt genomes
Transposition mediated by RAG1 and RAG2 and its implications for the evolution of the immune system
RAG1 core and V(D)J recombination signal sequences were derived from Transib transposons
Reconstructing the evolution of vertebrate blood coagulation from a consideration of the amino acid sequences of clotting proteins
Evolutionary links between FliH/YscL-like proteins from bacterial type III secretion systems and second-stalk components of the F₀F₁ and vacuolar ATPases
Collaboration of RAG2 with RAG1-like proteins during the evolution of V(D)J recombination
Bioinformatics, genomics and evolution of non-flagellar type-III secretion systems: a Darwinian perspective