Overview
- Natural selection has acted on human populations within the past 10,000 years, producing genetically distinct adaptations to local environments — among the clearest examples are lactase persistence in dairying cultures, EPAS1-mediated altitude tolerance in Tibetans derived from Denisovan introgression, and multiple independent mutations conferring partial resistance to malaria in populations across sub-Saharan Africa and the Mediterranean.
- Many adaptive traits have evolved convergently in separated populations facing the same selective pressure: lactase persistence arose independently at least three times, high-altitude hypoxia tolerance has been solved differently in Tibetans, Andeans, and Ethiopians, and at least six distinct genetic mechanisms protect against malaria across different endemic regions.
- The breadth and recency of these adaptations demonstrate that human evolution did not cease with the emergence of anatomically modern Homo sapiens; populations under strong and sustained environmental pressure continue to evolve, and the collective pattern reveals a species still actively shaped by natural selection.
Human genetic adaptation refers to the process by which anatomically modern Homo sapiens has undergone measurable evolutionary change in response to local environmental pressures during the Holocene and Late Pleistocene. The view that natural selection ceased to act meaningfully on our species once cultural buffering — fire, clothing, medicine, agriculture — reduced the direct impact of environmental hazards has not been borne out by genomic evidence. Whole-genome surveys of human populations consistently identify signatures of recent positive selection, the telltale patterns of allele-frequency change and reduced haplotype diversity that reveal genetic variants swept to high frequency because they improved survival or reproduction in a particular ecological context.12, 15 These adaptations span domains as varied as diet, altitude, pathogen exposure, and ultraviolet radiation, and they collectively demonstrate that the human species has diversified genetically in step with the ecological diversity of the environments it has colonised.
Several properties of these adaptive events are particularly instructive. Many involve not single-point mutations but copy number variation, regulatory changes, or entire haplotypes transferred from other hominin lineages. Many have evolved convergently — the same functional outcome achieved by different molecular means in populations that faced the same selective pressure independently and in geographic isolation. This convergence is one of the strongest pieces of evidence that selection, rather than genetic drift, is responsible: when three populations on three continents independently arrive at the same phenotypic solution, the probability that chance alone explains the outcome is vanishingly small.12
Lactase persistence and the dairying revolution
Lactase is an intestinal enzyme that cleaves lactose, the primary sugar in mammalian milk, into glucose and galactose that can be absorbed across the gut wall. In virtually all mammals, including most humans, the gene encoding lactase (LCT) is downregulated sharply after weaning, and adults are therefore unable to digest fresh milk without experiencing the bloating, cramping, and diarrhoea of lactose intolerance. Lactase persistence — the retention of high lactase activity into adulthood — is a derived trait absent in hunter-gatherer populations and common only in populations with long histories of cattle, sheep, or camel pastoralism.1
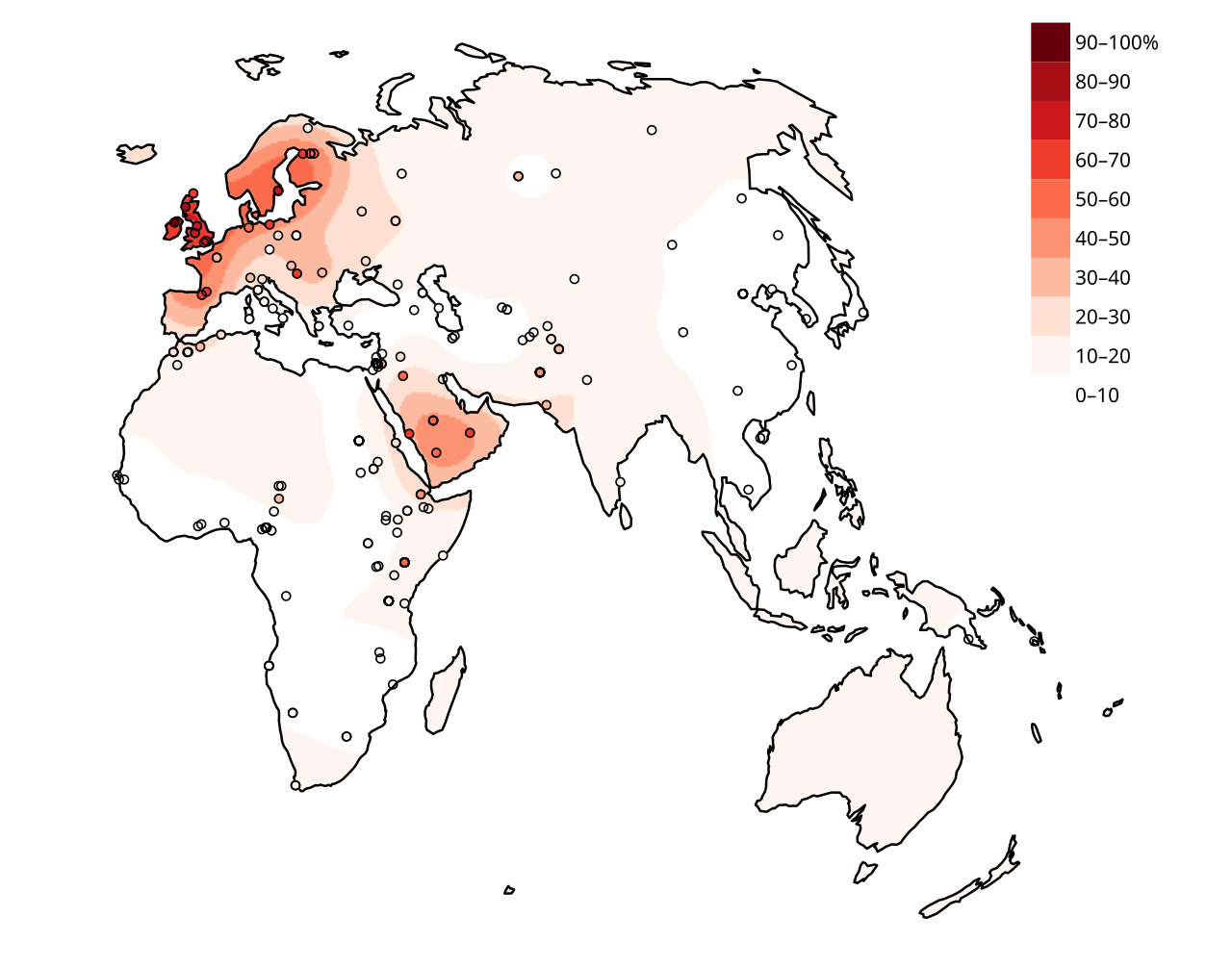
The dominant European variant responsible for this trait is a single C-to-T transition at position −13,910 upstream of LCT, in a regulatory region that controls gene transcription. This mutation, designated LCT*−13910T, disrupts a binding site that would otherwise silence LCT expression in adults. The allele is found at frequencies exceeding 80 percent in populations of northern and western European descent but is nearly absent in East Asian and most African populations. Population genetic analyses estimate that this variant rose from rare frequency to near-fixation in Europe in roughly 5,000 to 10,000 years, implying a selection coefficient of 0.01 to 0.05 — among the strongest sustained selective pressures documented in the modern human genome.1
Crucially, the European variant is not the only solution to the problem of adult lactose digestion.2 Tishkoff and colleagues identified three independent regulatory mutations in East African pastoralist populations — at positions −13,915, −14,010, and −13,907 — that confer lactase persistence by distinct molecular mechanisms and carry distinct haplotypic backgrounds, confirming that they arose as independent mutational events rather than shared descent from a single common ancestor.2 A fourth independent variant has been identified in populations of the Arabian Peninsula. The geographic distribution of each variant maps closely onto the history of pastoralism in its region, with the East African mutations correlating with the spread of cattle-herding cultures in the Sahel and East African Rift zones beginning approximately 7,000 years ago. This is a textbook case of convergent evolution: the same trait, achieved by different mutations, in populations facing the same selective challenge — the nutritional advantage of being able to consume the calorie- and calcium-rich milk available from domesticated livestock.2, 12
High-altitude adaptation: three solutions to hypoxia
The permanent settlement of high-altitude environments — the Tibetan Plateau above 4,000 metres, the Andean altiplano above 3,500 metres, and the Ethiopian highlands above 3,000 metres — poses one of the most severe physiological challenges humans have faced during recent history. At these elevations, the partial pressure of oxygen is 40 to 60 percent of sea-level values, and the low-oxygen (hypoxic) environment threatens aerobic metabolism, pregnancy outcomes, and infant survival. Despite occupying comparable altitudes, Tibetan, Andean, and Ethiopian highlanders have adapted through largely distinct genetic mechanisms, providing an unusually clear natural experiment in convergent evolution.4, 5, 6
The Tibetan adaptation centres on the EPAS1 gene, which encodes one of the subunits of hypoxia-inducible factor 2 (HIF-2α), a master transcriptional regulator of the cellular oxygen-sensing pathway. A distinctive haplotype of EPAS1 found in more than 80 percent of Tibetans is associated with blunted erythropoietic responses to hypoxia — that is, Tibetans do not raise haemoglobin concentration as sharply as lowlanders at altitude, which prevents the blood-thickening that leads to chronic mountain sickness and vascular complications.4 The origin of this haplotype is remarkable: it was shown to derive from Denisovan introgression, having been incorporated into the Tibetan gene pool from archaic humans who occupied the region long before the Holocene.3 The haplotype is essentially absent in Han Chinese lowlanders, the population most closely related to Tibetans, demonstrating that it was present at low frequency in the ancestral population and rose dramatically under the selection pressure imposed by high-altitude habitation.
Andean highlanders arrived at altitude tolerance by a different route. Genome-wide scans of Quechua and Aymara populations have identified signals of selection at loci including EGLN1, PKLR, and NOS2, which are involved in oxygen delivery, erythrocyte metabolism, and nitric oxide signalling respectively. Unlike Tibetans, Andean populations tend to maintain elevated haemoglobin concentrations at altitude, tolerate the higher blood viscosity, and compensate through other cardiovascular adjustments.5 The adaptive architecture is therefore functionally convergent — both populations survive and reproduce at altitude more effectively than un-adapted lowlanders — but mechanistically distinct, operating through different steps of the same physiological pathway.
Ethiopian highlanders present yet a third pattern. Selection signals in Ethiopian genomes have been detected at BHLHE41, ADORA1, and other loci not implicated in either the Tibetan or Andean adaptations, and the physiological profile differs again: Ethiopian highlanders maintain relatively normal haemoglobin levels and achieve altitude tolerance partly through increased red blood cell volume and altered nitric oxide bioavailability.6 That three geographically isolated human populations, each colonising high-altitude environments at different times over the past 10,000 to 25,000 years, have each evolved measurable genetic adaptations to hypoxia through distinct molecular mechanisms stands as compelling evidence that altitude imposes a genuine and persistent fitness cost in the absence of genetic compensation.
Malaria resistance: a catalogue of independent solutions
No infectious disease has exerted stronger and more sustained selection on human populations than malaria caused by Plasmodium falciparum, the protozoan parasite transmitted by Anopheles mosquitoes. Malaria remains one of the leading causes of childhood mortality in sub-Saharan Africa, and the history of its endemicity in tropical and subtropical regions over the past several thousand years is written clearly in the genomes of the populations most exposed to it. At least six distinct genetic mechanisms conferring partial resistance to malaria have been identified in human populations, each geographically localised to malaria-endemic regions and maintained by balancing or positive selection.7, 9, 10
The most extensively studied is sickle cell disease, caused by a point mutation in codon 6 of the HBB gene that substitutes valine for glutamic acid, producing the aberrant haemoglobin S (HbS). Homozygotes for this allele develop severe haemolytic anaemia, but heterozygous carriers — those with sickle cell trait — have measurably reduced parasitaemia and are substantially protected against severe malaria and malaria-associated death. The distribution of the HbS allele across sub-Saharan Africa, the Mediterranean, and South Asia tracks malaria endemicity with striking fidelity, and the allele has risen to frequencies of 10 to 25 percent in hyperendemic zones despite the severe fitness cost imposed on homozygotes. Allison documented this protective association as early as 1954, in one of the earliest demonstrations of balancing selection in humans.7
Haemoglobin C (HbC), produced by a different mutation at the same codon 6 position, is geographically concentrated in West Africa, particularly in Burkina Faso and surrounding populations.8 Unlike HbS, HbC has minimal pathological effects in homozygotes and confers protection against malaria in both heterozygous and homozygous carriers through a mechanism involving altered display of parasite-derived antigens on the red cell surface.8 The geographic separation of HbS (broadly distributed across the malaria belt) and HbC (largely confined to West Africa) reflects the independent origin of each mutation and the regional history of malaria transmission intensity. Haemoglobin E (HbE), yet another variant at a different position in HBB, is the most common structural haemoglobin variant in Southeast Asia and similarly associated with reduced malaria severity.
Alpha-thalassaemia, caused by deletions of one or both copies of the alpha-globin genes on chromosome 16, is the world's most common monogenic disease, reaching frequencies of 50 to 80 percent in malaria-endemic populations of sub-Saharan Africa, the Mediterranean, the Middle East, and Southeast Asia. The protective mechanism appears to involve altered red cell properties that impair parasite invasion and development, and the geographic and demographic distribution of the deletions is strongly correlated with historical malaria endemicity.16
Glucose-6-phosphate dehydrogenase (G6PD) deficiency, encoded by the X-linked G6PD gene, affects more than 400 million people worldwide and reaches frequencies of 20 to 30 percent in some malaria-endemic regions. Deficient red cells produce inadequate amounts of the antioxidant cofactor NADPH, making them more susceptible to oxidative stress and hostile to Plasmodium replication. Because the gene is X-linked, hemizygous males are fully deficient while heterozygous females show a mosaic phenotype, and it is heterozygous females who appear to benefit most from malaria protection relative to fitness cost. Over 180 distinct G6PD variants conferring deficiency have been described, and the multiplicity of independent mutations across populations in Africa, the Mediterranean, and Asia confirms convergent selection.9
A qualitatively different mechanism involves the Duffy blood group antigen, encoded by the DARC gene, which serves as the primary receptor on red blood cells for Plasmodium vivax, a second malaria-causing species. A regulatory mutation that ablates DARC expression on red blood cells (while preserving it on other cell types) is found at frequencies approaching fixation across sub-Saharan Africa, rendering those populations almost entirely resistant to P. vivax infection. This is perhaps the most complete selective sweep documented in the human genome, with the Duffy-negative allele rising to near-100 percent frequency in West and Central Africa in a pattern that leaves no ambiguity about the selective agent.10
Genetic adaptations to malaria by region and mechanism7, 8, 9, 10, 16
| Variant | Gene | Primary region | Peak frequency | Mechanism |
|---|---|---|---|---|
| HbS (sickle cell) | HBB | Sub-Saharan Africa, S. Asia | ~25% | Impairs parasite replication in RBCs |
| HbC | HBB | West Africa | ~15% | Alters antigen display on RBC surface |
| HbE | HBB | Southeast Asia | ~50% | Reduced RBC surface area; instability |
| Alpha-thalassaemia | HBA1/2 | Africa, Mediterranean, SE Asia | ~80% | Impaired RBC invasion by parasite |
| G6PD deficiency | G6PD | Africa, Mediterranean, Asia | ~30% | Oxidative stress hostile to parasite |
| Duffy-negative | DARC | West and Central Africa | ~100% | Eliminates P. vivax receptor |
Skin pigmentation and the ultraviolet radiation gradient
Human skin colour varies more dramatically across geographic populations than almost any other visible trait, and this variation is the product of natural selection acting on the balance between two conflicting ultraviolet (UV) radiation pressures. Strong UV radiation at equatorial latitudes degrades folate, a B vitamin critical for DNA synthesis and fetal neural tube development, favouring heavy melanisation as a photoprotectant. At high latitudes, reduced UV radiation limits cutaneous vitamin D synthesis, placing a premium on lighter skin that permits adequate photoproduction of vitamin D with scarce sunlight.17 The latitudinal gradient of skin reflectance observed in human populations closely mirrors the gradient of annual UV irradiance, a pattern consistent with selection rather than genetic drift as the primary driver of variation.
The genetic basis of this variation has been substantially resolved. The SLC24A5 gene, which encodes a cation exchanger involved in melanosome biogenesis, carries a derived Ala111Thr variant that is nearly fixed in European and South Asian populations but nearly absent in African and East Asian populations. This single variant accounts for approximately 25 to 38 percent of the difference in skin melanin content between European and West African individuals and shows one of the strongest signatures of positive selection in the human genome, with an estimated selective sweep beginning roughly 6,000 to 12,000 years ago in European ancestral populations.11 A related gene, SLC45A2, shows similar patterns of population differentiation and contributes an additional component of the European depigmentation phenotype.
Variation at the MC1R gene, which encodes the melanocortin 1 receptor that switches melanin synthesis between the photoprotective eumelanin and the less protective phaeomelanin, underlies much of the variation in hair and skin colour among European populations and is associated with red hair, fair skin, and increased UV sensitivity. Unlike SLC24A5, MC1R shows high diversity in Europeans but is highly conserved in African populations, a pattern consistent with strong purifying selection maintaining dark pigmentation at low latitudes and relaxed constraint at high latitudes permitting the accumulation of loss-of-function variants.17 The full picture of skin colour genetics involves dozens of loci, but the strongest selective signals at SLC24A5, SLC45A2, and MC1R collectively document ongoing adaptation to the UV environment encountered during the out-of-Africa dispersal into Europe and Asia.11, 17 A more detailed treatment of skin colour genetics appears in the companion article on the evolution of human skin color.
Amylase copy number and starch digestion
Copy number variation — the presence of more or fewer copies of a genomic segment than the standard diploid complement — provides a rapid mechanism for quantitative adaptation when more enzyme product would be beneficial. Salivary amylase, encoded by the AMY1 gene on chromosome 1, initiates starch digestion in the mouth, breaking down the α-1,4 glycosidic bonds of amylopectin into maltose and smaller oligosaccharides before the food bolus reaches the intestine. Humans show extraordinary variation in AMY1 copy number, ranging from two to eighteen copies per diploid genome.13
Perry and colleagues demonstrated that this variation is non-random with respect to diet: populations with high-starch diets — agricultural societies subsisting on cereals and root vegetables — carry on average nearly twice as many AMY1 copies as populations with traditionally low-starch diets, such as the Biaka forest foragers of Central Africa or the Yakut pastoralists of Siberia.13 Higher AMY1 copy number correlates with higher salivary amylase protein concentration and activity, which facilitates the rapid hydrolysis of starch and may improve glucose absorption and glycaemic regulation from starchy foods. The increase in AMY1 copy number in agricultural populations began no earlier than the Neolithic, consistent with the hypothesis that the transition to cereal agriculture imposed new selection on digestive capacity. This is a striking example of gene-culture co-evolution: a human cultural innovation — the cultivation and consumption of starchy crops — created a new selective environment that in turn drove genetic adaptation.13
Alcohol metabolism and ADH1B
Ethanol is metabolised primarily by alcohol dehydrogenases (ADH), a family of enzymes that oxidise ethanol to acetaldehyde, which is then further oxidised to acetate by aldehyde dehydrogenase (ALDH). A coding variant in the ADH1B gene, Arg47His (sometimes designated ADH1B*47His or ADH1B*2), produces an enzyme with approximately 40 to 70 times greater catalytic activity than the ancestral form, resulting in more rapid conversion of ethanol to acetaldehyde. Because acetaldehyde is toxic and causes flushing, nausea, and tachycardia, carriers of the fast-acting allele who consume alcohol experience an unpleasant response that acts as an aversive conditioning signal.14
The ADH1B*47His allele reaches frequencies of 70 to 90 percent in East Asian populations but is uncommon in European and African populations. Population genetic analyses have identified a signal of positive selection at this locus in East Asian genomes, with the allele apparently rising steeply in frequency during or after the Neolithic expansion of rice agriculture in China approximately 7,000 to 9,000 years ago.14 The selective advantage of an allele that discourages alcohol consumption is plausibly connected to the production of fermented beverages from rice: as alcohol became increasingly available with the development of brewing technology, selection may have favoured genotypes that limited excessive consumption and the associated mortality and morbidity. Whether the primary mechanism is protection from alcohol toxicity directly, or some other correlated physiological effect of rapid acetaldehyde production, remains under investigation, but the population-genetic signal for selection is robust.
What these examples tell us about human evolution
The examples surveyed in this article are united by a set of recurring themes that illuminate the dynamics of human adaptation more broadly. The most important is the reality of ongoing evolution. Each of the adaptations described — lactase persistence, altitude tolerance, malaria resistance, depigmentation, elevated amylase, rapid alcohol metabolism — has a detectable selective history within the past 10,000 years, and in many cases the sweep is estimated to have occurred within the past 5,000 years. Selection coefficients sufficient to drive these changes are on the order of 0.01 to 0.10 per generation, well within the range that population genetics predicts can produce rapid allele-frequency shifts in populations of realistic size. The assumption that cultural innovation freed humans from evolutionary pressure is not supported by the genomic evidence; rather, cultural changes such as agriculture, pastoralism, and high-altitude settlement created novel selective environments that drove new rounds of genetic adaptation.12, 15
The prevalence of convergent evolution is equally significant. Lactase persistence arose at least four times independently; hypoxia tolerance has evolved through different molecular mechanisms in three separate highland populations; at least six distinct genetic variants independently protect against malaria across populations that were geographically separated during the period of adaptation. This convergence is the hallmark of natural selection rather than genetic drift: drift produces different outcomes in different populations, while selection repeatedly favours any mutation that achieves the adaptive phenotype, regardless of molecular mechanism.2, 12
Several adaptations also illustrate gene-culture co-evolution, the dynamic in which human cultural practices alter the selective environment for genetic variants, which in turn influence the cultural practices that spread most successfully. The tight correlation between dairying cultures and lactase persistence, and between starch-based agriculture and high AMY1 copy number, demonstrates that biological and cultural evolution in humans are not independent processes but reciprocally entangled ones. Culture changes the genome; the genome shapes which cultures are viable.1, 2, 13
The malaria resistance variants highlight another dimension: balancing selection and the cost of adaptation. Most of the haemoglobin variants that protect against malaria impose substantial costs in homozygotes — sickle cell anaemia, haemoglobin C disease, severe thalassaemia. Natural selection maintains these variants in populations not because they are beneficial in all genotypic configurations but because the heterozygote advantage outweighs the homozygote cost in environments where malaria is lethal without protection. These are not perfect adaptations but compromises, and they illustrate that evolution does not produce optimal solutions but only solutions sufficient to outcompete the alternatives available in a given population at a given time.7, 9
Finally, the EPAS1 case in Tibetans introduces a dimension that underscores the complexity of human evolutionary history: the role of archaic introgression as a source of adaptive variation. Tibetans did not wait for a beneficial mutation to arise de novo; they acquired a pre-adapted allele from an archaic population — the Denisovans — that had been resident at high altitude far longer. This represents a form of evolutionary shortcut in which gene flow from a divergent lineage provides access to genetic variants shaped by a very different evolutionary history. The frequency with which adaptive introgression contributed to human environmental success across different contexts remains an active research question, but the EPAS1 case demonstrates that the genetic legacy of archaic hominins in modern human genomes is not merely a historical curiosity but a functional contribution to present-day adaptation.
Taken together, the catalogue of human genetic adaptations documents a species that is neither evolutionarily static nor uniquely exempt from the forces that shape all living populations. Each geographically localised genome carries the record of the selective pressures its ancestors encountered — the latitudinal gradient of sunlight, the altitude of the plateau, the pathogen load of the tropical environment, the subsistence strategy of the culture — written in the frequencies of alleles that were or were not favoured. Reading that record is one of the central projects of modern evolutionary genetics, and the picture it reveals is of a profoundly varied but still unified species, diversified at the surface by the demands of local ecology, unified at the core by shared origins and the universal mechanisms of natural selection.12, 15
References
A single nucleotide polymorphism in the LCT gene explains 70% of lactase persistence in northern Europe
Andean adaptations to altitude are not associated with increased haemoglobin concentration
Copy number variation of the salivary amylase gene AMY1 is associated with the number of copies in the genome
The ADH1B Arg47His polymorphism in East Asian populations and expansion of agriculture