Overview
- Endosymbiotic theory, first championed by Lynn Margulis in 1967, holds that mitochondria descended from an alphaproteobacterial endosymbiont and chloroplasts from a cyanobacterial one, a model now supported by overwhelming molecular, phylogenetic, and ultrastructural evidence.
- The acquisition of mitochondria was a singular, transformative event in the history of life that enabled aerobic metabolism in eukaryotes, while primary endosymbiosis of cyanobacteria gave rise to chloroplasts and brought oxygenic photosynthesis into the eukaryotic domain.
- Secondary and tertiary endosymbioses, in which eukaryotes engulfed other plastid-bearing eukaryotes, generated the extraordinary diversity of algal lineages, and ongoing gene transfer from organelles to the nucleus continues to reshape eukaryotic genomes today.
Endosymbiosis is the evolutionary process by which free-living prokaryotic cells were incorporated into ancestral eukaryotic cells and, over hundreds of millions of years, became permanent organelles. The endosymbiotic theory holds that mitochondria descended from an alphaproteobacterial ancestor and that chloroplasts descended from a cyanobacterial ancestor, each having been engulfed by a host cell in an ancient symbiotic partnership that became irreversible.1, 3 First formally proposed by Lynn Margulis in 1967 and initially met with widespread skepticism, endosymbiotic theory is now one of the best-supported frameworks in cell biology, underpinned by convergent lines of evidence from molecular phylogenetics, comparative genomics, ultrastructure, and biochemistry.2, 5 The acquisition of these organelles fundamentally transformed the trajectory of life on Earth: mitochondria enabled the efficient aerobic metabolism that powers virtually all complex eukaryotic life, and chloroplasts brought oxygenic photosynthesis into the eukaryotic domain, giving rise to the plant and algal lineages that sustain most terrestrial and aquatic ecosystems today.5, 6
Margulis and the modern proposal
The idea that organelles might have originated as independent organisms has deep roots. In the late nineteenth and early twentieth centuries, several biologists, including Andreas Schimper, Konstantin Mereschkowski, and Ivan Wallin, noticed that chloroplasts and mitochondria resembled free-living bacteria in size, shape, and mode of division, and speculated that they might have once been autonomous organisms living inside host cells.9, 23 These early proposals were largely ignored or dismissed by the scientific mainstream, which lacked the molecular tools to test them. The hypothesis lay dormant for decades.
In 1967, Lynn Sagan (later Lynn Margulis) published a landmark paper, "On the Origin of Mitosing Cells," in the Journal of Theoretical Biology. In it, she marshalled a comprehensive argument that mitochondria, chloroplasts, and flagella had each originated through serial endosymbiotic events in which free-living prokaryotes were engulfed by a host cell and progressively integrated into its biology.1 The paper was reportedly rejected by more than a dozen journals before finding a publisher, reflecting the deep skepticism with which the hypothesis was regarded at the time.2, 23 Of her three proposed endosymbioses, the evidence for the mitochondrial and chloroplast claims proved overwhelming in subsequent decades, while the spirochete origin of flagella has not been supported and is no longer accepted.2
The turning point came with the advent of molecular biology. When researchers sequenced ribosomal RNA genes from mitochondria and chloroplasts in the 1970s and 1980s, they found that these organellar sequences did not resemble the nuclear genes of the host cell but instead clustered phylogenetically with specific groups of bacteria — alphaproteobacteria in the case of mitochondria and cyanobacteria in the case of chloroplasts.3 This molecular evidence, combined with the ultrastructural observations that Margulis had emphasised, rapidly converted the endosymbiotic theory from a fringe idea into the consensus view. By the early 1990s, it was a cornerstone of cell biology and evolutionary theory.2, 9
The alphaproteobacterial origin of mitochondria
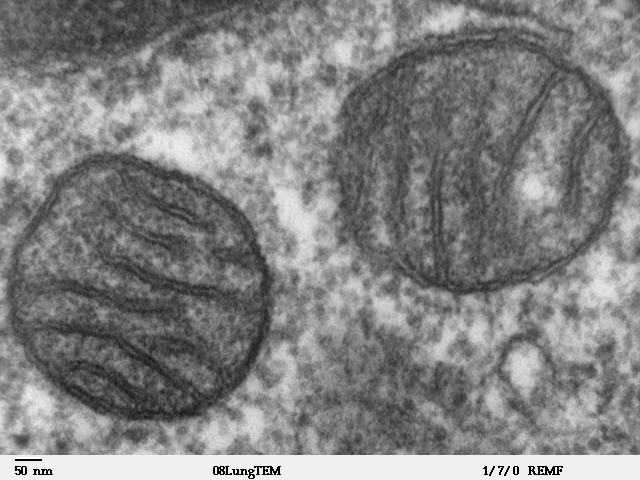
Mitochondria bear an unmistakable suite of bacterial traits that collectively constitute the primary evidence for their endosymbiotic origin. They are bounded by a double membrane, the inner membrane of which is thought to correspond to the original plasma membrane of the engulfed bacterium. They contain their own genome, a small circular DNA molecule that is structurally analogous to a bacterial chromosome rather than to the linear chromosomes of the eukaryotic nucleus. In humans, the mitochondrial genome is approximately 16,569 base pairs in length and encodes 13 proteins, 22 transfer RNAs, and 2 ribosomal RNAs.3, 5 Mitochondria possess their own ribosomes, which are 70S in sedimentation coefficient — the same as bacterial ribosomes — rather than the 80S ribosomes found in the eukaryotic cytoplasm. And mitochondria reproduce by binary fission, dividing independently of the host cell's nuclear division cycle, much as free-living bacteria do.3
Phylogenetic analyses have firmly placed the mitochondrial ancestor within the Alphaproteobacteria, the bacterial class that also includes such well-known genera as Rhizobium, Agrobacterium, and Rickettsia. In 1998, Andersson and colleagues sequenced the complete genome of Rickettsia prowazekii, the obligate intracellular parasite that causes epidemic typhus, and found striking similarities to mitochondrial genomes in gene content, gene order, and metabolic capacity, leading them to describe Rickettsia as the closest known free-living relative of mitochondria at that time.4 Although the precise phylogenetic position of mitochondria within the Alphaproteobacteria remains debated — with different analyses placing them near the Rickettsiales, the SAR11 clade of marine bacteria, or as a deep-branching sister to all sampled alphaproteobacteria — the affiliation with this bacterial class is not in doubt.3, 19
The endosymbiotic event that gave rise to mitochondria appears to have occurred only once in the history of life. All known eukaryotic lineages either possess mitochondria or possess organelles of clear mitochondrial descent that have been secondarily reduced (discussed below), indicating that the last eukaryotic common ancestor already harboured a mitochondrial endosymbiont.6, 19 No primitively amitochondriate eukaryotes have ever been identified, despite decades of searching among anaerobic protists that were once thought to lack mitochondria entirely.6
The cyanobacterial origin of chloroplasts
Chloroplasts arose through an analogous process: a heterotrophic eukaryotic cell, one that already possessed mitochondria, engulfed a cyanobacterium capable of oxygenic photosynthesis.
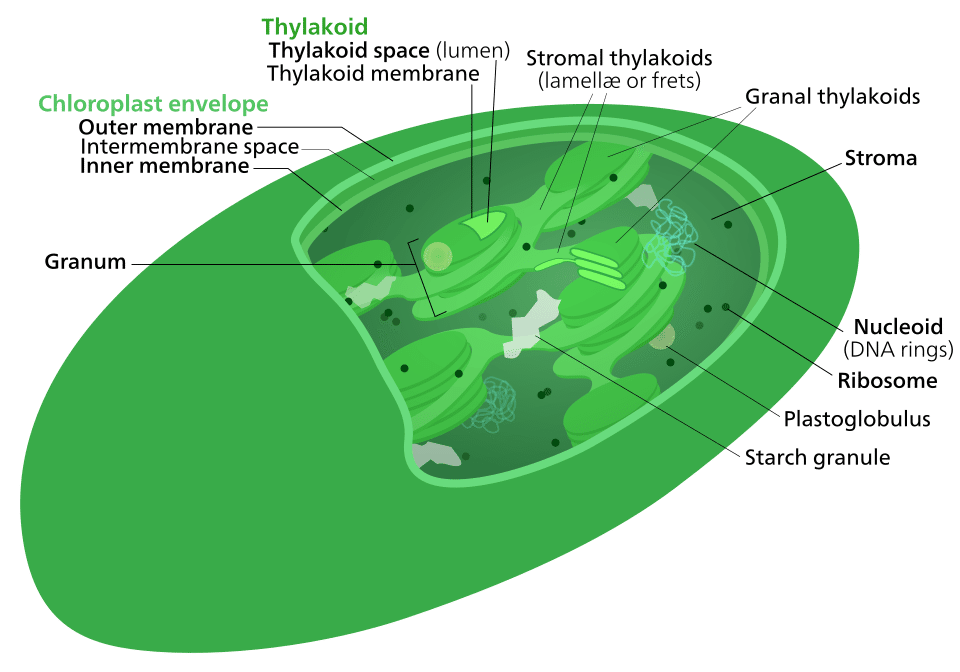
Over time, the cyanobacterium was converted into a permanent organelle, transferring most of its genes to the host nucleus while retaining a reduced genome devoted primarily to photosynthetic functions.10, 11 This process, called primary endosymbiosis, gave rise to the chloroplasts found in three major eukaryotic lineages: the Glaucophyta (glaucophyte algae), the Rhodophyta (red algae), and the Viridiplantae (green algae and land plants).10, 12
The evidence for the cyanobacterial origin of chloroplasts parallels that for mitochondria. Chloroplasts possess a double membrane, their own circular DNA genome (typically 120,000 to 160,000 base pairs in land plants), 70S ribosomes, and they divide by binary fission.11 Phylogenetic analyses of ribosomal RNA sequences, ribosomal proteins, and photosystem components unambiguously place chloroplast genomes within the cyanobacterial radiation. Syntenic gene clusters encoding ribosomal proteins, ATPase subunits, and RNA polymerase subunits are shared between chloroplast and cyanobacterial genomes, a pattern that would be difficult to explain by any mechanism other than descent from a common ancestor.11, 12
A particularly compelling piece of evidence comes from the glaucophyte algae, whose plastids (called cyanelles) retain a thin layer of peptidoglycan between their two bounding membranes, directly recalling the cell wall of free-living cyanobacteria. This retained peptidoglycan layer is widely interpreted as a molecular relic of the original cyanobacterial endosymbiont, preserved in a lineage that diverged early in plastid evolution.10, 12 As with mitochondria, phylogenetic and genomic evidence strongly supports a single origin for primary plastids, meaning that the Glaucophyta, Rhodophyta, and Viridiplantae collectively form a monophyletic group called the Archaeplastida that traces back to one ancestral endosymbiotic event.10
Secondary and tertiary endosymbiosis
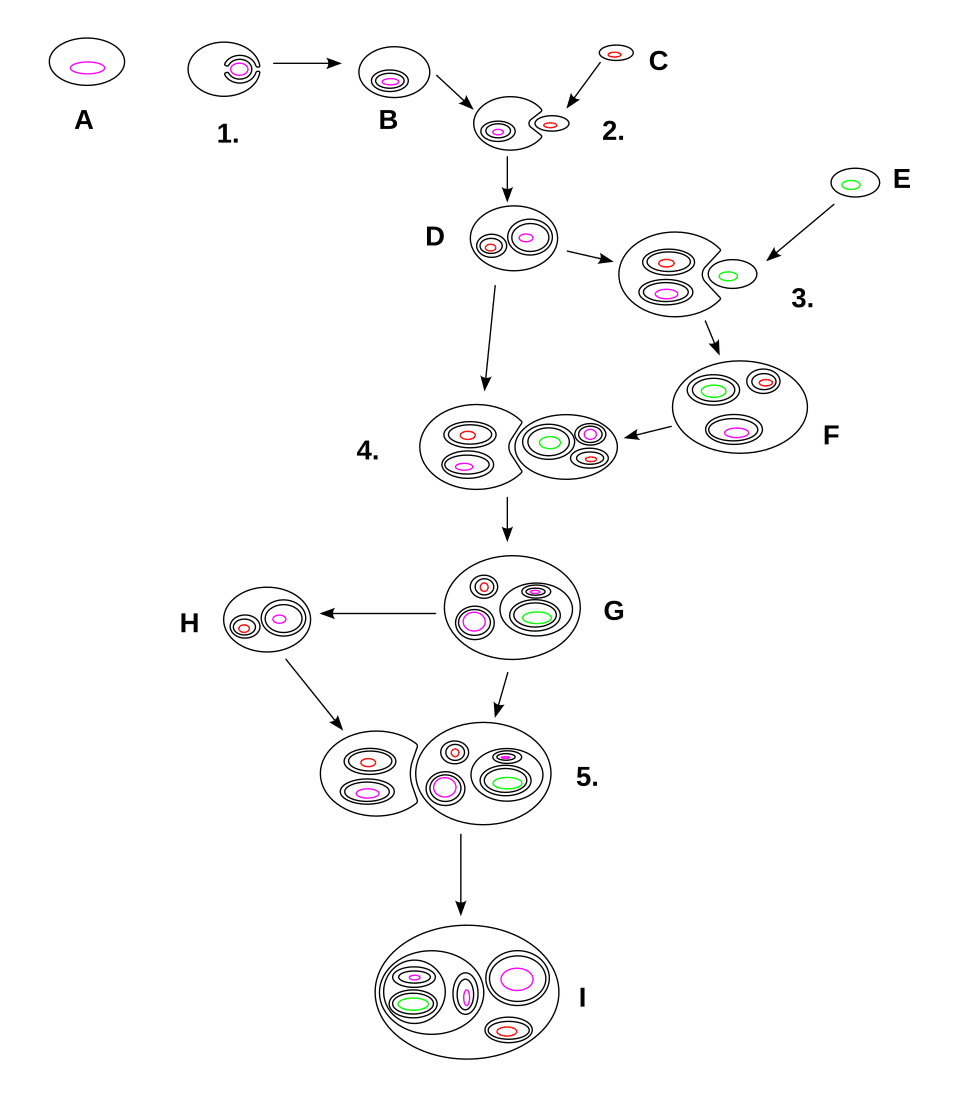
Primary endosymbiosis produced chloroplasts in only the three Archaeplastida lineages, yet photosynthetic organelles are found across a far wider swath of the eukaryotic tree of life, including diatoms, brown algae, dinoflagellates, cryptophytes, haptophytes, euglenids, and chlorarachniophytes. These organisms acquired their plastids not by engulfing a cyanobacterium directly but by engulfing a eukaryote that already possessed a primary chloroplast, in a process called secondary endosymbiosis.10, 13
Secondary endosymbiosis is recognisable by the number of membranes surrounding the resulting plastid. Whereas primary plastids are bounded by two membranes (the outer and inner envelope membranes corresponding to the host phagosomal membrane and the cyanobacterium's plasma membrane, respectively), secondary plastids are typically surrounded by three or four membranes, the additional membranes reflecting the plasma membrane and, in some cases, the endoplasmic reticulum membrane of the engulfed eukaryotic cell.12, 13 In cryptophytes, a remnant of the engulfed alga's nucleus, called a nucleomorph, persists as a tiny organelle between the second and third plastid membranes, providing direct cytological evidence that the plastid was once part of an independent eukaryotic cell.10
Two major classes of secondary endosymbiosis have been identified based on whether the engulfed alga was a green alga or a red alga. Euglenids and chlorarachniophytes acquired their plastids from green algal endosymbionts in two independent events.10, 22 A much larger and more ecologically important group of organisms — including diatoms, brown algae, cryptophytes, haptophytes, and the non-photosynthetic oomycetes — acquired plastids derived from red algal endosymbionts. Whether all red-algal-derived secondary plastids trace to a single endosymbiotic event, as proposed by the chromalveolate hypothesis, or arose through multiple independent uptakes remains actively debated, with recent phylogenomic analyses increasingly challenging the single-event model.22, 13
In some dinoflagellates, the complexity deepens further through tertiary endosymbiosis, in which a dinoflagellate that had already lost or replaced its original secondary plastid engulfed yet another eukaryotic alga, producing a plastid surrounded by as many as five membranes and a cell containing genetic material from multiple distinct evolutionary lineages.10 The genus Durinskia, for example, harbours a diatom endosymbiont with its own nucleus, mitochondria, and chloroplasts, resulting in a single cell with five genetically distinct compartments, each with its own evolutionary history.10
Levels of endosymbiosis and resulting plastid membrane numbers10, 12, 13
| Endosymbiosis type | Endosymbiont engulfed | Plastid membranes | Example lineages |
|---|---|---|---|
| Primary | Cyanobacterium | 2 | Green algae, red algae, glaucophytes, land plants |
| Secondary (green) | Green alga | 3–4 | Euglenids, chlorarachniophytes |
| Secondary (red) | Red alga | 3–4 | Diatoms, brown algae, cryptophytes, haptophytes |
| Tertiary | Eukaryote with secondary plastid | 4–5 | Some dinoflagellates (e.g., Durinskia) |
The host cell debate
While the bacterial identity of the mitochondrial endosymbiont is well established, the nature of the host cell that engulfed it has been one of the most contentious questions in evolutionary biology. For decades, two broad classes of models competed. The traditional archezoan hypothesis, championed by Thomas Cavalier-Smith, proposed that the host was a complex, nucleated, phagocytotic cell — essentially a primitive eukaryote that already possessed a nucleus and endomembrane system but lacked mitochondria. In this view, the eukaryotic cell plan evolved first, and mitochondria were acquired later as a refinement.9, 6 This model predicted the existence of primitively amitochondriate eukaryotes, but as molecular and genomic data accumulated through the 1990s and 2000s, every candidate "archezoan" was shown to harbour mitochondrial remnant organelles or mitochondrial genes in its nuclear genome, effectively falsifying the prediction.6
The alternative class of models holds that the host cell was a prokaryote — specifically an archaeon — and that the acquisition of the mitochondrial endosymbiont was not a late addition but rather the founding event of eukaryogenesis itself. The most influential of these is the hydrogen hypothesis, proposed by William Martin and Miklós Müller in 1998. It envisions the host as a hydrogen-dependent, autotrophic archaeon (similar in metabolism to modern methanogens) that entered into a syntrophic partnership with a facultatively anaerobic alphaproteobacterium that produced molecular hydrogen as a metabolic waste product. The metabolic dependence of the host on the symbiont's hydrogen drove increasingly tight physical association, culminating in the engulfment of the symbiont.8 An attractive feature of the hydrogen hypothesis is that it accounts for the existence of both aerobic mitochondria and anaerobic mitochondrial derivatives (hydrogenosomes) from a single endosymbiotic ancestor, because the original alphaproteobacterial endosymbiont was metabolically versatile.8, 9
A related model, the syntrophy hypothesis, proposed by Purificación López-García and David Moreira, invokes a tripartite metabolic symbiosis among a methanogenic archaeon, a fermentative deltaproteobacterium, and a metabolically versatile alphaproteobacterium that eventually became the mitochondrion. In this scenario, the deltaproteobacterial partner gave rise to the eukaryotic cytoplasm, while the archaeon gave rise to the nucleus.17 Both the hydrogen and syntrophy hypotheses share the fundamental insight that metabolic interdependence, rather than predatory phagocytosis, may have been the driving force behind the initial endosymbiotic association.9, 17
The discovery of the Asgard archaea, beginning with the identification of Lokiarchaeota from deep-sea sediments near a hydrothermal vent field in 2015, dramatically advanced this debate. The genomes of Asgard archaea encode an unexpectedly large repertoire of proteins previously considered unique to eukaryotes, including homologues of the ESCRT membrane-remodelling complex, actin, and ubiquitin-system components.15, 16 Phylogenomic analyses consistently place eukaryotes within or as the sister group of the Asgard archaea, supporting the view that the host cell was an archaeon closely related to this superphylum.16, 20 The expanded genomic sampling of Asgard archaea, including the description of six additional phyla through metagenomic surveys, has further reinforced this relationship, although the exact branching order remains sensitive to analytical methods.20
Gene transfer from organelle to nucleus
One of the most consequential outcomes of endosymbiosis has been the massive transfer of genes from the genomes of the endosymbionts to the nuclear genome of the host cell, a process termed endosymbiotic gene transfer (EGT). Over the roughly 1.5 to 2 billion years since the mitochondrial endosymbiosis, the vast majority of the genes originally present in the alphaproteobacterial genome have been either lost outright or relocated to the host nucleus. Modern mitochondrial genomes are drastically reduced: the human mitochondrial genome retains only 37 genes, compared with the approximately 800 to 1,500 protein-coding genes in a typical free-living alphaproteobacterium.3, 7 Chloroplast genomes have undergone a similar, though somewhat less extreme, reduction, retaining roughly 80 to 120 protein-coding genes compared with the 2,000 to 3,000 found in free-living cyanobacteria.11, 12
The proteins encoded by transferred genes are not lost to the organelle; rather, they are synthesised in the cytoplasm on 80S ribosomes and then imported back into the organelle using specialised protein import machineries, the TOM/TIM complexes of mitochondria and the TOC/TIC complexes of chloroplasts. These import systems, which have no counterpart in free-living bacteria, are eukaryotic innovations that evolved to maintain organelle function despite genome reduction.5, 7 The net result is that 93 to 99 percent of the proteins found in mitochondria and chloroplasts are encoded in the nucleus, synthesised in the cytoplasm, and post-translationally imported, with only a small core of highly hydrophobic membrane proteins and ribosomal RNAs still encoded locally in the organellar genome.7
Endosymbiotic gene transfer is not merely a historical event; it is an ongoing process. Experimental studies in yeast and tobacco have demonstrated that DNA is transferred from organelles to the nucleus at surprisingly high frequencies in laboratory settings.7 Genomic analyses have revealed numerous recent insertions of mitochondrial and plastid DNA into nuclear chromosomes, termed NUMTs (nuclear mitochondrial DNA segments) and NUPTs (nuclear plastid DNA segments), which are found in nearly all eukaryotic genomes examined. In the model plant Arabidopsis thaliana, approximately 18 percent of nuclear genes appear to be of cyanobacterial origin, reflecting the cumulative impact of endosymbiotic gene transfer from the chloroplast ancestor.7, 18 A comprehensive phylogenomic analysis of 55 eukaryotic genomes found that the vast majority of eukaryotic genes of prokaryotic origin entered the nuclear genome in two discrete episodes corresponding to the origins of mitochondria and chloroplasts, followed by extensive differential gene loss among descendant lineages, rather than through a continuous trickle of horizontal gene transfer from diverse bacterial donors.18
Reduced mitochondrial derivatives
Not all mitochondrial descendants look or function like the textbook aerobic organelle. In many eukaryotic lineages that inhabit permanently anoxic or microaerophilic environments, the ancestral mitochondrion has been modified by reductive evolution into functionally and morphologically distinct organelles collectively known as mitochondrion-related organelles (MROs). These include hydrogenosomes, mitosomes, and a spectrum of intermediate forms.14, 19
Hydrogenosomes, found in organisms such as the parabasalid Trichomonas vaginalis and certain anaerobic fungi and ciliates, generate ATP through substrate-level phosphorylation and produce molecular hydrogen and carbon dioxide as metabolic byproducts. They lack the electron transport chain and the citric acid cycle of aerobic mitochondria, and most have lost their genome entirely, but they retain double membranes and key mitochondrial proteins that betray their ancestry.14 Mitosomes, found in diplomonads such as Giardia intestinalis and microsporidians, represent an even more extreme reduction. They do not produce ATP at all and appear to function primarily in iron-sulfur cluster assembly, an essential biochemical process that is the most conserved function across all known mitochondrial derivatives.14, 6
The discovery of hydrogenosomes and mitosomes in lineages once considered "primitively amitochondriate" was a pivotal moment in endosymbiotic theory. Through the 1980s and early 1990s, certain anaerobic protists such as Giardia, Trichomonas, and microsporidia were placed at the base of the eukaryotic tree in molecular phylogenies, and their apparent lack of mitochondria was interpreted as evidence that they had diverged from the eukaryotic lineage before the mitochondrial endosymbiosis occurred. These organisms were called archezoans, and their existence was cited as support for the idea that eukaryotic cell complexity preceded the acquisition of mitochondria.6 However, improved phylogenetic methods revealed that the basal branching of these organisms was an artefact of long-branch attraction, and molecular studies progressively uncovered mitochondrial genes in their nuclear genomes and mitochondrial remnant organelles in their cytoplasm.6, 14 Today, no known eukaryote is considered to be primitively amitochondriate. Every eukaryotic lineage examined possesses either canonical mitochondria or a mitochondrion-related organelle, firmly establishing that the mitochondrial endosymbiosis preceded the diversification of all extant eukaryotes.6, 19
Functional spectrum of mitochondrial derivatives14, 19
Timing of eukaryogenesis
Constraining the date of the endosymbiotic events that gave rise to mitochondria and chloroplasts is inherently difficult, because the fossil record of early eukaryotes is sparse and the molecular clock calibrations carry large uncertainties. Nonetheless, several independent lines of evidence converge on an approximate window for the origin of the eukaryotic cell and its mitochondrion. The oldest fossils that are confidently attributed to eukaryotes, based on cell size, wall ultrastructure, and morphological complexity, appear in the geological record at approximately 1.6 to 1.8 billion years ago (Ga) in Paleoproterozoic and early Mesoproterozoic sediments.21 Molecular clock analyses, which use calibrated rates of molecular evolution to estimate divergence times, generally place the last eukaryotic common ancestor in the range of 1.0 to 1.9 Ga, with most estimates clustering around 1.5 to 2.0 Ga.21, 19
Because all known eukaryotes possess mitochondria or mitochondrial remnants, the mitochondrial endosymbiosis must have occurred at or before the divergence of the last eukaryotic common ancestor, placing it no later than approximately 1.5 to 2.0 Ga.6, 19 The primary chloroplast endosymbiosis appears to have occurred somewhat later, after the divergence of the Archaeplastida from other eukaryotic supergroups. Molecular clock estimates for the origin of primary plastids vary but generally fall in the range of 1.0 to 1.5 Ga.10, 21
These dates place the endosymbiotic events within the Proterozoic eon, a time when atmospheric oxygen levels were rising but had not yet reached modern concentrations, and when the oceans were still chemically stratified with anoxic, often sulfidic deep waters. The ecological conditions of the Proterozoic ocean — with fluctuating oxygen availability and abundant opportunities for metabolic interactions between archaeal and bacterial cells — may have been precisely the environment in which the syntrophic partnerships envisioned by the hydrogen and syntrophy hypotheses could have arisen and deepened into obligate endosymbiosis.8, 17, 21
Transformative impact on life
The endosymbiotic acquisition of mitochondria and chloroplasts is widely regarded as one of the most consequential events in the history of life, comparable in significance to the origin of life itself and the colonisation of land. Aerobic respiration via the mitochondrial electron transport chain and oxidative phosphorylation generates roughly 15 to 18 times more ATP per molecule of glucose than anaerobic fermentation, providing eukaryotic cells with a vastly greater energy budget.5, 8 This energetic surplus is thought to have been a necessary precondition for the evolution of large cell size, complex internal membrane systems, a dynamic cytoskeleton, and ultimately multicellularity, all of which are hallmarks of the eukaryotic grade of cellular organisation.6, 9
The chloroplast endosymbiosis was equally transformative for the biosphere. By incorporating an oxygenic photosynthetic organelle, the ancestors of modern algae and land plants gained the ability to fix atmospheric carbon dioxide into organic matter using sunlight, a capacity that had previously been restricted to cyanobacteria and other prokaryotic phototrophs. The subsequent diversification and ecological expansion of photosynthetic eukaryotes, particularly during the Neoproterozoic and Phanerozoic, profoundly increased global primary productivity, accelerated the oxygenation of the atmosphere and oceans, and established the trophic foundations of modern marine and terrestrial ecosystems.10, 12
Endosymbiosis also introduced a novel mode of evolutionary innovation: the chimeric genome. Modern eukaryotic cells are genetic mosaics, their nuclear genomes containing thousands of genes inherited from the alphaproteobacterial and cyanobacterial endosymbionts alongside the archaeal genes inherited from the host.7, 18 This genomic chimerism, continually reshaped by ongoing organelle-to-nucleus gene transfer, has provided eukaryotes with a uniquely rich genetic toolkit that has been co-opted for functions far beyond the metabolic roles of the original endosymbionts. In this sense, endosymbiosis did not merely add new organelles to an existing cell; it fundamentally rewired the genetic architecture of the eukaryotic lineage and opened evolutionary trajectories that would have been inaccessible to organisms that remained purely archaeal or purely bacterial.9, 13
References
On the origin of mitosing cells: a historical appraisal of Lynn Margulis endosymbiotic theory