Overview
- Glycolysis is likely one of the most ancient metabolic pathways, predating the divergence of the three domains of life and possibly originating in a prebiotic or early cellular context on an anoxic Earth, using substrates available from abiotic chemistry.
- The citric acid cycle and oxidative phosphorylation evolved in stages, with ancestral partial pathways serving biosynthetic rather than energy-generating roles, and were later elaborated into the full aerobic respiratory chain after the rise of atmospheric oxygen.
- Metabolic pathway evolution proceeds through enzyme recruitment (co-option of enzymes from unrelated pathways), gene duplication and divergence, and retrograde pathway extension, producing the highly interconnected metabolic networks observed in modern organisms.
The metabolic pathways that sustain all living organisms — glycolysis, the citric acid cycle, oxidative phosphorylation, photosynthesis, nitrogen fixation, and hundreds of others — are themselves products of evolution. Like morphological structures, metabolic pathways have evolutionary histories that can be reconstructed through comparative biochemistry, phylogenetic analysis of enzyme sequences, and analysis of the chemical logic underlying pathway architecture.5, 14 Understanding how metabolism evolved illuminates the conditions on the early Earth, the constraints that shaped the biochemistry shared by all living organisms, and the mechanisms by which new metabolic capabilities arise.
The antiquity of glycolysis
Glycolysis — the ten-step enzymatic pathway that converts glucose to pyruvate, generating two molecules of ATP and two of NADH per molecule of glucose — is present in virtually all living organisms and is widely regarded as one of the most ancient metabolic pathways.4, 8 Several lines of evidence support this antiquity. Glycolysis is cytoplasmic and requires no membrane-bound organelles, consistent with an origin before the evolution of intracellular compartments. It operates anaerobically, consistent with an origin on an Earth whose atmosphere lacked significant free oxygen. And the enzymes of glycolysis are among the most universally conserved across the three domains of life (Bacteria, Archaea, and Eukarya), although some of the specific enzymes used differ between domains, suggesting that the pathway may have been assembled from non-homologous enzymes performing equivalent chemical transformations in different lineages.4, 7
The question of whether glycolysis evolved as a degradative (catabolic) pathway or was originally part of a biosynthetic (anabolic) network running in reverse is debated. Some researchers have proposed that the earliest carbon metabolism was autotrophic, with the reactions of gluconeogenesis (the reverse of glycolysis) serving to synthesise sugars from simpler carbon compounds available in the prebiotic environment, and that glycolysis evolved later as a means of extracting energy from accumulated organic molecules.3, 16 Others argue that glycolysis itself is ancient, with some of its intermediates potentially formed by abiotic chemistry at alkaline hydrothermal vents.3
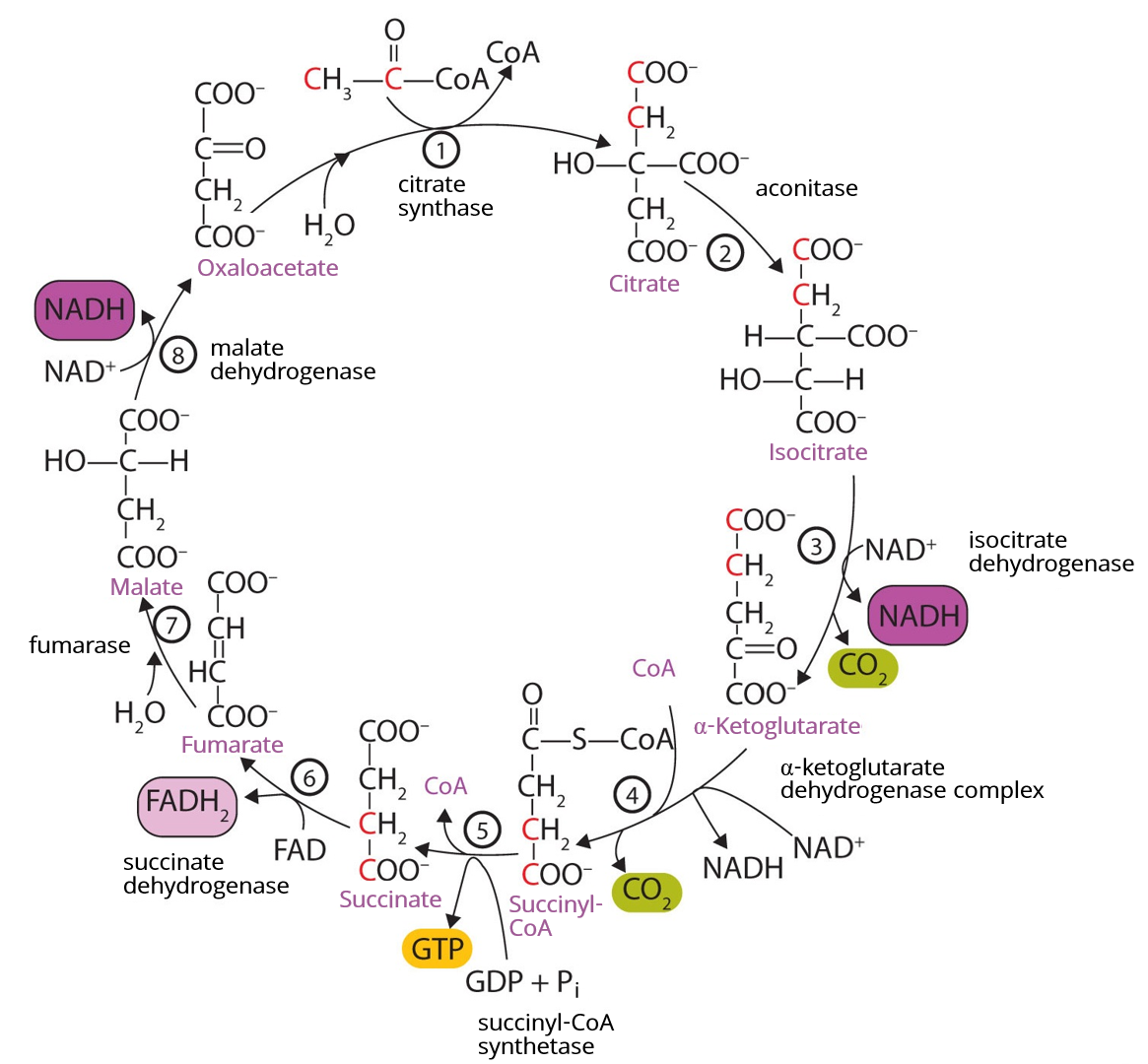
Evolution of the citric acid cycle
The citric acid cycle (also called the tricarboxylic acid or TCA cycle, or the Krebs cycle after its discoverer Hans Krebs) is the central metabolic hub that oxidises acetyl-CoA to CO2, generating reducing equivalents (NADH and FADH2) that feed into the electron transport chain for ATP production.1, 8 Unlike glycolysis, the citric acid cycle in its complete oxidative form requires molecular oxygen as the terminal electron acceptor (via oxidative phosphorylation), raising the question of how it evolved on an anoxic Earth.
Comparative biochemistry has revealed that many organisms possess incomplete or modified versions of the cycle. Some anaerobic bacteria and archaea operate a reductive (reverse) citric acid cycle, in which the reactions run in the opposite direction to fix CO2 into organic molecules rather than oxidise organic molecules to CO2. Smith and Morowitz proposed that the reductive citric acid cycle is the ancestral form, serving as an autocatalytic carbon fixation pathway in the earliest cellular life before being converted to the oxidative direction after the rise of atmospheric oxygen.2
Meléndez-Hevia and colleagues analysed the chemical logic of the citric acid cycle and concluded that it represents the optimal solution for converting a two-carbon unit (acetyl-CoA) into CO2 while maximising the yield of reducing equivalents, given the constraints of biochemistry.6 This optimality argument suggests that the cycle was shaped by natural selection for energetic efficiency. However, the evolutionary pathway from a simple, incomplete proto-cycle to the full eight-enzyme cycle remains uncertain. One proposal is that the cycle was assembled from pre-existing enzymatic reactions that originally served independent biosynthetic functions, with the cyclisation occurring relatively late as a refinement of energy metabolism.6, 9
The evolution of oxidative phosphorylation
Oxidative phosphorylation, the process by which the electron transport chain in the inner mitochondrial membrane (or the plasma membrane of aerobic bacteria) transfers electrons from NADH and FADH2 to molecular oxygen, generating a proton gradient that drives ATP synthesis by ATP synthase, is the most efficient energy-generating pathway in biology, yielding approximately 30 to 32 molecules of ATP per molecule of glucose when coupled with glycolysis and the citric acid cycle.8
The individual components of the electron transport chain — NADH dehydrogenase (Complex I), succinate dehydrogenase (Complex II), the cytochrome bc1 complex (Complex III), and cytochrome c oxidase (Complex IV) — have distinct evolutionary histories and did not arise as a coordinated system. Phylogenetic analyses by Castresana and Saraste showed that the complexes evolved at different times, with some components (such as cytochrome bc1) predating the availability of atmospheric oxygen and originally functioning with alternative electron acceptors such as sulphate, nitrate, or ferric iron.12 Cytochrome c oxidase, which catalyses the terminal reduction of O2 to water, evolved after the Great Oxidation Event approximately 2.4 billion years ago, when the accumulation of photosynthetically produced oxygen made aerobic respiration thermodynamically favourable.11, 12
ATP synthase itself appears to be extremely ancient. The F1Fo-ATP synthase is found in bacteria, archaea, and the mitochondria and chloroplasts of eukaryotes, and its rotary mechanism — in which the flow of protons through the Fo subunit drives the physical rotation of the central stalk, which in turn drives conformational changes in the F1 subunit that catalyse ATP synthesis — is conserved across all domains of life.8 The universality of ATP synthase suggests that chemiosmotic coupling (the use of a proton gradient to drive ATP synthesis) was established very early in the history of life, possibly even before the divergence of the last universal common ancestor (LUCA).3, 17
Mechanisms of metabolic pathway evolution
Three principal mechanisms have been proposed for the evolutionary assembly of metabolic pathways: retrograde evolution, enzyme recruitment (the patchwork model), and gene duplication and divergence.5, 13
The retrograde hypothesis, proposed by Horowitz in 1945, posits that metabolic pathways evolved backwards from their end products. If a cell depends on a particular nutrient available in the environment, and that nutrient becomes depleted, selection favours organisms that can synthesise the nutrient from a precursor. Depletion of the precursor then favours organisms that can synthesise it from an earlier precursor, and so on, extending the pathway one step at a time in the reverse direction from product to substrate.13 While the retrograde hypothesis is logically elegant, empirical evidence for it is limited, and few metabolic pathways show the pattern of sequentially related enzymes that the hypothesis predicts.
The enzyme recruitment (patchwork) model, proposed by Jensen in 1976, holds that early enzymes had broad substrate specificities and could catalyse multiple related reactions. As metabolic needs became more complex, these promiscuous enzymes were recruited into different pathways, and subsequent mutations narrowed their substrate specificities to the reactions of particular pathways.5 This model is supported by the observation that many enzymes in different metabolic pathways share structural folds and catalytic mechanisms despite having no obvious functional relationship, suggesting that they were co-opted from a common ancestral enzyme. The "patchwork" nature of modern metabolic networks, in which the same protein fold families appear in unrelated pathways, is consistent with this model.5, 14
Gene duplication provides the raw material for both retrograde evolution and enzyme recruitment. When a gene encoding a metabolic enzyme is duplicated, one copy can maintain the original function while the other is free to accumulate mutations that alter its substrate specificity, catalytic activity, or regulation, potentially adapting it to a new role in a different pathway. Comparative genomic studies have confirmed that gene duplication has been a major source of metabolic innovation across all domains of life, with paralogous enzyme families populating multiple pathways in modern organisms.10
The evolution of carbon fixation pathways
At least six distinct pathways for autotrophic carbon fixation are known in extant organisms: the Calvin-Benson cycle, the reductive citric acid cycle, the Wood-Ljungdahl (reductive acetyl-CoA) pathway, the 3-hydroxypropionate bicycle, the 3-hydroxypropionate/4-hydroxybutyrate cycle, and the dicarboxylate/4-hydroxybutyrate cycle.15, 16 The existence of multiple independent solutions to the problem of fixing inorganic carbon into organic molecules indicates that carbon fixation has evolved repeatedly, with each pathway reflecting the particular thermodynamic and environmental constraints faced by the organisms that use it.
Braakman and Smith proposed that the reductive citric acid cycle and the Wood-Ljungdahl pathway are the most ancient carbon fixation routes, potentially deriving from abiotic chemistry at hydrothermal vents where geochemically produced H2 and CO2 could drive reductive carbon chemistry.16 The Calvin-Benson cycle, which depends on the enzyme RuBisCO and is the dominant carbon fixation pathway in photosynthetic organisms, is considered a later evolutionary innovation that arose in conjunction with photosynthesis.16
Significance for evolutionary biology
The evolution of metabolism demonstrates that the biochemical machinery of life, like its morphological structures, has been shaped by the same evolutionary forces of mutation, selection, gene duplication, and horizontal transfer. The universality of core metabolic pathways such as glycolysis, the citric acid cycle, and chemiosmotic ATP synthesis across all domains of life provides powerful evidence for common descent, while the patchwork assembly of pathways from enzymes with diverse evolutionary origins illustrates how natural selection can build complex systems incrementally from pre-existing components.5, 14 The history of metabolism also records the history of the Earth: the evolution of anaerobic pathways before aerobic ones, the development of oxygen-using enzymes after the Great Oxidation Event, and the repeated evolution of carbon-concentrating mechanisms in response to declining atmospheric CO2 are all cases where the trajectory of biochemical evolution was shaped by planetary geochemistry.11, 16
References
Evolution of the enzymes of the citric acid cycle and the glyoxylate cycle of higher plants: a case study of endosymbiotic gene transfer