Overview
- Gene flow is the transfer of genetic material between populations through migration, and it acts as a homogenizing force that reduces genetic differentiation among populations while introducing novel alleles that increase local genetic variation.
- The interplay between gene flow and natural selection determines whether populations diverge or remain genetically cohesive: moderate gene flow can spread beneficial alleles across a species' range, while excessive gene flow can swamp locally adapted genotypes and impose a migration load.
- Interspecific gene flow through introgressive hybridization has been a significant source of adaptive variation in numerous lineages, including the transfer of high-altitude EPAS1 alleles from Denisovans to Tibetans and wing-pattern mimicry genes among Heliconius butterflies.
Gene flow, also called gene migration, is the transfer of genetic material from one population to another through the movement of individuals or their gametes. Along with natural selection, mutation, and genetic drift, gene flow is one of the four fundamental mechanisms of evolution, and it plays a distinctive dual role: it acts as a homogenizing force that reduces genetic differentiation among populations while simultaneously introducing novel alleles that may increase the adaptive potential of recipient populations.1, 2 Theodosius Dobzhansky, in his foundational synthesis of genetics and evolutionary biology, recognised migration as a major factor shaping the genetic architecture of species, noting that the movement of individuals between populations prevents them from diverging indefinitely through drift or local selection alone.21 The study of gene flow has since become central to population genetics, conservation biology, and our understanding of how species maintain cohesion across their geographic ranges or, alternatively, how the interruption of gene flow sets the stage for speciation.
Definition and mathematical foundations
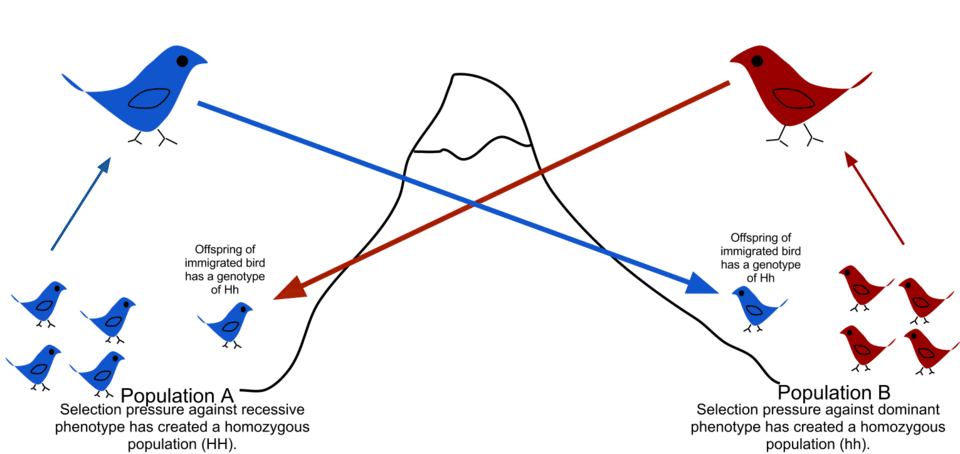
In population genetics, gene flow refers to any mechanism by which alleles are transferred from one population to another. The most straightforward form is the physical immigration of breeding individuals into a new population, but gene flow can also occur through the dispersal of gametes (such as pollen carried by wind or insects in plants), the movement of seeds, or even the horizontal transfer of genetic material between organisms, as occurs routinely among bacteria.1, 11 Regardless of the mechanism, the evolutionary consequence is the same: the introduction of alleles from one gene pool into another, altering allele frequencies in the recipient population.
Sewall Wright's island model, one of the earliest and most influential mathematical treatments of population structure, provides the theoretical foundation for understanding gene flow's effects on genetic differentiation.3 In this model, a species is divided into numerous subpopulations (demes), each of effective size N, that exchange migrants at a rate m per generation. The equilibrium level of genetic differentiation among demes, measured by Wright's fixation index FST, is given by the approximation FST ≈ 1 / (4Nm + 1), where Nm represents the absolute number of migrants exchanged per generation.3 This equation reveals a fundamental insight: even very modest levels of migration can prevent substantial genetic differentiation. When Nm = 1 — that is, when just one migrant enters each subpopulation per generation — the expected FST is only 0.20, meaning that 80 percent of the total genetic variation is shared among populations rather than distributed between them.1, 3
The simplicity of the island model masks considerable complexity in natural populations, where migration rates are rarely uniform, populations differ in size, and gene flow may be asymmetric or restricted by geographic distance. Montgomery Slatkin's influential reviews of gene flow in natural populations demonstrated that real patterns of genetic differentiation reflect the interplay of migration, drift, selection, and the spatial arrangement of populations across the landscape.1, 2 Indirect methods of estimating gene flow from allele-frequency data, while widely used, are subject to the assumptions of the underlying models and may overestimate or underestimate actual migration rates depending on the demographic history and spatial configuration of the populations studied.9 Andrew Bohonak's comprehensive review found that direct estimates of dispersal often diverge from indirect genetic estimates, emphasising the need for integrating ecological data on movement with population genetic analyses.9
Gene flow as a homogenizing force
The most fundamental effect of gene flow is the genetic homogenization of populations. In the absence of natural selection and genetic drift, gene flow between populations drives allele frequencies toward a common equilibrium equal to the average frequency across all connected populations.1 This homogenizing effect means that gene flow opposes the genetic differentiation that would otherwise accumulate through the independent action of drift and local selection in isolated subpopulations. Dobzhansky recognized this principle early in the modern synthesis, arguing that the "cohesion" of a species — the fact that geographically widespread populations remain sufficiently similar to interbreed — depends in large part on the connecting thread of gene flow.21
The homogenizing power of gene flow has been quantified in numerous empirical systems. Morjan and Rieseberg's review of published FST values across a wide range of taxa found that gene flow is considerably higher in many species than earlier studies had suggested, with a large proportion of surveyed species showing FST values below 0.10, indicating that populations share the vast majority of their genetic variation.10 In plants, gene flow through pollen and seed dispersal can operate over surprisingly long distances: Ellstrand's review of gene flow in plants documented cases of pollen-mediated gene flow occurring over kilometres, far exceeding the spatial scale of individual plant neighbourhoods, and concluded that gene flow through both pollen and seed is "idiosyncratic, varying with specific populations involved, and typically occurring at evolutionarily significant rates and distances."12
When gene flow is high enough relative to the effective population sizes of the connected demes, the subpopulations function as a single panmictic (randomly mating) unit, and genetic differentiation is negligible. When gene flow is very low or absent, each population evolves independently, accumulating distinct allele frequencies through drift and local selection.1, 3 The critical threshold lies in the vicinity of one migrant per generation (Nm ≈ 1), below which drift can overwhelm the homogenizing effect of migration and populations diverge substantially.1
Effect of migration rate on population differentiation (FST)1, 3
Gene flow and natural selection
The relationship between gene flow and natural selection is among the most consequential interactions in evolutionary biology. When environmental conditions vary across a species' range, natural selection favours different alleles in different populations, driving local adaptation. Gene flow, by continuously introducing alleles from other populations where different conditions prevail, can either facilitate or constrain this adaptive divergence depending on its magnitude relative to the strength of selection.4, 5
Thomas Lenormand's influential review established that gene flow sets a limit on local adaptation when migrants carry alleles that are advantageous in their source population but deleterious in the recipient population.4 If gene flow exceeds a critical threshold relative to the selection coefficient acting against immigrant alleles, it can "swamp" local adaptation entirely, preventing populations from evolving optimal genotypes for their local environments. This phenomenon produces a migration load — a reduction in the mean fitness of a population caused by the continual influx of maladapted alleles from elsewhere.4, 18 Bolnick and Nosil demonstrated the migration load empirically using Timema cristinae walking-stick insects, which inhabit two host plant species that favour different cryptic colour patterns. Populations receiving more immigrants from the alternative host-plant environment exhibited higher levels of maladaptation and consequently experienced stronger natural selection against the immigrant phenotypes.18
Conversely, gene flow can promote adaptation when it introduces globally advantageous alleles into populations that have not yet acquired them through local mutation. Morjan and Rieseberg argued that given sufficiently strong selection for universally beneficial mutations, gene flow enables their rapid spread across a species' range, allowing species to "evolve collectively" even when individual populations are connected by relatively low levels of migration.10 This spreading function of gene flow is particularly important for large, geographically subdivided species in which the same beneficial mutation is unlikely to arise independently in every population through local mutation alone.
Garant, Forde, and Hendry synthesised the empirical literature on the effects of dispersal and gene flow on contemporary adaptation, finding that the outcome depends critically on the rate of gene flow, the strength and spatial heterogeneity of selection, and the degree of environmental matching between source and recipient populations.5 Low to moderate gene flow tends to increase genetic variation within populations and can supply the raw material for adaptive responses to changing conditions. High gene flow, however, can overwhelm local selection, preventing populations from tracking their local fitness optima and reducing the overall adaptive fit of the species across heterogeneous environments.4, 5
Gene flow and genetic drift
Gene flow and genetic drift are antagonistic forces in the evolution of population structure. Genetic drift causes random fluctuations in allele frequencies that are most pronounced in small populations, leading over time to the loss of alleles and the fixation of others purely by chance. In isolated small populations, drift can rapidly erode genetic diversity and cause substantial differentiation from other populations of the same species.3 Gene flow counteracts these effects by reintroducing alleles that have been lost to drift and by maintaining genetic connectivity among populations.
Wright's mathematical framework established that the balance between gene flow and drift determines the equilibrium level of population differentiation, as captured by the FST equation described above.3 When the number of migrants per generation (Nm) is large, gene flow overwhelms drift and populations remain genetically similar. When Nm is small, drift dominates and populations diverge. The transition between these regimes is remarkably steep: increasing migration from Nm = 0.1 to Nm = 1 — a tenfold increase — reduces the expected FST from approximately 0.71 to 0.20, while a further tenfold increase to Nm = 10 reduces it to only 0.02.1, 3
The counterbalancing effect of gene flow on drift has profound implications for the maintenance of genetic diversity, particularly in small or fragmented populations. Slatkin demonstrated that the geographic structure of genetic variation in natural populations is shaped primarily by the interaction between migration and drift, with selection playing a secondary role for most loci.2 Empirical studies have confirmed this pattern across a wide range of organisms: populations that are well connected by gene flow retain higher levels of heterozygosity and allelic richness than isolated populations of comparable size, because migrants continually replenish the genetic variation that drift removes.9, 10
In the context of human evolution, recent analyses of ancient DNA have quantified the relative contributions of gene flow, selection, and drift to allele frequency change over the past five thousand years in European populations. Mathieson and Terhorst found that gene flow — driven largely by major migration events, including the Steppe migration into Europe during the Bronze Age — was by far the dominant force shaping allele frequency change, dwarfing the contributions of both selection and drift at most loci across the genome.22
Barriers to gene flow
Because gene flow requires the physical movement of organisms or gametes between populations, any factor that impedes this movement constitutes a barrier to gene flow. Barriers may be extrinsic (imposed by the environment) or intrinsic (arising from genetic or behavioural differences between populations), and they range in effectiveness from partial reduction of gene flow to its complete cessation.8
Geographic barriers are the most obvious impediments to gene flow. Mountain ranges, bodies of water, deserts, and other features of the physical landscape can divide a previously continuous population into isolated fragments, preventing or severely reducing the exchange of migrants. Ernst Mayr's classic model of allopatric speciation, elaborated in his 1963 treatise Animal Species and Evolution, held that geographic isolation is the primary initiating event in speciation, because by eliminating gene flow between populations it allows them to diverge independently through the accumulation of genetic differences by drift and selection.8 Even incomplete geographic barriers — a mountain range with passable valleys, or a river that is occasionally crossed — can substantially reduce gene flow and permit measurable genetic differentiation on either side.
Ecological barriers restrict gene flow between populations occupying different habitats, even when they are not geographically separated. If natural selection against immigrants from a different habitat is strong enough, maladapted migrants will fail to survive or reproduce in the recipient population, effectively preventing gene exchange. Nosil, Vines, and Funk demonstrated that immigrant inviability — the reduced fitness of individuals moving between divergent habitats — constitutes a significant form of reproductive isolation that reduces gene flow even in the absence of geographic separation.20
Reproductive barriers encompass any mechanism that prevents or reduces hybridization between divergent populations. Prezygotic barriers, such as differences in mating season, courtship behaviour, or pollinator preference, prevent the formation of hybrid zygotes. Postzygotic barriers, such as hybrid inviability or sterility, reduce the fitness of hybrid offspring. Both types of barriers restrict the effective transfer of alleles between populations, even when individuals from different populations come into physical contact.8, 21 The accumulation of reproductive barriers is central to the process of speciation, and the degree to which these barriers are "leaky" — allowing some gene flow to persist between diverging lineages — has important consequences for the rate and mode of species formation.
Distance itself functions as a barrier to gene flow. In continuously distributed species, the probability that any two individuals exchange genes declines with the geographic distance between them, producing a pattern of isolation by distance in which nearby populations are genetically more similar than distant ones. This pattern, first formalised by Wright, is one of the most commonly observed signatures of spatially restricted gene flow in nature and has been documented across virtually all major taxonomic groups.3, 9
Gene flow and speciation
The relationship between gene flow and speciation is one of the central questions in evolutionary biology. Because gene flow homogenizes populations, it has traditionally been viewed as the primary force opposing the divergence that leads to new species. Mayr's allopatric model explicitly requires the elimination of gene flow by geographic barriers as the first step in speciation, and this model remains the most widely accepted framework for understanding how new species arise.8 However, a growing body of theoretical and empirical evidence demonstrates that speciation can occur in the face of ongoing gene flow under certain conditions, challenging the strict requirement for complete geographic isolation.
In parapatric speciation, divergence occurs between adjacent populations that exchange migrants across a zone of contact. John Endler's theoretical treatment showed that strong divergent selection along an environmental gradient can produce sharp genetic discontinuities — clines — between populations, even when individuals disperse freely across the boundary.7 If selection is strong enough to overcome the homogenizing effect of gene flow, the cline steepens over time, and the accumulation of genetic differences across the contact zone may eventually lead to reproductive isolation and the formation of distinct species. Endler demonstrated mathematically that the width of a cline maintained by the balance between selection and gene flow is proportional to the ratio of dispersal distance to the square root of the selection intensity, providing a quantitative framework for predicting when divergence can proceed despite migration.7
Hybrid zones — narrow geographic regions where genetically distinct populations meet, interbreed, and produce hybrid offspring — provide natural laboratories for studying the tension between gene flow and divergent selection. Barton and Hewitt's landmark review of hybrid zones introduced the concept of the tension zone, a hybrid zone maintained by a balance between dispersal (which broadens the zone by mixing parental genotypes) and selection against hybrids (which narrows it by removing unfit recombinant genotypes).6 Tension zones are expected to be narrow, to coincide with regions of low population density where few individuals are available to move across the boundary, and to persist stably over long periods unless the balance of forces changes.6 The study of hybrid zones has revealed that gene flow across species boundaries is not all-or-nothing: some genomic regions cross freely between species, while others — particularly those containing genes under divergent selection or involved in reproductive isolation — show steep clines and resist introgression, producing a "mosaic" or "semipermeable" genome in which different loci have different effective migration rates.6, 7
Seehausen and colleagues demonstrated that the relationship between gene flow and speciation can operate in reverse: when environmental conditions change in a way that weakens divergent selection or removes ecological barriers to hybridization, previously distinct species may collapse back into a single interbreeding population through a process called speciation reversal or despeciation.19 This phenomenon has been documented in cichlid fish in African lakes, where increased water turbidity — caused by eutrophication from human activity — disrupted the visual cues used by females to distinguish conspecific from heterospecific males, leading to the breakdown of assortative mating and the merger of previously distinct species through hybridization.19
Adaptive introgression

While gene flow between species has traditionally been viewed primarily as an impediment to divergence, a rapidly growing body of research has revealed that interspecific gene flow — introgression — can also be a powerful source of adaptive variation. When alleles that have been shaped by natural selection in one species are transferred to another through hybridization and backcrossing, the recipient species can acquire pre-tested adaptations far more rapidly than if it had to wait for the same mutations to arise and be selected independently.17
One of the most striking examples of adaptive introgression involves the high-altitude adaptation of Tibetan populations. Huerta-Sánchez and colleagues discovered that the EPAS1 gene — which encodes a transcription factor in the hypoxia response pathway and is associated with lower haemoglobin concentrations at high altitude in Tibetans — carries a haplotype that is virtually absent in all other modern human populations but is closely related to sequences found in the genome of a Denisovan, an archaic hominin known primarily from a finger bone found in Siberia.15 The most parsimonious explanation is that the adaptive EPAS1 haplotype was acquired by the ancestors of modern Tibetans through gene flow from Denisovans or a closely related archaic population, and was subsequently driven to high frequency by natural selection in the hypoxic environment of the Tibetan Plateau.15
More broadly, the sequencing of the Neandertal genome revealed that all modern non-African human populations carry approximately 1 to 4 percent Neandertal ancestry, the legacy of gene flow between Neandertals and the ancestors of modern Eurasians approximately 50,000 to 60,000 years ago.14 Subsequent analyses have identified specific Neandertal-derived alleles that appear to have been positively selected in modern human populations, including variants affecting immune function, skin and hair morphology, and metabolism, suggesting that introgression from archaic humans provided a reservoir of adaptive genetic variation that facilitated the colonisation of new environments outside Africa.14
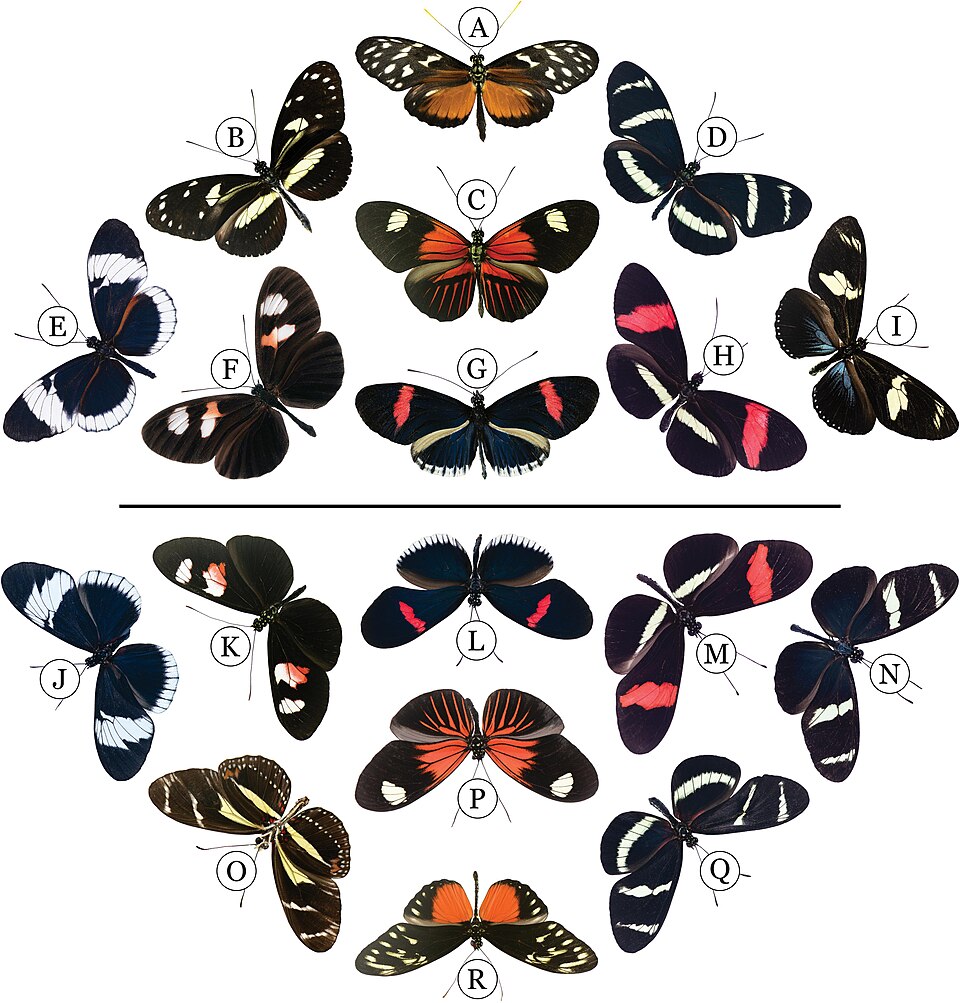
Adaptive introgression is not unique to humans. In Heliconius butterflies, which are renowned for their Müllerian mimicry — a phenomenon in which multiple distasteful species converge on the same warning colour patterns to reinforce predator avoidance — Pardo-Diaz and colleagues demonstrated that mimicry-associated wing-pattern alleles have been transferred between species through introgressive hybridization.16 The red wing-pattern locus in Heliconius timareta was acquired from Heliconius melpomene through gene flow, allowing H. timareta to rapidly adopt the mimicry pattern of its co-mimic without waiting for the independent evolution of the same complex colour-pattern phenotype.16 Hedrick's review of adaptive introgression across animal taxa concluded that introgression can introduce large sets of new alleles simultaneously at multiple unlinked loci, providing a mechanism for rapid adaptation that is qualitatively different from, and potentially faster than, adaptation based on new mutations or standing genetic variation within a species.17
Gene flow in natural populations
The consequences of gene flow are best understood through detailed studies of natural populations in which the movement of individuals, the exchange of genetic material, and the resulting effects on allele frequencies and adaptation can be directly observed or inferred.

Darwin's finches on the Galápagos Islands provide one of the most thoroughly studied examples of the interplay between gene flow, hybridization, and adaptive evolution. Peter and Rosemary Grant's decades-long field study on the small island of Daphne Major documented regular hybridization between Geospiza fortis (the medium ground finch) and G. scandens (the cactus finch), producing hybrid offspring that are viable and fertile.13 This interspecific gene flow introduced alleles from one species into the other, and Grant, Grant, and Petren demonstrated that introgressive hybridization, combined with natural selection, drove convergent morphological evolution between the two species over a period of two decades, with G. scandens evolving a blunter beak shape that approached the morphology of G. fortis as gene flow moved alleles from the larger species into the smaller.23 The finch study illustrates how gene flow can reshape the trajectory of adaptive evolution, producing morphological convergence rather than the divergence that is typically associated with speciation.
Walking-stick insects (Timema cristinae) inhabiting different host plants in California provide a well-documented example of gene flow constraining local adaptation. Populations on the shrub Adenostoma are typically green and striped, while those on Ceanothus are typically unstriped, reflecting selection for crypsis on each host plant. Where the two host plants grow in close proximity, gene flow between adjacent populations on different hosts introduces maladapted colour-pattern alleles, reducing the frequency of the locally optimal morph and producing a measurable migration load.18 Populations receiving higher rates of immigration show greater phenotypic mismatch to their local environment and experience correspondingly stronger directional selection to restore the locally adapted phenotype.18
Human populations provide a uniquely data-rich system for studying gene flow at continental scales. The analysis of ancient DNA from thousands of individuals spanning the past ten thousand years has revealed that the genetic landscape of Europe, Central Asia, and beyond was shaped overwhelmingly by massive population movements — the Neolithic expansion of farming populations from Anatolia, the Bronze Age migration of steppe-related pastoralists into Europe, and subsequent population movements — each of which introduced large quantities of new genetic material and dramatically altered allele frequencies across entire regions.22 At a deeper timescale, gene flow from archaic human populations, including Neandertals and Denisovans, left a lasting imprint on the genomes of all non-African modern humans, contributing functional alleles that have influenced traits ranging from immune defence to skin pigmentation to high-altitude adaptation.14, 15
Plants experience gene flow through mechanisms distinct from those of most animals. Pollen dispersal, mediated by wind, insects, birds, or other vectors, can transfer genetic material over distances far exceeding the dispersal range of seeds, making pollen flow the primary conduit of gene exchange in many plant species. Ellstrand's review documented pollen-mediated gene flow between cultivated crops and their wild relatives occurring at distances of hundreds of metres to several kilometres, raising concerns about the unintended spread of transgenes from genetically engineered crops into wild populations.12 In natural plant populations, long-distance pollen flow has been shown to maintain genetic connectivity among geographically separated populations, counteracting the effects of habitat fragmentation and genetic drift on the scale of entire landscapes.12
Horizontal gene transfer
In prokaryotes, gene flow takes a fundamentally different form from the migration-based transfer that characterises sexual organisms. Bacteria and archaea exchange genetic material through horizontal gene transfer (HGT), also called lateral gene transfer, in which DNA is transmitted between organisms that are not parent and offspring. The three principal mechanisms of HGT are transformation (uptake of free DNA from the environment), transduction (transfer mediated by bacteriophages), and conjugation (direct transfer of DNA between cells through a physical connection called a pilus).11
Ochman, Lawrence, and Groisman's review in Nature established that horizontal gene transfer has been a pervasive force in bacterial evolution, responsible for the acquisition of entirely new metabolic capabilities, virulence factors, and antibiotic resistance determinants.11 Unlike eukaryotes, which evolve primarily through the modification of existing genetic information passed vertically from parent to offspring, bacteria have acquired a substantial proportion of their genetic repertoire from distantly related organisms, producing extremely dynamic genomes in which large segments of DNA are regularly introduced and deleted.11 In some bacterial lineages, horizontally acquired genes constitute 10 to 20 percent or more of the genome, and many of the traits that define the ecological and pathogenic character of particular bacterial species — including the ability to fix nitrogen, degrade novel substrates, or resist antibiotics — were acquired through horizontal transfer rather than vertical inheritance.11
The prevalence of horizontal gene transfer in prokaryotes complicates the classical concept of gene flow as migration between populations of the same species, because HGT routinely crosses not only population boundaries but species and even kingdom boundaries. It also means that the tree-of-life metaphor, in which lineages diverge and evolve independently after separation, is an incomplete description of prokaryotic evolution; a more accurate image is a web or network of genetic connections, with horizontal transfers linking even distantly related branches.11
Modes of gene flow across major groups of organisms1, 11, 12
| Group | Primary mode of gene flow | Typical distance | Key feature |
|---|---|---|---|
| Terrestrial mammals | Individual migration | Metres to hundreds of km | Often sex-biased (male dispersal) |
| Birds | Individual migration | Km to thousands of km | Long-distance natal dispersal |
| Marine invertebrates | Larval dispersal | Metres to hundreds of km | Planktonic larvae carried by currents |
| Wind-pollinated plants | Pollen dispersal | Metres to hundreds of km | Pollen travels far beyond seed range |
| Insect-pollinated plants | Pollen dispersal by vectors | Metres to several km | Depends on pollinator flight range |
| Bacteria | Horizontal gene transfer | Not distance-limited | Crosses species and kingdom boundaries |
Evolutionary significance
Gene flow occupies a unique position among the four mechanisms of evolution. Unlike mutation, which generates new genetic variation, or natural selection and genetic drift, which sort that variation within populations, gene flow redistributes existing variation among populations, binding them together into a genetic network whose structure determines the evolutionary trajectory of the species as a whole.1, 2 Its effects are context-dependent: it can promote adaptation by spreading beneficial alleles, constrain adaptation by swamping locally favoured genotypes, maintain species cohesion by homogenizing gene pools, or facilitate speciation by introducing novel genetic combinations through hybridization.
The recognition that gene flow is rarely an all-or-nothing phenomenon — that genomes are semipermeable to introgression, with different loci experiencing different effective migration rates depending on their proximity to genes under selection — has transformed the study of speciation in the genomic era.6, 7 Speciation with gene flow, once considered theoretically implausible by many evolutionary biologists, is now recognised as a common mode of diversification, and the detection of genomic regions that resist gene flow (so-called "islands of divergence") has become a major focus of speciation genomics.
The study of gene flow also carries practical implications. In conservation biology, understanding the patterns and magnitude of gene flow among fragmented populations is essential for designing wildlife corridors, managing captive breeding programmes, and predicting the genetic consequences of habitat loss.9 In agriculture, the potential for gene flow from genetically modified crops to wild relatives raises questions about the ecological consequences of transgene escape.12 And in public health, horizontal gene transfer among bacteria underlies the rapid spread of antibiotic resistance genes across pathogenic species, one of the most urgent challenges in modern medicine.11 In all of these domains, the principles first articulated by Wright, Dobzhansky, Mayr, and Slatkin continue to provide the conceptual foundation for understanding how the movement of genes shapes the course of evolution.
References
How species evolve collectively: implications of gene flow and selection for the spread of advantageous alleles
Adaptive introgression in animals: examples and comparison to new mutation and standing variation as sources of adaptive variation
Speciation reversal and biodiversity dynamics with hybridization in changing environments
Reproductive isolation caused by natural selection against immigrants from divergent habitats
The contribution of gene flow, selection, and genetic drift to five thousand years of human allele frequency change
Convergent evolution of Darwin's finches caused by introgressive hybridization and selection