Overview
- Hox genes are a family of homeobox-containing transcription factors that specify segment identity along the anterior-posterior axis of nearly all bilaterian animals, and their order on the chromosome mirrors the order of body regions they pattern, a property called colinearity that has been conserved for over 600 million years.
- Mutations in Hox genes produce homeotic transformations — the replacement of one body structure with another, such as legs growing where antennae should be — demonstrating that these genes act as master switches of regional identity rather than builders of individual structures.
- Whole-cluster duplications in the vertebrate lineage produced four paralogous Hox clusters (HoxA–D), providing raw genetic material for the elaboration of the vertebrate body plan, while changes in Hox gene expression boundaries across arthropod lineages account for much of the diversity in segment specialisation and appendage distribution.
Hox genes are a family of transcription factors that contain a highly conserved 180-base-pair DNA sequence called the homeobox, which encodes a 60-amino-acid DNA-binding domain known as the homeodomain.1 These genes function as master regulators of body segment identity along the anterior-posterior (head-to-tail) axis in nearly all bilaterian animals, from insects and nematodes to fish, mice, and humans. The discovery that the same family of regulatory genes controls body plan specification across phyla separated by more than 600 million years of evolution was one of the most transformative findings in modern biology, establishing that animal diversity arises not from fundamentally different genetic programmes but from the differential deployment of a deeply conserved developmental toolkit.3, 4
The study of Hox genes emerged from classical genetic work on the fruit fly Drosophila melanogaster. In the early twentieth century, geneticists identified mutations that caused dramatic homeotic transformations — the conversion of one body part into the likeness of another. The molecular basis of these transformations was elucidated over subsequent decades, culminating in the cloning and characterisation of the Hox gene clusters in the 1980s, work that earned Edward B. Lewis, Christiane Nüsslein-Volhard, and Eric Wieschaus the 1995 Nobel Prize in Physiology or Medicine.5, 7
Homeotic transformations
A homeotic transformation occurs when the identity of one body segment is replaced by that of another. The most famous example in Drosophila involves the Ultrabithorax (Ubx) gene, which specifies the identity of the third thoracic segment. In wild-type flies, the third thoracic segment bears a pair of halteres — small balancing organs derived from the ancestral hindwing. Loss-of-function mutations in Ubx cause the third thoracic segment to adopt the identity of the second thoracic segment, resulting in a four-winged fly with a complete second pair of wings replacing the halteres.5 Conversely, the Antennapedia mutation causes legs to grow in place of antennae on the head, because the gene that normally specifies leg identity in thoracic segments is ectopically expressed in the antennal segment.1, 6
These transformations revealed a critical principle: Hox genes do not build structures directly. They do not encode the proteins that make up legs, wings, or antennae. Instead, they act as selector genes that activate downstream batteries of structural genes appropriate to a particular segment identity. When a Hox gene is misexpressed, the segment in question executes the developmental programme of whichever identity the Hox gene specifies, producing a morphologically normal structure in an abnormal location.1, 14 This distinction between identity specification and structure construction is fundamental to understanding how Hox genes generate body plan diversity.
The Hox cluster and colinearity
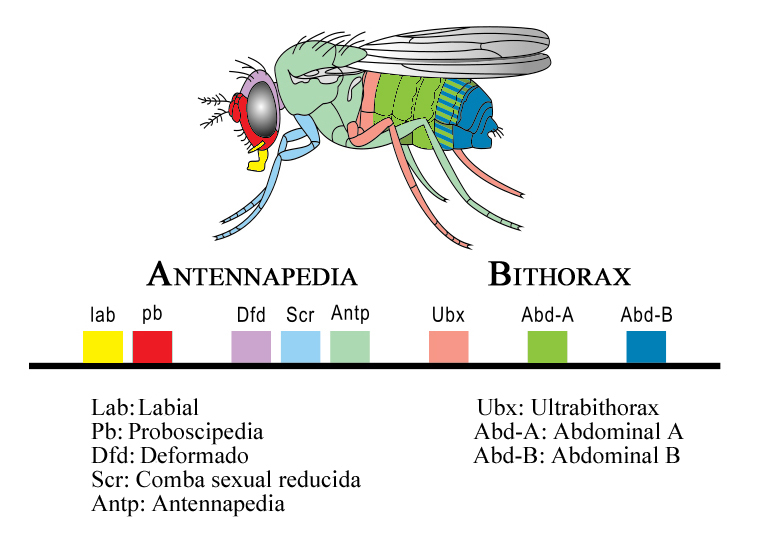
One of the most remarkable features of Hox genes is their chromosomal organisation. In Drosophila, the Hox genes are arranged in two clusters on chromosome 3: the Antennapedia complex (ANT-C), whose genes pattern the head and anterior thorax, and the bithorax complex (BX-C), whose genes pattern the posterior thorax and abdomen.5 In vertebrates, the Hox genes are organised into four paralogous clusters (HoxA, HoxB, HoxC, and HoxD), each located on a different chromosome.2 In all cases, the order of genes along the chromosome corresponds to the order of body regions they pattern along the anterior-posterior axis, a property called spatial colinearity. Genes at the 3' end of the cluster are expressed in anterior body regions and are activated earlier in development, while genes at the 5' end are expressed in posterior regions and are activated later, a property called temporal colinearity.2, 1
This colinear relationship between chromosomal position and expression domain has been conserved across bilaterians for over 600 million years, implying strong functional constraints on cluster organisation. In organisms where the cluster has become fragmented — as in Drosophila, where the ancestral single cluster split into two complexes — colinearity is partially disrupted, and gene regulation relies more heavily on individual enhancer elements. In vertebrates, where the clusters remain intact, the chromatin architecture of the cluster itself plays a regulatory role: the progressive opening of chromatin from 3' to 5' during embryonic development controls the sequential activation of Hox genes in increasingly posterior body regions.2, 13
Deep conservation across phyla
The conservation of Hox genes extends far beyond Drosophila and vertebrates. Hox gene clusters have been identified in virtually all bilaterian phyla, including annelids, molluscs, nematodes, and echinoderms.16 Even cnidarians (jellyfish, corals, and sea anemones), which diverged from bilaterians before the evolution of bilateral symmetry, possess homeobox-containing genes related to the bilaterian Hox family, though their organisation and function differ.4 The ancestral bilaterian is inferred to have possessed a single cluster of at least seven Hox genes, with subsequent lineage-specific duplications and losses producing the diversity of cluster configurations seen today.11
This deep conservation demonstrates that the Hox patterning system was in place before the Cambrian explosion, when the major animal body plans first appeared in the fossil record approximately 540 to 520 million years ago.15 The implication is profound: the genetic toolkit for specifying body segment identity predates the morphological diversity it now generates. The diversity of animal body plans arose not through the invention of new types of patterning genes but through modifications in how existing Hox genes are regulated and in what downstream targets they activate in different lineages.3, 12
Hox cluster duplications in vertebrates
Vertebrates possess four Hox gene clusters, designated HoxA through HoxD, located on four different chromosomes. These clusters arose through two rounds of whole-genome duplication early in vertebrate evolution, an event supported by extensive genomic evidence and often referred to as the 2R hypothesis.11 Following duplication, individual Hox genes within the paralogous clusters underwent a combination of subfunctionalisation (partitioning of ancestral functions among duplicates) and neofunctionalisation (acquisition of novel functions by one duplicate), producing a more complex and versatile Hox code than that of invertebrates.2, 11
The expanded Hox complement of vertebrates has been linked to several features of the vertebrate body plan. The HoxD cluster, for instance, is deployed in two successive phases during limb development: an early phase that patterns the proximal limb (upper arm and forearm) and a later phase, driven by a distinct set of regulatory elements, that patterns the distal autopod (wrist, hand, and digits).4, 17 This second phase of HoxD expression appears to be a vertebrate innovation absent in ray-finned fishes, and it has been proposed as a key regulatory change underlying the evolutionary origin of the tetrapod hand and foot.17 Teleost fishes, which experienced an additional round of genome duplication, possess seven or eight Hox clusters, and the subsequent divergence of duplicated Hox genes has been associated with the morphological diversification of fish body plans.11
Hox genes and arthropod segment diversity
Arthropods display extraordinary variation in the number, type, and specialisation of their body segments. Insects have three pairs of legs restricted to the thorax; crustaceans may have dozens of appendage-bearing segments; myriapods possess numerous leg-bearing segments of similar identity. Evo-devo research has shown that much of this variation can be attributed to differences in the expression boundaries of Hox genes along the anterior-posterior axis.8
In insects, the Hox gene Ultrabithorax and the related gene abdominal-A are expressed in abdominal segments, where they repress the limb-patterning gene Distal-less, preventing leg formation. In crustaceans, however, the expression domains of these same Hox genes differ, and Distal-less is not repressed in the corresponding segments, allowing appendages to develop throughout much of the body.8 Averof and Patel demonstrated that the transition from a crustacean-like body plan with many appendage-bearing segments to an insect-like body plan with appendages restricted to the thorax can be explained largely by shifts in the posterior expression boundary of Ubx and abd-A.8
Butterfly wing patterns provide another illustration. In nymphalid butterflies, Ultrabithorax is expressed in the hindwing, where it specifies hindwing-specific features that distinguish it from the forewing. Weatherbee and colleagues showed that different butterfly species modulate Ubx expression to produce species-specific differences in forewing-hindwing morphology, demonstrating that variation in Hox gene regulation contributes to the diversification of wing patterns within a single insect order.9
Hox protein specificity and cofactors
A longstanding puzzle in Hox biology is how proteins with very similar homeodomains achieve different regulatory outcomes in different segments. The homeodomains of paralogous Hox proteins bind similar DNA sequences in vitro, yet each Hox gene specifies a distinct segmental identity in vivo. The resolution of this paradox lies in the interaction of Hox proteins with cofactors, particularly the TALE-class homeodomain proteins Extradenticle (Exd) and Homothorax (Hth) in Drosophila, and their vertebrate orthologues Pbx and Meis.14 These cofactors bind cooperatively with Hox proteins to form heteromeric complexes that recognise composite DNA binding sites with greater specificity than either protein alone. Different Hox-cofactor combinations recognise different composite sites, enabling each Hox protein to regulate a distinct set of downstream target genes and thereby specify a unique segmental identity.14, 13
Recent research has expanded the view of Hox protein function beyond simple transcriptional activation or repression. Hox proteins have been shown to act as "micromanagers" that regulate not only the broad identity of body segments but also fine-grained aspects of cell behaviour, including cell proliferation, cell adhesion, apoptosis, and cell migration within developing segments.13 This dual role — as both master selectors of segment identity and fine-scale regulators of cell biology — helps explain how Hox genes can generate the detailed morphological differences between segments that go beyond gross anatomical identity.
Hox genes within gene regulatory networks
Hox genes do not operate in isolation. They function as components of hierarchical gene regulatory networks (GRNs) that integrate positional information from upstream signals, process it through combinatorial transcription factor interactions, and activate downstream effector genes that execute the developmental programme.10 Davidson and Erwin proposed that the GRN architecture underlying body plan specification has a layered structure: a deeply conserved "kernel" of interconnected regulatory genes, including Hox factors, that defines fundamental body plan features, underlain by more evolutionarily labile subcircuits and peripheral gene batteries.10
The kernel is highly resistant to evolutionary modification because its internal connectivity means that perturbation of any single component destabilises the entire network. This explains the deep conservation of Hox gene function: the core role of Hox genes in body plan specification has been maintained across bilaterians because the regulatory networks in which they are embedded are so tightly integrated that fundamental changes are developmentally lethal.10, 15 At the same time, the peripheral layers of the network — the downstream targets and tissue-specific enhancers — are free to evolve, and it is changes at this level that produce the morphological differences between species while leaving the core Hox code intact.12
Significance for evolutionary biology
The discovery of Hox genes and their conservation transformed evolutionary biology in several ways. It provided a molecular explanation for homology: structures in different organisms are homologous because they are patterned by the same deeply conserved regulatory genes inherited from a common ancestor.17 It revealed that the genetic potential for complex body plans existed long before the morphological complexity itself appeared in the fossil record, implying that the Cambrian explosion was driven not by the invention of new gene families but by the elaboration of pre-existing regulatory interactions.15 And it showed that major morphological transformations — the suppression of limbs in snake evolution, the modification of arthropod appendages, the origin of the tetrapod hand — can be traced to relatively simple changes in the regulation of a small number of master control genes, rather than requiring wholesale genetic innovation.4, 18
Perhaps most importantly, the Hox gene story demonstrated that the divide between organisms as different as a fruit fly and a human is not one of genetic content but of genetic regulation. The same toolkit builds radically different bodies depending on when, where, and how much each gene is expressed — a principle that lies at the heart of modern evolutionary developmental biology and has reshaped our understanding of how evolution generates the diversity of animal form.3, 12
References
Molecular and genetic organization of the Antennapedia gene complex of Drosophila melanogaster
Hox gene expression in a sea urchin (Strongylocentrotus purpuratus): determination of colinear expression along the oral-aboral axis