Overview
- Mutation is the ultimate source of all genetic variation, generating the raw material upon which natural selection, genetic drift, and other evolutionary forces act — without mutation, evolution could not occur.
- Cells deploy an elaborate suite of DNA repair mechanisms — including base excision repair, nucleotide excision repair, mismatch repair, and double-strand break repair — that correct the vast majority of DNA lesions before they become heritable mutations, reducing replication error rates by several orders of magnitude.
- Spontaneous mutation rates vary enormously across the tree of life but follow predictable patterns: per-nucleotide rates scale inversely with genome size (Drake's rule in microbes) and are constrained by a lower bound set by the power of random genetic drift (the drift-barrier hypothesis), while most molecular-level substitutions are selectively neutral rather than adaptive.
Mutation is the process by which the nucleotide sequence of DNA is altered, producing heritable changes in genetic information that constitute the ultimate source of all genetic variation. Without mutation, there would be no allelic differences for natural selection to act upon, no raw material for genetic drift to sample, and no possibility of adaptive evolution or speciation. Every evolutionary change, from the acquisition of antibiotic resistance in bacteria to the diversification of vertebrate body plans, traces back to one or more mutations that introduced novel genetic variants into a population.6, 15 Yet DNA is not passively subject to degradation. Cells possess an elaborate and highly conserved suite of DNA repair mechanisms that detect and correct the vast majority of lesions before they become permanent mutations, maintaining genome integrity across billions of cell divisions. The interplay between the generation of mutations and the fidelity of DNA repair determines the mutation rate of an organism — a fundamental parameter that shapes the pace and pattern of molecular evolution.4, 5
The study of mutation has been central to genetics since Hermann Muller's demonstration in 1927 that X-rays could induce heritable changes in Drosophila, but the molecular understanding of how mutations arise and how cells repair them has advanced enormously in recent decades. The 2015 Nobel Prize in Chemistry was awarded to Tomas Lindahl, Paul Modrich, and Aziz Sancar for their mechanistic studies of DNA repair, recognising the fundamental importance of these processes to the maintenance of life.22 This article examines the types of mutations, the agents that cause them, the repair systems that counteract them, the rates at which mutations accumulate, and the evolutionary consequences of mutation as a force in its own right.
Types of mutations
Mutations range in scale from single-nucleotide changes to rearrangements affecting entire chromosomes, and their molecular character determines both their likelihood of occurrence and their phenotypic consequences. The simplest and most common class consists of point mutations, in which one nucleotide is substituted for another. Point mutations are divided into two categories based on the chemistry of the substitution: transitions, which exchange a purine for a purine (A to G or G to A) or a pyrimidine for a pyrimidine (C to T or T to C), and transversions, which exchange a purine for a pyrimidine or vice versa. Although there are twice as many possible transversions as transitions, transitions are observed at substantially higher frequencies in most genomes, reflecting the biochemistry of nucleotide misincorporation during replication and the elevated rate of cytosine deamination at CpG dinucleotides.9, 15
Insertions and deletions (collectively termed indels) add or remove one or more nucleotides from the DNA sequence. When an indel occurs within a protein-coding region and the number of nucleotides inserted or deleted is not a multiple of three, the reading frame of the downstream codons is shifted, producing a frameshift mutation that typically results in a truncated or nonfunctional protein. Frameshifts are among the most functionally disruptive mutations because they alter every codon downstream of the indel, usually generating a premature stop codon.6, 21 Indels are frequently caused by slippage of the DNA polymerase during replication of repetitive sequences, where short tandem repeats provide opportunities for the template and nascent strands to misalign.
At a larger scale, chromosomal mutations rearrange the order or orientation of substantial segments of DNA. Inversions reverse the orientation of a chromosomal segment without changing its content, potentially disrupting genes at the breakpoints or altering gene expression by repositioning regulatory elements. Translocations move a segment from one chromosome to another, sometimes fusing parts of two genes to create a novel chimeric gene. Duplications copy a segment, ranging from a single gene to an entire chromosome arm, providing redundant genetic material that can diverge in function over evolutionary time. Susumu Ohno argued in his influential 1970 monograph that gene duplication has been the single most important factor in evolution, because a duplicated gene is freed from the constraint of maintaining its original function and can accumulate mutations that lead to novel capabilities.17 Polyploidy — the duplication of the entire genome — is rare in animals but widespread in plants, where it has been a major driver of speciation and adaptation. In vertebrate evolution, two rounds of whole-genome duplication early in the chordate lineage are thought to have provided the genetic raw material for the elaboration of the vertebrate body plan.17
Sources of spontaneous DNA damage
DNA, despite its role as the stable repository of genetic information, is chemically fragile. Tomas Lindahl's landmark 1993 review demonstrated that DNA undergoes continuous spontaneous decay through hydrolysis, oxidation, and nonenzymatic methylation at rates sufficient to generate thousands of lesions per cell per day in the absence of any external damaging agent.
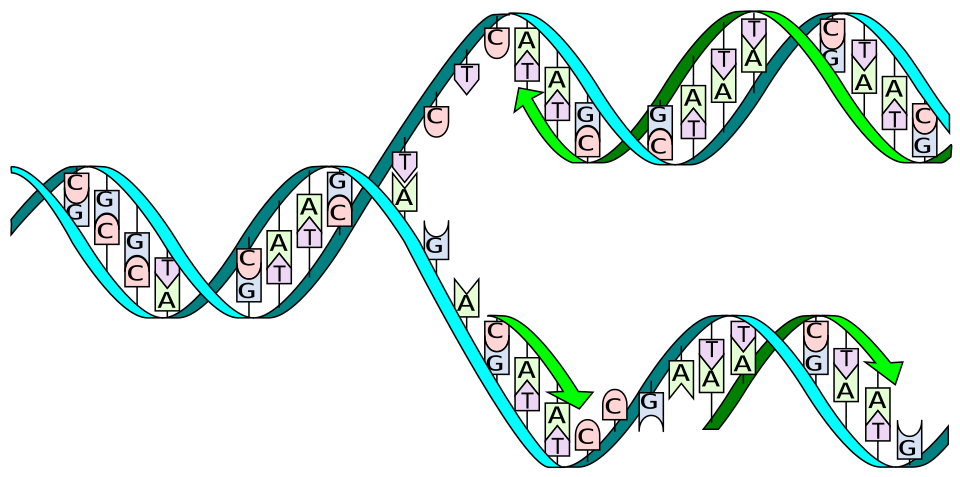
The two most common hydrolytic reactions are depurination, in which the glycosidic bond between a purine base (adenine or guanine) and its deoxyribose sugar is cleaved to produce an abasic (apurinic/apyrimidinic) site, and deamination, in which cytosine loses its amino group and is converted to uracil. In mammalian cells under physiological conditions, an estimated 10,000 to 20,000 depurination events and 100 to 500 cytosine deamination events occur per cell per day.4 If left unrepaired, an abasic site can block replication or cause the insertion of an incorrect nucleotide, while cytosine deamination to uracil produces a G:U mismatch that, if replicated, becomes a C-to-T transition mutation.
Reactive oxygen species (ROS), generated as by-products of aerobic metabolism in the mitochondrial electron transport chain, constitute another major source of endogenous DNA damage. The most prevalent oxidative lesion is 8-oxoguanine (8-oxoG), a modified form of guanine that can mispair with adenine during replication, leading to G-to-T transversion mutations. It is estimated that several hundred to several thousand 8-oxoG lesions arise per cell per day in human tissues, depending on the metabolic activity of the tissue.4, 21 The cumulative burden of oxidative DNA damage has been implicated in ageing, cancer, and neurodegenerative disease.
Errors introduced during DNA replication represent a third major source of mutation. Replicative DNA polymerases insert incorrect nucleotides at a frequency of approximately one per 104 to 105 nucleotides polymerised. The intrinsic 3'-to-5' exonuclease (proofreading) activity of these polymerases then removes most of these misincorporations, improving fidelity by roughly 100-fold to an error rate of approximately one per 107 nucleotides.6 Post-replicative mismatch repair further reduces the error rate by another two to three orders of magnitude, yielding a final per-nucleotide error rate of approximately 10−9 to 10−10 per base pair per cell division in organisms with functional repair systems.5, 6
Exogenous mutagenic agents
In addition to the spontaneous lesions generated by normal cellular chemistry, DNA is vulnerable to damage from a range of environmental agents. Ultraviolet (UV) radiation, particularly the UV-B component of sunlight (wavelengths 280 to 315 nanometres), is absorbed directly by DNA bases and induces the formation of cyclobutane pyrimidine dimers (CPDs) and 6-4 photoproducts between adjacent pyrimidines on the same strand. These bulky photoproducts distort the double helix, blocking transcription and replication and causing C-to-T transitions at dipyrimidine sites if they are bypassed by error-prone translesion synthesis polymerases.5, 21 The characteristic UV mutation signature — C-to-T transitions at CC and TC dinucleotides — is the dominant mutational signature in sun-exposed human skin and in skin cancers such as melanoma and squamous cell carcinoma.19
Ionising radiation (X-rays, gamma rays, and cosmic rays) damages DNA primarily through the generation of free radicals in water molecules surrounding the DNA, producing a spectrum of lesions that includes single-strand breaks, double-strand breaks, base modifications, and DNA-protein crosslinks. Double-strand breaks are the most dangerous of these lesions because both complementary strands are severed, leaving no intact template for error-free repair.7, 21
Chemical mutagens encompass a diverse array of compounds that damage DNA through distinct mechanisms. Alkylating agents such as ethyl methanesulfonate (EMS) and N-methyl-N-nitrosourea (MNU) add methyl or ethyl groups to DNA bases, producing miscoding lesions such as O6-methylguanine, which pairs with thymine instead of cytosine. Intercalating agents such as ethidium bromide and acridine orange insert themselves between adjacent base pairs, distorting the helix and causing insertions or deletions during replication. Certain industrial and dietary chemicals, including polycyclic aromatic hydrocarbons and aflatoxin B1, are metabolised into reactive intermediates that form bulky covalent adducts with DNA bases, particularly guanine.21 Each class of mutagen produces a characteristic spectrum of mutations, and the identification of these mutational signatures in cancer genomes has become a powerful tool for tracing the environmental exposures that drive tumour development.19
DNA repair mechanisms
The high rate of spontaneous and environmentally induced DNA damage would rapidly overwhelm genome integrity if cells lacked the means to detect and correct it.
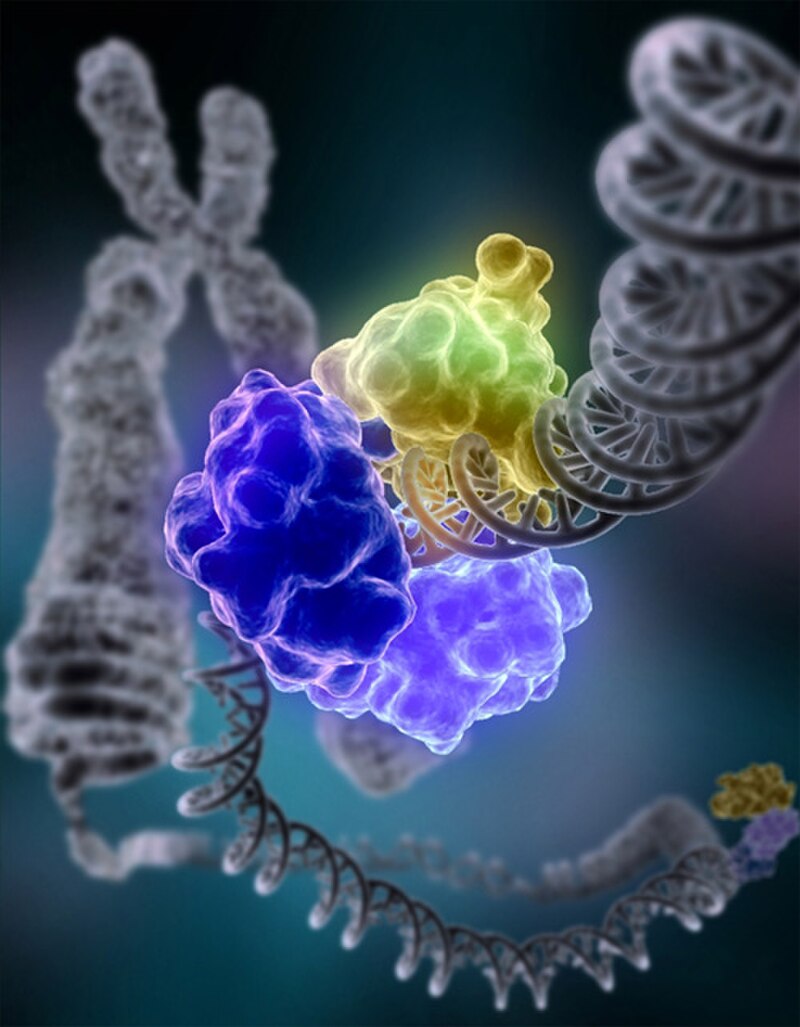
Over the course of evolution, all living organisms have evolved a network of repair pathways, each specialised for a particular class of lesion. The importance of these pathways was recognised with the 2015 Nobel Prize in Chemistry, awarded for the elucidation of three of the principal repair mechanisms: base excision repair, nucleotide excision repair, and mismatch repair.22
Base excision repair (BER) corrects small, non-helix-distorting lesions such as oxidised bases (including 8-oxoguanine), deaminated bases (uracil arising from cytosine deamination), and alkylated bases. The pathway is initiated by a DNA glycosylase that recognises the damaged base and cleaves the glycosidic bond linking it to the sugar-phosphate backbone, leaving an abasic site. An AP endonuclease then incises the backbone adjacent to the abasic site, and DNA polymerase and ligase fill the resulting gap. The human genome encodes at least eleven different DNA glycosylases, each recognising a distinct set of base lesions, providing broad coverage against the diverse products of oxidative and hydrolytic damage.4, 5, 22
Nucleotide excision repair (NER) removes bulky, helix-distorting lesions such as UV-induced pyrimidine dimers and chemical adducts. Unlike BER, which excises only the damaged base, NER removes a short oligonucleotide segment — typically 12 to 13 nucleotides in prokaryotes and 24 to 32 nucleotides in eukaryotes — spanning the lesion. The resulting gap is filled by a DNA polymerase using the undamaged strand as a template and sealed by DNA ligase. In humans, inherited defects in NER genes cause xeroderma pigmentosum, a devastating syndrome characterised by extreme sensitivity to sunlight and a more than 1,000-fold increase in the risk of UV-induced skin cancers.5, 22
Mismatch repair (MMR) corrects errors that escape the proofreading activity of the replicative DNA polymerase, including base-base mismatches and small insertion-deletion loops. The pathway employs specialised proteins (MutS and MutL homologues in eukaryotes) that scan newly replicated DNA for mismatches, distinguish the newly synthesised strand from the parental template, excise the erroneous segment, and resynthesize the correct sequence. MMR improves replication fidelity by approximately 100- to 1,000-fold.5, 6 Inherited defects in human MMR genes cause Lynch syndrome (hereditary nonpolyposis colorectal cancer), which confers a substantially elevated lifetime risk of colorectal and other cancers due to the accumulation of replication errors throughout the genome.5
Double-strand break repair addresses the most dangerous class of DNA damage through two principal pathways. Non-homologous end joining (NHEJ) directly ligates the broken ends with minimal processing, operating throughout the cell cycle. NHEJ is fast but inherently error-prone, frequently introducing small insertions or deletions at the junction site.7 Homologous recombination (HR) uses an intact homologous sequence (typically the sister chromatid in S or G2 phase) as a template to restore the broken region with high fidelity. HR involves end resection to generate single-stranded DNA, strand invasion of the homologous template mediated by the Rad51 recombinase, DNA synthesis using the template, and resolution of the recombination intermediates.8 The choice between NHEJ and HR depends on the cell cycle stage and the nature of the break, with HR predominating when a sister chromatid template is available.7, 8
Spontaneous mutation rates across organisms
The spontaneous mutation rate — the probability that a given nucleotide will be altered per replication or per generation — varies across many orders of magnitude among different organisms, reflecting differences in genome size, generation time, DNA polymerase fidelity, and the efficiency of repair systems. One of the most striking regularities in this variation was identified by John Drake in 1991, who showed that for DNA-based microbes (bacteriophages, bacteria, and simple eukaryotes), the per-nucleotide mutation rate scales inversely with genome size such that the per-genome mutation rate per replication is approximately constant at roughly 0.003 mutations per genome per generation.1 This observation, now known as Drake's rule, implies that microbes with larger genomes have evolved proportionally more accurate replication machinery, maintaining a nearly invariant total genomic mutation rate despite orders-of-magnitude differences in genome size.
In multicellular eukaryotes, mutation rates per nucleotide site per generation are generally higher than in microbes, in part because these organisms undergo many more cell divisions per generation (with more opportunities for replication errors) and in part because their smaller effective population sizes weaken the efficiency of selection for replication fidelity.14, 20 In humans, early indirect estimates based on comparisons of pseudogene sequences between humans and chimpanzees placed the per-nucleotide mutation rate at approximately 2.5 × 10−8 per site per generation, corresponding to roughly 175 mutations per diploid genome per generation.9 Direct whole-genome sequencing of Icelandic parent-offspring trios subsequently confirmed a rate of approximately 1.2 × 10−8 per nucleotide per generation, with a mean of about 70 de novo single-nucleotide mutations per offspring. This study also demonstrated that the number of de novo mutations increases with paternal age at a rate of roughly two additional mutations per year, reflecting the continued division of spermatogonial stem cells throughout male adulthood.10
Spontaneous mutation rates across the tree of life1, 9, 10, 14, 16
| Organism | Genome size (Mb) | Per-nucleotide rate (per generation) | Per-genome rate (per generation) |
|---|---|---|---|
| Bacteriophage λ | 0.049 | ~7.7 × 10−8 | ~0.004 |
| Escherichia coli | 4.6 | ~5.4 × 10−10 | ~0.003 |
| Saccharomyces cerevisiae | 12.1 | ~3.3 × 10−10 | ~0.004 |
| Caenorhabditis elegans | 100 | ~2.7 × 10−9 | ~0.27 |
| Drosophila melanogaster | 140 | ~3.5 × 10−9 | ~0.49 |
| Mus musculus (mouse) | 2,700 | ~5.4 × 10−9 | ~14.6 |
| Homo sapiens | 3,200 | ~1.2 × 10−8 | ~38–70 |
The table illustrates the enormous range of per-nucleotide rates across organisms and the approximate constancy of the per-genome rate in microbes (Drake's rule), contrasted with the much higher per-genome rates in multicellular eukaryotes. The higher per-genome rate in organisms like humans reflects both the larger genome and the greater number of cell divisions per generation, particularly in the male germline.10, 15
Mutation rate and genome size
The observation that mutation rates are not fixed but have themselves evolved under natural selection raises a fundamental question: what determines the mutation rate of a species? Natural selection generally favours lower mutation rates because most non-neutral mutations are deleterious — estimates from a range of organisms suggest that the distribution of fitness effects of new mutations is strongly skewed toward harmful effects, with beneficial mutations constituting a small minority.18 Selection should therefore drive the evolution of ever more accurate replication and repair machinery. Yet mutation rates cannot be reduced to zero, because the molecular machinery of replication fidelity and DNA repair has finite costs in terms of energy, time, and the enzymes required.
Michael Lynch and colleagues formalised this reasoning in the drift-barrier hypothesis, which proposes that the lower bound on the mutation rate is set not by molecular constraints but by the power of random genetic drift. In populations with small effective population sizes, the selective advantage of a further incremental improvement in replication fidelity may be so small that it is overwhelmed by drift, preventing selection from driving the mutation rate any lower.13, 14 This framework explains a key empirical pattern: organisms with large effective population sizes, such as bacteria with populations of 108 or more, tend to have much lower per-nucleotide mutation rates than organisms with small effective population sizes, such as vertebrates with effective populations on the order of 104 to 105.14, 20 The drift barrier thus provides a unifying explanation for the inverse correlation between mutation rate and effective population size observed across the tree of life.
The drift-barrier hypothesis also accounts for the relationship between mutation rate and genome size. Organisms with large genomes tend to have small effective population sizes (partly because of the increased energetic cost of maintaining a large genome and partly because of ecological constraints on large-bodied organisms), and therefore have higher per-nucleotide mutation rates. Conversely, organisms with small genomes tend to have large effective population sizes and lower per-nucleotide rates. The net result is that the total mutational burden per genome per generation increases with genome size, but the per-nucleotide rate decreases, as captured by Drake's rule for microbes.1, 14, 20
The neutral theory of molecular evolution
The recognition that mutation rates are high enough to generate enormous numbers of new variants in every generation prompted a fundamental re-evaluation of the role of mutation in evolution. In 1968, Motoo Kimura proposed the neutral theory of molecular evolution, arguing that the great majority of evolutionary substitutions at the molecular level are not driven by natural selection but instead result from the random fixation of selectively neutral (or very nearly neutral) mutations through genetic drift.2 Kimura's argument was based on the observation that the rate of amino acid substitution in proteins, as inferred from comparative sequence data, appeared to be remarkably constant per year across diverse lineages — a pattern more consistent with a steady rain of neutral mutations fixed by drift than with the episodic action of positive selection.
The neutral theory does not deny the importance of natural selection in shaping organismal phenotypes, nor does it claim that all mutations are neutral. Rather, it asserts that among the mutations that actually become fixed in populations over evolutionary time, the vast majority are neutral with respect to fitness, while strongly deleterious mutations are eliminated by purifying selection and strongly beneficial mutations, though important, are sufficiently rare that they account for only a small fraction of total substitutions.2, 3 The theory predicts that the rate of neutral molecular evolution should be equal to the neutral mutation rate, independent of population size — a prediction that has been broadly confirmed by comparative genomic data showing approximately clock-like substitution rates in non-functional sequences such as pseudogenes and intergenic regions.
Kimura expanded his framework in his 1983 book The Neutral Theory of Molecular Evolution, incorporating the concept of nearly neutral mutations whose selective effects are so small that their fate is determined primarily by drift in finite populations.3 The nearly neutral theory predicts that organisms with smaller effective population sizes should fix a greater proportion of slightly deleterious mutations, leading to a higher rate of molecular evolution in functional sequences — a prediction supported by evidence from island populations and organisms with small population sizes. The neutral theory remains one of the foundational frameworks of molecular evolution, serving as the null hypothesis against which evidence for positive selection is evaluated.2, 3, 18
Somatic versus germline mutations
An essential distinction in the biology of mutation is between germline mutations, which occur in the cells that give rise to gametes and are therefore heritable, and somatic mutations, which occur in the non-reproductive cells of the body and affect only the individual in which they arise. From the standpoint of evolution, only germline mutations matter: they are the source of the heritable genetic variation upon which natural selection and drift operate. Somatic mutations, by contrast, cannot be transmitted to offspring and are therefore invisible to evolutionary forces acting across generations.15, 19
In the human male germline, spermatogonial stem cells divide continuously from puberty onward, accumulating roughly two replication-associated mutations per year. This continuous division explains the well-documented paternal age effect: children of older fathers carry more de novo mutations than children of younger fathers, with the average number increasing by approximately two mutations per additional year of paternal age.10, 15 In the female germline, by contrast, oocytes complete the majority of their DNA replication during fetal development and then arrest until ovulation, resulting in a much smaller contribution of maternal age to de novo mutation counts. The sex difference in mutation rate has evolutionary implications: the male germline is the predominant source of new point mutations in the population, a pattern sometimes summarised as the male mutation bias.15
Although somatic mutations do not contribute to heritable evolution, they are of immense biomedical significance. Somatic mutations accumulate throughout the human lifespan at rates that vary by tissue type, with rapidly dividing tissues such as the intestinal epithelium, blood-forming cells, and skin acquiring mutations at particularly high rates. Deep sequencing of normal human tissues has revealed that phenotypically normal cells can carry surprisingly high burdens of somatic mutations, including driver mutations in known cancer genes, which accumulate with age and are subject to positive selection within tissues.19 This process of somatic evolution — mutation, selection, and clonal expansion within the body — mirrors the evolutionary dynamics of populations and is a central mechanism of cancer development.
Mutation accumulation experiments
Much of what is known about spontaneous mutation rates and spectra comes from mutation accumulation (MA) experiments, in which organisms are propagated under conditions that minimise the efficiency of natural selection, allowing mutations to accumulate at rates close to the underlying mutation rate. By maintaining very small population sizes (often single individuals or pairs passed through repeated bottlenecks), researchers ensure that only lethal or nearly lethal mutations are removed by selection, while the great majority of spontaneous mutations — including mildly deleterious ones — drift to fixation and can be detected by subsequent sequencing.16

One of the most important MA experiments in multicellular organisms was conducted by Denver and colleagues using the nematode Caenorhabditis elegans. By propagating MA lines through single-individual bottlenecks for hundreds of generations and then sequencing their genomes, the researchers obtained a direct and unbiased estimate of the nuclear mutation rate: approximately 2.7 × 10−9 per nucleotide site per generation, with a mutational spectrum dominated by insertions and featuring a pronounced bias toward transitions over transversions. This rate was roughly tenfold higher than previous indirect estimates, demonstrating the value of direct genomic approaches for measuring mutation rates.16
The most celebrated long-running evolution experiment is Richard Lenski's Long-Term Evolution Experiment (LTEE) with Escherichia coli, initiated in 1988 and now exceeding 80,000 generations. Twelve initially identical populations have been propagated in a simple glucose-limited medium, and their evolutionary trajectories have been tracked through frozen fossil records and whole-genome sequencing.23 The LTEE has yielded several landmark findings directly relevant to mutation biology. Six of the twelve populations independently evolved mutator phenotypes — elevated mutation rates caused by defects in DNA repair genes, particularly in the mismatch repair system. In most cases, the mutator alleles appear to have risen to fixation not because high mutation rates are intrinsically beneficial, but because they were hitchhiked to fixation by linkage with strongly beneficial mutations that arose in the mutator genetic background.11
The LTEE also produced one of the most dramatic examples of evolutionary innovation through mutation. After more than 30,000 generations, one of the twelve populations evolved the ability to utilise citrate as a carbon source under aerobic conditions — a capacity that E. coli normally lacks and that defines the species. Replay experiments using the frozen fossil record demonstrated that this innovation required the prior occurrence of specific potentiating mutations that created the genetic background necessary for the key citrate-utilising mutation to have its phenotypic effect, providing direct evidence for historical contingency in the mutational pathways available to evolving populations.12
The distribution of fitness effects
Not all mutations are equal in their consequences. The distribution of fitness effects (DFE) of new mutations describes the probability that a random new mutation will have a given effect on the fitness of the organism that carries it. Extensive empirical work across bacteria, yeast, Drosophila, nematodes, and other model organisms has established that the DFE is strongly skewed: the majority of new mutations that have any effect on fitness are deleterious, while beneficial mutations are rare.18 A substantial fraction of mutations, particularly those in non-coding DNA and at synonymous sites in protein-coding genes, appear to be effectively neutral, consistent with the predictions of Kimura's neutral theory.
Studies using site-directed mutagenesis and deep mutational scanning have refined this picture. For protein-coding genes, most amino acid substitutions are at least mildly deleterious, reducing protein stability, catalytic efficiency, or folding kinetics. A smaller fraction are strongly deleterious, eliminating protein function altogether. Beneficial mutations exist but typically represent only a small percentage of the total spectrum, and their effects tend to be modest compared with the effects of the most deleterious mutations.18 The shape of the DFE has profound evolutionary implications: it determines the rate at which populations can adapt (limited by the supply of beneficial mutations), the genetic load imposed by the accumulation of deleterious mutations, and the fraction of molecular substitutions that are driven by positive selection versus genetic drift.
The DFE also depends on the environment and the current state of adaptation of the organism. Populations far from their adaptive optimum tend to encounter beneficial mutations more frequently and with larger effect sizes, because there are many possible improvements available. As populations approach the optimum, beneficial mutations become rarer and smaller in effect, a pattern consistent with Fisher's geometric model of adaptation.18 This diminishing-returns epistasis has been observed directly in the Lenski LTEE, where the rate of fitness improvement has decelerated over tens of thousands of generations as the populations have become progressively better adapted to their laboratory environment.23
Distribution of fitness effects of new mutations18
The approximate proportions shown above reflect estimates from a range of organisms and loci; exact values vary by species, gene, and environmental context. The key pattern is consistent across studies: the vast majority of non-neutral mutations are deleterious, neutral and nearly neutral mutations are abundant, and beneficial mutations are rare but disproportionately important for adaptive evolution.18
References
The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway
Historical contingency and the evolution of a key innovation in an experimental population of Escherichia coli
High mutation rate and predominance of insertions in the Caenorhabditis elegans nuclear genome