Overview
- All living organisms fall into a nested hierarchical classification—groups within groups within groups—exactly the pattern predicted by descent with modification from a common ancestor through a branching evolutionary tree.
- DNA sequences, protein sequences, morphological traits, and biogeographic distributions independently produce the same nested hierarchy, a concordance that would be astronomically improbable if species were designed independently.
- Where conflicts between data sets exist, they are small and explicable by known biological processes such as horizontal gene transfer and convergent evolution—they do not disrupt the overall hierarchical pattern.
When biologists classify living organisms, a striking pattern emerges: all life falls into a nested hierarchy of groups within groups within groups. Mammals are a subset of vertebrates, which are a subset of chordates, which are a subset of animals. Within mammals, primates are a subset; within primates, great apes; within great apes, the genus Homo. This is not an arbitrary filing system imposed by human convention—it reflects a real, discoverable structure in the natural world. The same hierarchical pattern is recovered independently by anatomical comparisons, DNA sequences, protein structures, embryological development, and biogeographic distributions.1, 2 This concordance across independent lines of evidence is one of the most fundamental predictions of evolutionary theory and one of the strongest confirmations of universal common descent.
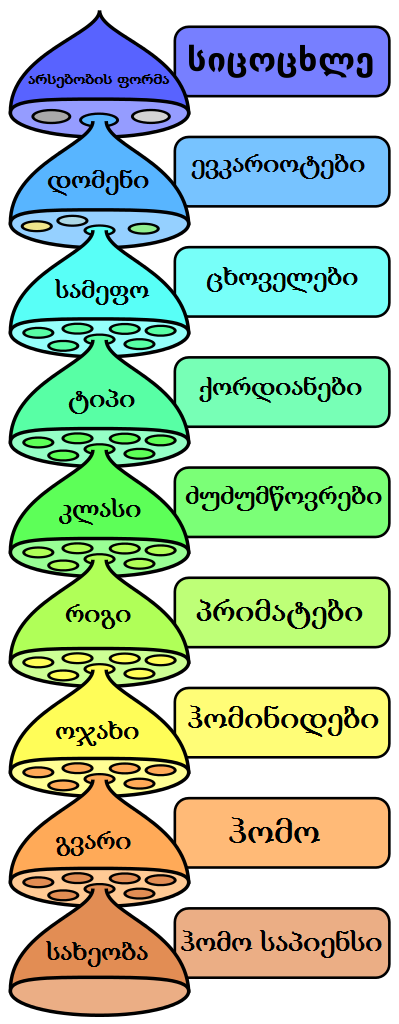
The prediction of descent with modification
Darwin recognized that the hierarchical pattern of life is a natural consequence of descent with modification through a branching tree.1 When a population splits into two lineages that evolve independently, the resulting species share all the characteristics they inherited from their common ancestor while differing in whatever traits changed after the split. When those descendant lineages themselves split, the pattern repeats at a finer scale. The result, accumulated over billions of years, is a tree-like structure in which organisms at any level of analysis can be grouped by shared derived characters into nested sets—precisely the pattern that Linnaeus described in the eighteenth century, decades before Darwin proposed a mechanism to explain it.2, 5
This prediction is specific. Evolution by descent from a common ancestor requires that all organisms fit into a single, consistent, branching hierarchy. If organisms were instead designed independently, with features mixed and matched according to the requirements of each species, there is no reason the resulting pattern should be hierarchical at all. A designer could give wings to mammals, photosynthesis to animals, or the echolocation system of bats to birds. The features of different organisms could be combined in any arrangement, producing a reticulate pattern rather than a nested one.3, 12
Concordance across independent data sets
The most compelling aspect of the nested hierarchy of life is not that one data set produces it, but that every independent data set produces the same hierarchy. Morphological comparisons among vertebrates, for example, group mammals together based on features such as hair, mammary glands, and three middle ear bones. Molecular comparisons of cytochrome c protein sequences group the same organisms together based on amino acid similarities. Ribosomal RNA sequences produce the same groupings. Immunological distances, chromosome banding patterns, and the distribution of endogenous retroviruses all independently recover the same tree.6, 7, 10
This concordance is the key observation. Any single data set might conceivably produce a hierarchical pattern for reasons unrelated to common ancestry. But when dozens of independent data sources—each subject to different evolutionary pressures, different rates of change, and different methodological biases—converge on the same branching tree, the probability that the pattern is an artifact becomes negligibly small. Theobald formally tested the hypothesis of universal common ancestry against alternative models of independent origin and found that common ancestry was supported over independent origin by a factor of at least 102,860—a number so large as to represent statistical certainty.9
Molecular phylogenetics and the hierarchy
The revolution in molecular biology that began in the 1960s provided a powerful new way to test the hierarchical pattern. If organisms truly share a nested genealogy, then comparisons of homologous DNA and protein sequences should produce branching trees that match those derived from anatomy, and the degree of sequence divergence should correlate with the time since two lineages separated. This prediction has been confirmed across every group of organisms examined.6, 7, 10
When phylogenomic studies compare hundreds or thousands of genes simultaneously, the resulting trees achieve high statistical support and agree with morphology-based classifications at virtually every major node. Mammals group with other vertebrates, arthropods group with other ecdysozoans, and flowering plants group with other seed plants—exactly as predicted by anatomy, fossils, and biogeography.10, 11 The few remaining areas of genuine uncertainty (such as the precise branching order among the earliest animal phyla or certain rapid radiations within mammals) involve cases where lineages diverged in rapid succession, leaving little time for molecular differences to accumulate between splitting events. These are technical challenges of resolution, not challenges to the hierarchical pattern itself.10, 11
Apparent conflicts and their resolution
Opponents of evolution sometimes point to cases where different data sets produce conflicting trees as evidence against the hierarchical pattern. Such conflicts do exist, but they are small, localized, and explicable by known biological mechanisms—they do not disrupt the overall nested structure.9, 8
Horizontal gene transfer (HGT) is the most significant source of genuine conflict. In bacteria and archaea, genes are frequently transferred between distantly related lineages, blurring the tree-like pattern. Even in eukaryotes, HGT occurs through mechanisms such as endosymbiosis, viral integration, and parasitic gene capture.9, 8 However, HGT produces a characteristic signature: a small minority of genes in an organism may have a phylogenetic history that conflicts with the majority. The underlying hierarchical pattern is not destroyed but rather overlaid with a secondary reticulate signal that can be identified and accounted for through careful analysis.8
Convergent evolution—the independent evolution of similar traits in unrelated lineages—is another source of superficial conflict. The wings of bats, birds, and pterosaurs are structurally similar but evolved independently. If one classified organisms by the presence of wings alone, one would incorrectly group these animals together. But convergent traits are always a minority of total characters, and they are readily identified by their lack of detailed similarity at deeper anatomical and molecular levels. Bat wings are modified mammalian forelimbs with five fingers; bird wings are modified dinosaurian forelimbs with fused digits; the underlying skeletal architectures are entirely different.3, 7
The mathematical improbability of concordance under design
The strength of the nested hierarchy argument can be appreciated by considering the combinatorial mathematics. For any set of species, there are an astronomically large number of possible branching trees. For just 30 species, the number of possible unrooted bifurcating trees exceeds 1036.2 If species were designed independently with features assigned without regard to phylogenetic history, the probability that DNA sequences, protein sequences, morphological characters, and shared pseudogenes would all converge on the same tree out of these trillions upon trillions of possibilities is effectively zero.
Under common descent, however, concordance is not just expected—it is required. Every heritable trait in an organism was transmitted through the same lineage of ancestors, so every independent data set must recover the same branching history (subject to the minor perturbations from HGT and convergence noted above). The fact that this prediction is confirmed across all of life, from bacteria to whales, from DNA sequences to skeletal anatomy, represents one of the most thoroughly tested and strongly supported conclusions in all of science.7, 9, 12
From Linnaeus to phylogenetics
It is worth noting that the nested hierarchical pattern of life was recognized long before Darwin. Carl Linnaeus, working in the eighteenth century within a creationist framework, organized all known organisms into a hierarchy of kingdom, class, order, genus, and species. He did so because the pattern was empirically obvious—organisms naturally cluster into nested groups based on shared characteristics. What Linnaeus could not explain was why this pattern existed. Why should nature be organized as a tree rather than a web, a continuum, or a random scatter?2, 5
Darwin's theory of descent with modification provided the answer: the nested hierarchy exists because it is the genealogical record of life's actual history. Modern phylogenetics and cladistic methods have refined Linnaeus's classification into an explicit reconstruction of evolutionary relationships, but the fundamental observation remains the same. Life is organized into a tree because life evolved as a tree, with each branching point representing the splitting of one ancestral population into two independently evolving lineages.2, 7
Shared derived characters and nested sets
The nested hierarchy is not built from just any similarities between organisms but specifically from shared derived characters (synapomorphies)—traits that originated in a common ancestor and were inherited by all of its descendants. Hair and mammary glands are synapomorphies that define the group Mammalia; feathers are a synapomorphy of Dinosauria (including birds); a bony internal skeleton is a synapomorphy of Vertebrata. The distinction between shared derived characters and shared ancestral characters (symplesiomorphies) is critical: possessing a vertebral column is shared among all mammals, but it cannot be used to define the mammalian group because it was inherited from a much earlier ancestor and is shared with fish, amphibians, and reptiles as well.2, 13
This distinction is what gives the nested hierarchy its diagnostic power. If organisms were assembled independently rather than descending from common ancestors, the distribution of characters across organisms would not form nested sets. Instead, characters would be distributed in overlapping, conflicting patterns that could not be resolved into a single branching tree. The fact that characters do form nested sets—that every group defined by one set of characters is fully contained within a larger group defined by more ancient characters—is precisely the pattern produced by a branching genealogy and no other known process.5, 13
The three-domain classification of life proposed by Carl Woese illustrates nested hierarchy at the deepest level. Within Eukarya, animals, plants, and fungi each form nested groups defined by their own suites of synapomorphies. Within animals, chordates form a nested group; within chordates, vertebrates; within vertebrates, tetrapods; and so on down to individual species. At every level of this hierarchy, the groupings recovered from molecular data match those recovered from morphological and developmental data, confirming that the pattern reflects a single underlying genealogical history.7, 14 Rapid radiations—periods in which many lineages diverged in quick succession—can make certain nodes difficult to resolve, producing "bushes" rather than cleanly bifurcating branches, but these represent limitations of resolution at specific timescales rather than failures of the hierarchical pattern itself.15
References
Resolution of a concatenation/coalescence kerfuffle: partitioned coalescence support and a robust family-level tree for Mammalia