Overview
- Cospeciation (parallel cladogenesis) occurs when a host lineage and its associated parasite or symbiont lineage speciate in tandem, producing mirror-image phylogenetic trees that provide some of the strongest evidence for long-term coevolutionary associations.
- Phylogenetic congruence between host and associate trees is assessed using methods such as reconciliation analysis, ParaFit, and cophylogenetic mapping, which distinguish cospeciation events from host switches, duplications, losses, and failure to speciate.
- Classic examples include pocket gophers and their chewing lice, figs and fig wasps, and great apes and their lentiviruses, though strict cospeciation is rare in nature and most host-associate systems show a mixture of cospeciation and host-switching events.
Cospeciation, also called parallel cladogenesis, is the process by which two ecologically associated lineages — typically a host and its parasite, symbiont, or obligate mutualist — speciate in synchrony, so that each speciation event in the host lineage is mirrored by a corresponding speciation event in the associate lineage.11, 12 When cospeciation occurs consistently over evolutionary time, it produces a pattern of phylogenetic congruence in which the branching order (topology) and relative timing of divergences in the host phylogeny closely match those in the associate phylogeny. This mirror-image relationship, sometimes called Fahrenholz's rule after the parasitologist who first articulated the expectation, provides powerful evidence that the two lineages have been intimately associated throughout their evolutionary history and that the speciation of one partner has driven the speciation of the other.4, 12
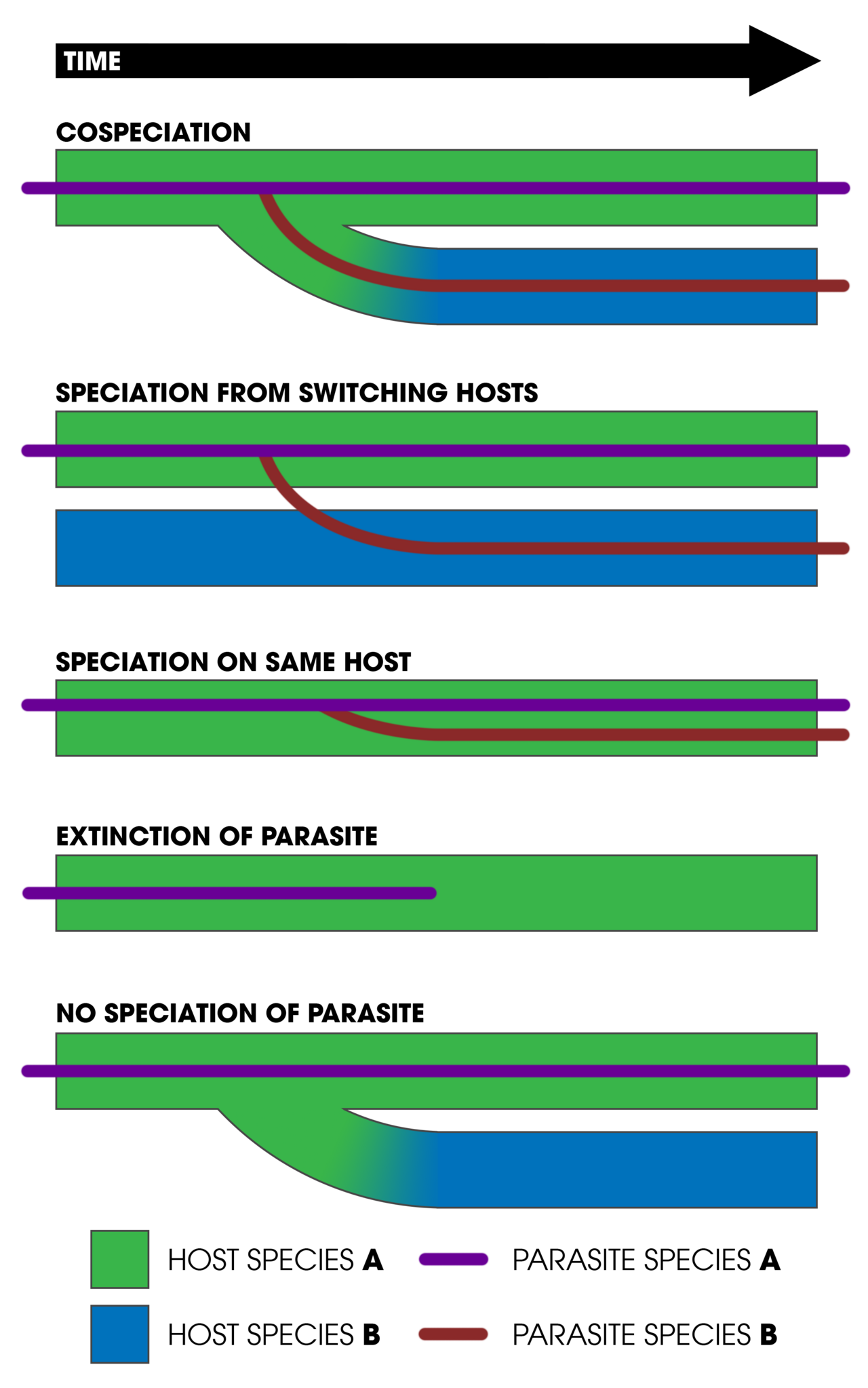
Cospeciation is of broad interest in evolutionary biology because it provides a test of the intimacy and duration of ecological associations, offers an independent means of calibrating molecular clocks, and furnishes evidence for the role of coevolution in generating biodiversity. However, strict cospeciation — perfect topological congruence between host and associate phylogenies — is rarely observed in practice. Most systems exhibit a mixture of cospeciation events, host switches (the associate colonises a new host lineage), duplications (the associate speciates independently of the host), lineage sorting (the associate goes extinct in one host lineage), and failure to speciate (the host speciates but the associate does not).12, 16
Methods for detecting cospeciation
Assessing whether two phylogenies are more congruent than expected by chance requires formal statistical methods, because some degree of topological similarity can arise by coincidence even between unrelated trees. Several analytical frameworks have been developed to distinguish true cospeciation from other processes that can produce apparent congruence.11
Reconciliation analysis, pioneered by Page and Ronquist, maps the associate phylogeny onto the host phylogeny and identifies the minimum combination of cospeciation events, host switches, duplications, and losses required to reconcile any incongruences between the two trees. Each type of event is assigned a cost, and the optimal reconciliation is the one that minimises total cost. If the number of inferred cospeciation events significantly exceeds the number expected under a random association, the null hypothesis of no cospeciation is rejected.4, 11
ParaFit, developed by Legendre, Desdevises, and Bazin, takes a different approach. Rather than comparing tree topologies directly, it tests whether the host-associate links (the specific pairings of host species with their associates) are significantly correlated with the phylogenetic positions of those species in their respective trees. This method can identify individual host-associate links that contribute to the overall cophylogenetic signal and those that do not, allowing researchers to pinpoint which parts of the association reflect cospeciation and which reflect host switching.5
Event-based methods such as Jane and eMPRess extend reconciliation analysis by incorporating temporal information (divergence dates) and allowing more complex models of host-associate interaction, including tanglegram-based approaches that visualise the correspondence between host and associate phylogenies.3, 12, 16
Pocket gophers and chewing lice
The association between pocket gophers (Geomyidae) and their chewing lice (Trichodectidae) is one of the most thoroughly documented cases of cospeciation. Hafner and Nadler published the first formal comparison of gopher and louse phylogenies in 1988, demonstrating a striking degree of topological congruence: the branching order of the louse phylogeny closely mirrored that of the gopher phylogeny, with most host-louse pairs representing cospeciation events.1 Subsequent work incorporating additional taxa and molecular data confirmed the overall pattern, while also identifying a small number of host-switching events that introduced incongruences into the otherwise congruent trees.2
The gopher-louse system owes its exceptional cospeciation signal to the biology of both partners. Pocket gophers are highly fossorial (burrowing) rodents with limited dispersal and minimal interspecific contact, which restricts the opportunities for lice to transfer between host species. The lice themselves are permanent ectoparasites with no free-living stage, spending their entire life cycle on the host, and they are transmitted primarily through direct physical contact between mother and offspring. These life-history features create strong barriers to host switching and favour the parallel diversification of host and parasite lineages.1, 12
An additional finding from the gopher-louse system was that rates of molecular evolution differ between hosts and parasites. Hafner and colleagues estimated that the lice have evolved approximately 2.8 times faster than their gopher hosts at mitochondrial loci, a pattern attributed to the shorter generation times and smaller effective population sizes of the parasites. This rate difference has implications for using cospeciation to calibrate molecular clocks, as it demonstrates that host and associate lineages can diverge at different rates even when their speciation events are synchronised.2
Figs and fig wasps
The mutualism between figs (Ficus) and their pollinator wasps (Agaonidae) is another classic system for studying cospeciation. Each of the roughly 750 fig species is pollinated by one or a few species of agaonid wasp, and the wasps can reproduce only inside the enclosed inflorescences of their host fig, creating an obligate reciprocal dependence that has persisted for an estimated 60 to 90 million years.6 Early phylogenetic analyses revealed broadly congruent patterns of branching between fig and wasp phylogenies, consistent with long-term cospeciation.6
More recent and more densely sampled phylogenetic studies, however, have revealed a more complex picture. While the overall correspondence between fig and wasp trees remains statistically significant, there are numerous incongruences attributable to host switching, duplication, and extinction. Machado and colleagues found that strict one-to-one cospeciation accounts for only a fraction of the observed diversity, with host switches occurring at appreciable rates, particularly in tropical regions where multiple fig species co-occur and wasps have opportunities to colonise new hosts.7 Cruaud and colleagues confirmed this mixed pattern with an expanded dataset, concluding that the fig-wasp association exhibits "fuzzy" cospeciation: a statistically significant signal of parallel diversification overlaid with substantial host switching and occasional periods of independent diversification.10
Primates and their microbes
Cospeciation is not limited to macroscopic parasites and mutualists. Studies of primate-associated microorganisms have revealed cospeciation at multiple levels of the microbial world. The simian immunodeficiency viruses (SIVs) that infect African primates show a pattern of partial phylogenetic congruence with their primate hosts, suggesting that some SIV lineages have cospeciated with their hosts over millions of years while others represent cross-species transmission events. The lentiviruses that gave rise to HIV-1 and HIV-2, for example, originated through relatively recent host switches from chimpanzees and sooty mangabeys, respectively, to humans.8, 9
At the level of the gut microbiome, Moeller and colleagues demonstrated in 2016 that specific lineages of gut bacteria have cospeciated with their great ape hosts over a timescale of millions of years. By comparing the phylogenies of Bacteroidaceae bacterial strains isolated from humans, chimpanzees, bonobos, and gorillas with the known phylogeny of the great apes, they found significant topological congruence, indicating that these bacterial lineages have been vertically transmitted from mother to offspring and have diversified in parallel with their hosts since the divergence of the great ape species.14 This finding extended the concept of cospeciation to the microbiome, demonstrating that host speciation can drive the diversification of entire microbial communities, not just individual parasite or mutualist species.
Organellar cospeciation
An often-overlooked example of cospeciation involves the relationship between eukaryotic hosts and their mitochondria. Because mitochondria are maternally inherited and do not undergo horizontal transfer between species, the mitochondrial genome has effectively cospeciated with the host nuclear genome since the original endosymbiotic event. This obligate cospeciation has functional consequences: mitochondrial-encoded proteins must interact physically with nuclear-encoded proteins in the electron transport chain and other mitochondrial complexes, creating selection for coevolution between the two genomes. Rand and colleagues documented evidence for compensatory molecular evolution between mitochondrial and nuclear genes in multiple animal lineages, a signature of the tight functional integration that results from millions of years of obligate cospeciation.13
Limits and evolutionary significance
The study of cospeciation has matured considerably since the early expectation that intimate ecological associations should routinely produce matching phylogenies. It is now recognised that strict cospeciation is the exception rather than the rule: even highly specialised host-associate systems typically show a mixture of cospeciation, host switching, and other processes.16 The prevalence of host switching means that phylogenetic congruence alone is not sufficient evidence for cospeciation; statistical tests are required to determine whether the observed level of congruence exceeds what would be expected by chance.5, 12
Thompson's geographic mosaic theory provides a framework for understanding why strict cospeciation is uncommon. If the strength and nature of the interaction between host and associate varies across geography — with coevolutionary hotspots and coldspots — then the evolutionary trajectory of the association is not uniform across the species range, and different populations may experience cospeciation, host switching, or independent diversification depending on local ecological conditions.15
Despite these complexities, cospeciation remains a powerful concept in evolutionary biology. Where it occurs, it provides direct evidence that ecological associations can persist through speciation events and shape the diversification of lineages over millions of years. It offers an independent means of dating evolutionary events when fossil calibrations are unavailable. And it demonstrates that the evolutionary history of any organism is inextricably linked to the evolutionary histories of its ecological partners, reinforcing the view that coevolution is a fundamental driver of biodiversity.12, 15
References
Cospeciation vs host-shift speciation: methods for testing, evidence from natural associations and relation to coevolution