Overview
- Transposable elements—mobile DNA sequences that copy and paste themselves throughout the genome—make up approximately 45% of the human genome, including over one million Alu elements that alone account for 11% of human DNA.
- When a transposable element inserts at a specific chromosomal location, it creates a unique molecular marker; humans and chimpanzees share thousands of transposable element insertions at identical genomic positions, each one an independent confirmation of shared ancestry.
- The probability of two independent insertions landing at the same nucleotide position is negligibly small (approximately 1 in billions), making shared TE insertions among the most statistically powerful forms of evidence for common descent.
Nearly half of the human genome is composed of transposable elements—mobile DNA sequences that have copied and inserted themselves throughout our chromosomes over hundreds of millions of years. These "jumping genes," first discovered by Barbara McClintock in maize in the 1940s, include several major classes: DNA transposons that move by a cut-and-paste mechanism, and retrotransposons (LINEs, SINEs, and LTR retrotransposons) that copy themselves through an RNA intermediate.1, 5 Together, these elements account for approximately 45% of human genomic DNA—far more than the roughly 1.5% that encodes proteins.1 Like endogenous retroviruses, shared transposable element insertions at identical genomic positions across species constitute powerful molecular evidence for common descent, providing an independent line of confirmation from a different class of mobile DNA.
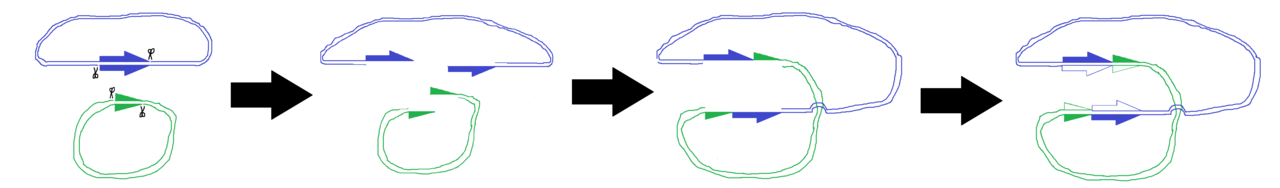
Classes of transposable elements
The human genome harbors several major families of transposable elements, each with a distinct replication mechanism and evolutionary history. Long interspersed nuclear elements (LINEs) are autonomous retrotransposons that encode the enzymes required for their own reverse transcription and integration. The LINE-1 (L1) family is the most abundant, with approximately 500,000 copies in the human genome, though the vast majority are truncated, mutated fragments. Only 80–100 full-length L1 elements remain capable of active retrotransposition in any given human genome.1, 9
Short interspersed nuclear elements (SINEs) are non-autonomous retrotransposons that depend on the enzymatic machinery of LINEs for their mobilization. The most prominent SINE family in primates is the Alu element, a roughly 300-base-pair sequence derived from the 7SL RNA gene. Alu elements are extraordinarily abundant: the human genome contains over one million copies, accounting for approximately 11% of total genomic DNA.1, 4 New Alu insertions continue to occur at an estimated rate of one new insertion per 20 births, making them one of the most active classes of mobile DNA in the human genome today.4
DNA transposons, unlike retrotransposons, move by a cut-and-paste mechanism that does not involve an RNA intermediate. They were highly active early in mammalian evolution but appear to have become largely inactive in the primate lineage tens of millions of years ago. Fossil DNA transposons now constitute approximately 3% of the human genome.1, 5 LTR retrotransposons, which include endogenous retroviruses, use a mechanism similar to retroviruses and are treated in a separate article.
Shared insertions as markers of common ancestry
When a transposable element inserts at a particular position in the genome, it creates a unique molecular marker at that site. The insertion generates a target site duplication—a short sequence of host DNA that is duplicated on either side of the inserted element—producing a molecular fingerprint that identifies both the element and the specific site of its insertion.5, 6 Because the human genome contains over three billion base pairs and TE insertion, while not perfectly random, shows enormous positional diversity, the probability that two independent insertion events would land at precisely the same nucleotide in two different species is negligibly small—on the order of one in several billion for any given site.6, 7
When researchers compare the genomes of humans and chimpanzees, they find thousands of Alu elements, L1 elements, and other transposable elements at identical chromosomal positions in both species, flanked by identical target site duplications.8 The same pattern extends across primate phylogeny: insertions shared by humans and chimpanzees but absent from gorillas mark events that occurred after the gorilla lineage diverged but before the human-chimpanzee split. Insertions shared by all great apes but absent from Old World monkeys date to the common ancestor of great apes. The distribution of shared insertions perfectly tracks the known phylogenetic tree, providing thousands of independent confirmations of the branching order.4, 7
Nearly homoplasy-free phylogenetic markers
Transposable element insertions are considered "nearly perfect" phylogenetic markers because they have several properties that make them exceptionally reliable for reconstructing evolutionary relationships. First, the probability of independent insertion at the same site is vanishingly small, as discussed above. Second, TE insertions are essentially irreversible—once an element is integrated, precise excision that restores the original pre-insertion sequence is extremely rare. Third, the ancestral state (absence of an insertion at a given locus) can be determined by comparison with outgroup species.6, 7
These properties mean that shared TE insertions are virtually free from homoplasy—the phenomenon in which the same character state arises independently in different lineages through convergent evolution. Homoplasy is a significant problem for many types of phylogenetic data (morphological characters, individual nucleotide substitutions), but it is negligible for TE insertions. A shared insertion at an orthologous locus in two species is, for all practical purposes, proof that the insertion occurred once in their common ancestor.6, 7
Resolving the phylogeny of whales
One of the most celebrated applications of shared TE insertions to phylogenetic questions involved the relationship of whales to other mammals. Morphological and some molecular data suggested that cetaceans (whales, dolphins, and porpoises) were closely related to artiodactyls (even-toed ungulates such as cows, pigs, and hippos), but the precise placement of whales within this group was debated. Nikaido, Rooney, and Okada used shared SINE insertions to demonstrate that whales are nested within the artiodactyls, with hippos as their closest living relatives.8
They identified specific SINE insertions shared by whales and hippos but absent from other artiodactyls, and additional insertions shared by the whale-hippo clade with ruminants to the exclusion of pigs. Because SINEs are essentially homoplasy-free markers, each shared insertion represented an unambiguous synapomorphy (shared derived character) linking these taxa. Subsequent phylogenomic analyses using larger data sets of retroposon insertions confirmed this topology with overwhelming statistical support.8, 9
Alu elements and primate evolution
Alu elements have been particularly informative for reconstructing primate phylogeny. Because Alu elements are primate-specific (they amplified to their current abundance only in the primate lineage), every Alu insertion shared by two primate species marks an event that occurred in their common primate ancestor. Batzer and Deininger documented hundreds of Alu insertions that are diagnostic for specific nodes in the primate tree: insertions shared by all catarrhines (Old World monkeys and apes), insertions shared by all hominoids (apes), insertions shared by all great apes, and insertions shared exclusively by humans and chimpanzees.4
The distribution of these lineage-specific Alu insertions is fully consistent with the primate phylogeny inferred from morphological, fossil, and other molecular evidence. Furthermore, the relative number of lineage-specific insertions on each branch of the tree is proportional to the estimated branch length, consistent with a roughly constant rate of Alu retrotransposition over primate evolution.4, 7
Occasional co-option of transposable elements
While the vast majority of TE insertions are selectively neutral or mildly deleterious, a small fraction have been co-opted (exapted) for functional roles in the host genome. Lowe, Bejerano, and Haussler identified thousands of TE-derived sequences in the human genome that have been conserved under strong purifying selection, many located near developmental genes where they appear to function as cis-regulatory enhancers.10 These co-opted elements illustrate how the random insertion of transposable elements can, on rare occasions, provide raw material for evolutionary innovation.
However, the functional co-option of individual TE insertions does not diminish their value as phylogenetic markers. Whether a shared insertion is functional or neutral, its presence at the same genomic locus in two species still indicates inheritance from a common ancestor. The evidence for common descent from shared TE insertions is based on the statistical impossibility of independent insertion at the same site, a property that holds regardless of whether the element subsequently acquired a function.6, 7
Independent confirmation of ERV evidence
Shared transposable element insertions provide evidence for common descent that is independent of, but concordant with, the evidence from shared endogenous retroviruses and shared pseudogenes. ERVs, SINEs, LINEs, and DNA transposons are different classes of mobile genetic elements with different replication mechanisms, different insertion preferences, and different evolutionary histories. Yet all four classes produce the same phylogenetic signal when their shared insertions are mapped across species.7, 9 The convergence of these independent lines of evidence on the same evolutionary tree is a hallmark of a well-supported scientific conclusion. Each class of shared insertion constitutes an independent test of the common descent hypothesis, and each test returns the same result.6, 7
A unified classification system for transposable elements
The diversity of transposable elements across genomes has prompted the development of systematic classification frameworks. Wicker and colleagues proposed a hierarchical classification that divides all TEs into two classes based on their transposition mechanism: Class I elements (retrotransposons), which mobilise through an RNA intermediate via a "copy-and-paste" mechanism, and Class II elements (DNA transposons), which move through a DNA intermediate via "cut-and-paste" or rolling-circle replication. Class I is further subdivided into LTR retrotransposons, LINEs, and SINEs, while Class II includes terminal inverted repeat (TIR) transposons and Helitrons.11 This classification has become the standard reference for TE annotation in genome projects and facilitates consistent comparison of TE content across species.
Genome-wide surveys of TE insertion polymorphisms within human populations have further extended the utility of transposable elements as evolutionary markers. Stewart and colleagues catalogued over 7,000 polymorphic TE insertions (sites where the insertion is present in some individuals but absent in others) across human populations, demonstrating that TE polymorphisms track population structure and migration history in the same way as single-nucleotide polymorphisms but with the added advantage of being effectively homoplasy-free.12 These population-level TE polymorphisms provide a bridge between the deep phylogenetic timescale (where fixed shared insertions mark species-level divergences) and the microevolutionary timescale of recent human demographic history.4, 12
Unexpected phylogenetic discoveries from retroposon data
Retroposon insertions have not only confirmed established phylogenies but also revealed unexpected evolutionary relationships that challenged previous classifications. Nishihara and colleagues used retroposon insertion data to identify Pegasoferae, a clade uniting bats, horses, and carnivorans within the Laurasiatherian mammals—a grouping that was not predicted by morphological analysis and was initially controversial but has since received support from additional genomic evidence.13 Similarly, retroposon data provided the definitive resolution of the placement of turtles within Archosauria, grouping them with birds and crocodilians rather than with lizards and snakes, overturning a long-held morphological classification.7, 9 These cases demonstrate that the virtually homoplasy-free nature of TE insertions gives them a unique capacity to resolve phylogenetic questions that remain ambiguous when analysed with other types of molecular or morphological data.
References
Phylogenomic analysis resolves the interordinal relationships and rapid diversification of the Laurasiatherian mammals
Thousands of human mobile element fragments undergo strong purifying selection near developmental genes
Pegasoferae, an unexpected mammalian clade revealed by tracking ancient retroposon insertions