Overview
- Epigenetics encompasses heritable changes in gene expression that occur without alterations to the underlying DNA sequence, mediated by mechanisms including DNA methylation, histone modification, chromatin remodeling, and non-coding RNAs.
- Transgenerational epigenetic inheritance has been documented in plants, nematodes, and mammals, though its prevalence and evolutionary significance in animals remain subjects of active scientific debate.
- The Extended Evolutionary Synthesis incorporates epigenetic inheritance alongside genetic assimilation, niche construction, and developmental bias as processes that may broaden our understanding of how organisms adapt and evolve.
Epigenetics is the study of heritable changes in gene expression that occur without alterations to the underlying DNA sequence. The term was coined by Conrad Hal Waddington in 1942, who defined the "epigenotype" as the complex of developmental processes lying between genotype and phenotype.1 In the decades since, the field has expanded dramatically, and a modern consensus defines an epigenetic trait as a stably heritable phenotype resulting from changes in a chromosome without alterations in the DNA sequence itself.2 Epigenetic mechanisms — including DNA methylation, histone modification, and regulation by non-coding RNAs — govern how genes are switched on and off in different cell types, at different developmental stages, and in response to environmental signals. They explain, for example, how genetically identical cells in a multicellular organism can differentiate into nerve cells, blood cells, and muscle cells, each expressing a distinct subset of the genome.
The evolutionary significance of epigenetics has become a subject of intense scientific interest and debate. If epigenetic modifications can be transmitted across generations — a phenomenon called transgenerational epigenetic inheritance — then heritable variation is not confined to DNA sequence alone, and the sources of variation upon which natural selection can act may be broader than the Modern Synthesis originally envisioned.4, 14 Evidence for transgenerational epigenetic inheritance is strongest in plants, well-documented in nematodes and rodents under laboratory conditions, and suggestive but more contested in humans. The degree to which these phenomena matter for evolution in the wild, and whether they necessitate a fundamental revision of evolutionary theory, remain open questions that lie at the heart of the ongoing conversation about the Extended Evolutionary Synthesis.16
Molecular mechanisms of epigenetic regulation
Epigenetic regulation operates through several distinct but interrelated molecular mechanisms, all of which modulate gene expression by altering the accessibility of DNA to the transcriptional machinery rather than by changing the DNA sequence itself.
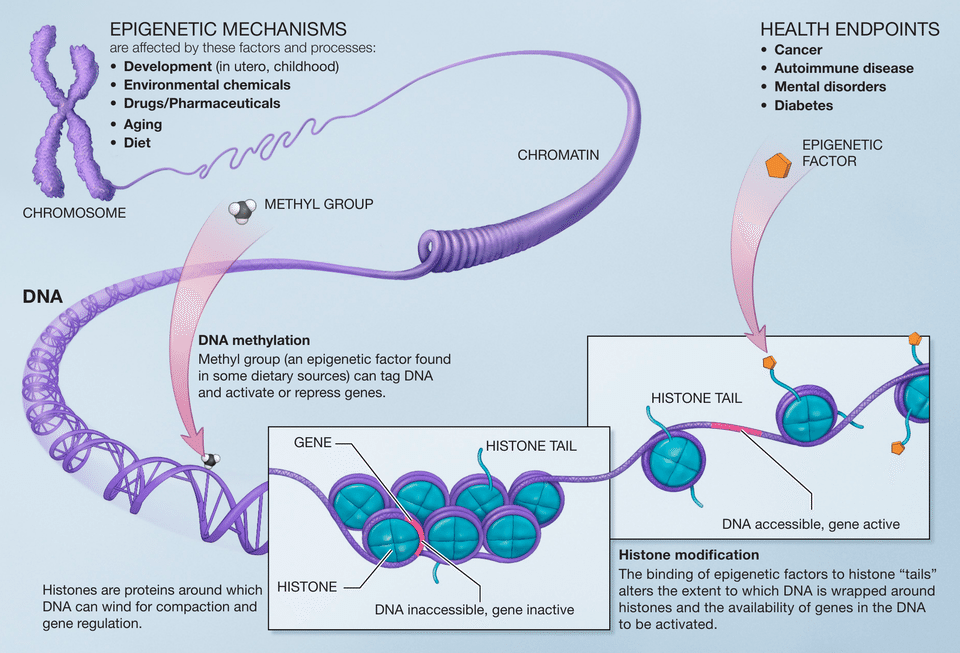
The most extensively studied of these mechanisms is DNA methylation, the covalent addition of a methyl group to the 5-carbon position of cytosine bases, most commonly at cytosine-guanine (CpG) dinucleotides in mammals. When CpG-rich regions called CpG islands in gene promoters are heavily methylated, the associated gene is typically silenced, because the methyl groups either directly block the binding of transcription factors or recruit methyl-binding proteins that remodel the surrounding chromatin into a repressive state.2, 4 DNA methylation patterns are maintained through cell division by the maintenance methyltransferase DNMT1, which copies methylation marks from the parental strand to the newly synthesised daughter strand, providing a mechanism for epigenetic memory across mitotic generations of cells.2
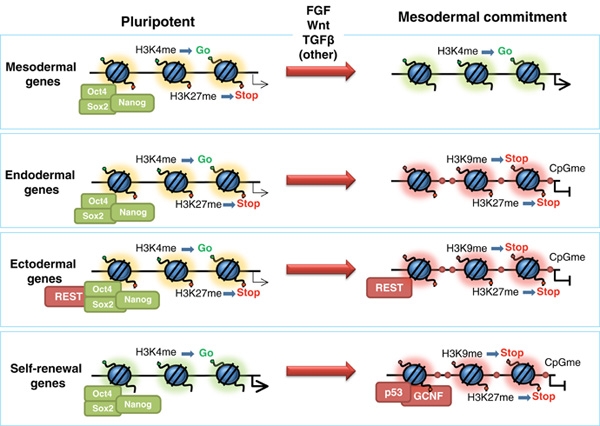
A second major layer of epigenetic information resides in the post-translational modification of histone proteins, the core structural components around which DNA is wrapped to form nucleosomes. Histones can be acetylated, methylated, phosphorylated, ubiquitinated, and subjected to numerous other chemical modifications, principally on their protruding N-terminal tails. Different combinations and patterns of these modifications influence whether the surrounding chromatin adopts an open, transcriptionally active conformation (euchromatin) or a compact, transcriptionally silent conformation (heterochromatin). Strahl and Allis proposed the histone code hypothesis in 2000, suggesting that specific patterns of histone modifications act in combination to form a regulatory language read by effector proteins to bring about distinct downstream events in gene expression, DNA repair, and chromosome condensation.3 Histone acetylation, for instance, is broadly associated with gene activation, whereas trimethylation of histone H3 at lysine 27 (H3K27me3) is associated with gene silencing by Polycomb group proteins.3
Additional epigenetic mechanisms include chromatin remodeling by ATP-dependent protein complexes that physically reposition or eject nucleosomes to expose or occlude regulatory DNA sequences, and non-coding RNAs — including microRNAs (miRNAs), long non-coding RNAs (lncRNAs), and small interfering RNAs (siRNAs) — that can guide chromatin-modifying complexes to specific genomic loci or directly modulate messenger RNA stability and translation.2, 12 Together, these mechanisms constitute an epigenetic regulatory system of enormous combinatorial complexity, capable of fine-tuning gene expression in response to developmental cues, cellular signals, and environmental stimuli.
Transgenerational epigenetic inheritance in plants
The strongest and most abundant evidence for transgenerational epigenetic inheritance comes from plants.

The relative stability of epigenetic marks in plant genomes, the absence of a segregated germline (meaning that somatic mutations and epimutations arising during vegetative growth can be transmitted to the next generation through meristematic tissue), and the well-characterised role of RNA-directed DNA methylation in plants all contribute to making epigenetic inheritance more readily observable in the plant kingdom than in animals.12, 13
One of the earliest and most striking examples is paramutation, first described by Alexander Brink in 1956 at the red1 (r1) locus in maize. In paramutation, one allele (termed the paramutagenic allele) heritably silences or reduces the expression of a homologous allele (the paramutable allele) when both are present in the same nucleus. The resulting silenced state is meiotically heritable: the newly silenced allele retains its reduced expression even after it segregates away from the paramutagenic allele in subsequent generations and can itself act as a paramutagenic allele, silencing further paramutable alleles. Molecular studies have revealed that paramutation at the maize b1 locus is mediated by tandem repeats that produce small interfering RNAs, which direct DNA methylation and chromatin compaction at the target locus.6 Paramutation-like phenomena have since been identified in other plant species, in fungi, and in mice, suggesting a broadly conserved mechanism involving small RNA-mediated chromatin silencing.6
A landmark study by Cubas, Vincent, and Coen in 1999 provided one of the first demonstrations that an epigenetic change in a single gene could produce a heritable, ecologically visible phenotype in a natural plant population. The researchers characterised a naturally occurring variant of Linaria vulgaris (toadflax) in which the flower symmetry is changed from the normal bilateral form to a radial form — a mutant originally described by Linnaeus more than 250 years earlier. They found that the Lcyc gene, a homologue of the cycloidea gene controlling floral asymmetry, was extensively methylated and transcriptionally silent in the mutant. The methylation was heritable and co-segregated with the mutant phenotype across generations, and occasional somatic reversion of the flower shape correlated with demethylation and restoration of Lcyc expression.5 The discovery that the first naturally occurring morphological variant to be molecularly characterised was epigenetic rather than genetic in origin underscored the potential evolutionary significance of epimutation.
Large-scale studies in the model plant Arabidopsis thaliana have further established the scope of heritable epigenetic variation. Schmitz and colleagues propagated Arabidopsis lines by single-seed descent for 30 generations and identified over 114,000 spontaneous single-methylation polymorphisms and nearly 2,500 differentially methylated regions (DMRs) that had arisen in the absence of any intentional selection or environmental perturbation. Strikingly, the spontaneous epimutation rate at CpG sites was several orders of magnitude higher than the genetic point mutation rate, and some of the observed DMRs were associated with measurable changes in gene expression.7 These findings demonstrated that epigenetic variation can arise spontaneously and accumulate over generations, potentially providing a substrate for natural selection independent of genetic mutation.
Evidence in animals
Transgenerational epigenetic inheritance in animals is more controversial than in plants, in part because the mammalian germline undergoes two waves of genome-wide epigenetic reprogramming — first in the primordial germ cells and again after fertilisation — that erase most existing methylation marks and re-establish them de novo.12
The agouti viable yellow (Avy) mouse is one of the best-studied models of epigenetic inheritance in mammals. In these mice, an intracisternal A-particle (IAP) retrotransposon inserted upstream of the agouti gene provides a cryptic promoter that drives ectopic agouti expression. The degree of DNA methylation at this IAP element varies among genetically identical individuals, producing a continuous spectrum of coat colour from yellow (unmethylated, with ectopic agouti expression causing obesity, diabetes, and tumour susceptibility) to brown or pseudoagouti (fully methylated, with normal agouti regulation and lean body habitus). Morgan and colleagues demonstrated in 1999 that the methylation state of the Avy allele could be transmitted from mother to offspring, with yellow dams tending to produce more yellow pups than pseudoagouti dams, constituting an example of epigenetic inheritance through the female germline.9 Waterland and Jirtle subsequently showed that maternal dietary supplementation with methyl donors (folic acid, vitamin B12, choline, and betaine) shifted the offspring coat-colour distribution toward brown by increasing CpG methylation at the Avy locus, demonstrating that a nutritional environmental exposure during gestation could alter the epigenetic state and phenotype of the next generation.10
The vinclozolin rat model provided some of the most dramatic and contested claims of transgenerational epigenetic inheritance in mammals. Anway and colleagues reported in 2005 that transient exposure of pregnant rats to vinclozolin, an endocrine-disrupting fungicide, during the period of fetal gonadal sex determination produced effects — including reduced sperm counts, increased rates of infertility, and elevated incidence of prostate and kidney disease — that persisted through the male germline for at least four generations (F1 through F4) without any further exposure. The transgenerational phenotype was associated with altered DNA methylation patterns in the male germline.8 These results generated intense debate. Subsequent independent attempts to replicate the findings produced mixed results, and critics noted methodological concerns including potential confounding by direct fetal exposure in the F1 and germ-cell exposure in the F2 generation, which complicates the attribution of effects in later generations to genuinely transgenerational epigenetic inheritance.12, 24
In humans, the best-known evidence for epigenetic effects of early-life environment comes from the Dutch Hunger Winter cohort. During the winter of 1944–1945, a Nazi-imposed food embargo subjected the western Netherlands to severe famine. Decades later, Heijmans and colleagues compared individuals who had been conceived during the famine with their unexposed same-sex siblings and found that periconceptional famine exposure was associated with a 5.2 percent reduction in DNA methylation at the imprinted IGF2 gene, a growth factor gene regulated by genomic imprinting — an effect that persisted six decades after the exposure.11 While this study demonstrated that early-life nutritional stress can produce lasting epigenetic changes in humans, it documented intragenerational persistence of epigenetic marks rather than their transmission to the next generation, and the distinction between intragenerational and transgenerational effects remains critical to the evolutionary debate.12
Major epigenetic mechanisms and their inheritance potential
Epigenetic mechanisms and evidence for transgenerational inheritance2, 4, 12
| Mechanism | Molecular basis | Transgenerational in plants | Transgenerational in animals |
|---|---|---|---|
| DNA methylation | Methyl group on cytosine (5mC), maintained by DNMT1 | Abundant evidence | Limited; most marks erased during reprogramming |
| Histone modifications | Acetylation, methylation, phosphorylation of histone tails | Some evidence (H3K27me3) | Emerging evidence in C. elegans; limited in mammals |
| Chromatin remodeling | ATP-dependent repositioning of nucleosomes | Indirect evidence via paramutation | Unclear |
| Small non-coding RNAs | siRNAs, piRNAs guide chromatin silencing | Well-documented (RNA-directed DNA methylation) | Evidence in C. elegans and mice (piRNAs, miRNAs in sperm) |
| Genomic imprinting | Parent-of-origin methylation at imprinted loci | Limited to certain loci | Maintained through reprogramming at ~100–200 genes |
The table above summarises the major molecular mechanisms of epigenetic regulation and the current state of evidence for their transgenerational transmission in plants and animals. As the table illustrates, the evidence is strongest for DNA methylation and small RNA-mediated silencing in plants, where mechanisms for the maintenance and re-establishment of epigenetic marks are well characterised. In animals, the two waves of genome-wide epigenetic reprogramming during gametogenesis and early embryogenesis present a substantial barrier to transgenerational transmission, though genomic imprints and certain retrotransposon-associated marks are known to resist reprogramming.12
The debate over evolutionary significance
The central question in the relationship between epigenetics and evolution is not whether epigenetic mechanisms exist — their role in development, cell differentiation, and gene regulation is firmly established — but whether transgenerational epigenetic inheritance is sufficiently widespread, stable, and responsive to selection to constitute an evolutionarily significant source of heritable variation. On this question, the scientific community is genuinely divided.
The most prominent advocates for the evolutionary importance of epigenetic inheritance have been Eva Jablonka and Marion Lamb, whose 2005 book Evolution in Four Dimensions argued that evolution operates through four channels of inheritance — genetic, epigenetic, behavioural, and symbolic — and that the Modern Synthesis, by focusing almost exclusively on genetic inheritance, has overlooked the contributions of the other three.14 In a comprehensive 2009 review, Jablonka and Raz catalogued over one hundred documented cases of transgenerational epigenetic inheritance across bacteria, protists, fungi, plants, and animals, arguing that the phenomenon is far more prevalent than commonly assumed and that it has the potential to influence evolutionary dynamics by generating heritable variation that is responsive to the environment and that can arise at rates far exceeding those of genetic mutation.13
Critics have challenged these claims on several grounds. Dickins and Rahman argued in 2012 that proponents of an extended evolutionary synthesis have conflated proximate and ultimate causation. In their view, epigenetic inheritance systems are best understood as proximate mechanisms — evolved responses that calibrate phenotypic development to environmental conditions in order to maximise fitness — rather than as a fundamentally new dimension of heredity that requires extending evolutionary theory. They contended that the existing framework of the Modern Synthesis, with its well-developed understanding of gene-environment interaction, phenotypic plasticity, and reaction norms, can accommodate epigenetic phenomena without requiring a conceptual revolution.15 Others have noted that many reported cases of transgenerational epigenetic inheritance in animals involve only two or three generations, making it difficult to distinguish genuinely transgenerational effects from direct exposure of fetal or germ-cell tissue, and that the long-term stability of epigenetic marks over evolutionary timescales remains poorly characterised.12
A measured assessment, articulated in a widely cited review by Heard and Martienssen, concludes that transgenerational epigenetic inheritance certainly occurs — particularly in plants — but that how much is attributable to environmental induction, how stable it is over many generations, and the extent to which it occurs in humans and other mammals remain open questions.12 The debate is not about whether epigenetics is real or important; it is about the scope of its contribution to heritable variation and adaptive evolution relative to the well-established primacy of DNA sequence variation.
Epigenetics and the Extended Evolutionary Synthesis
The question of epigenetic inheritance intersects with a broader conversation about whether the conceptual framework of evolutionary biology needs to be updated. The Extended Evolutionary Synthesis (EES) is a proposed expansion of standard evolutionary theory that incorporates processes including developmental bias, inclusive inheritance (genetic, epigenetic, ecological, and cultural), niche construction, and phenotypic plasticity as factors that can direct and shape evolutionary change in ways not fully captured by the gene-centred framework of the Modern Synthesis.16
Danchin and colleagues formalised the concept of inclusive inheritance in 2011, arguing that biological information is transmitted across generations not only through DNA sequence but also through epigenetic marks, parental effects, ecological inheritance (modifications of the environment that affect offspring), and cultural transmission. They proposed the concept of "inclusive heritability" to unify these multiple channels into a single quantitative framework, analogous to the way inclusive fitness unified direct and indirect effects on reproductive success.18 Epigenetic inheritance occupies a particularly interesting position in this framework because it is molecular and heritable — like genetic inheritance — but potentially responsive to environmental conditions in a way that DNA sequence is not.
Laland and colleagues articulated the full structure of the EES in 2015, identifying several core assumptions that distinguish it from the standard framework. Among these is the claim that developmental processes, including epigenetic modification, do not merely transmit variation but actively generate it in ways that may be biased toward functional outcomes — a concept termed developmental bias. If epigenetic changes tend to affect genes in regulatory networks that are already associated with environmental responsiveness, then the variation they produce may be more likely to be adaptive than random genetic mutations, which have no such directional tendency.16 Proponents of the EES do not argue that natural selection is unimportant, but rather that the processes generating and transmitting heritable variation are more diverse than traditionally recognised, and that incorporating these processes yields a richer and more accurate picture of evolutionary change.16
Genetic assimilation and the epigenetic landscape
The relationship between epigenetics and evolution has deep historical roots in the work of Waddington, who not only coined the term "epigenetics" but also developed the concepts of canalization and genetic assimilation that remain central to current discussions. Waddington proposed that developmental pathways are "canalised" — buffered against both genetic and environmental perturbation — so that the organism reliably produces the same phenotype despite minor variation in its genes or environment. He visualised this concept through the metaphor of the epigenetic landscape, in which a ball (representing the developing cell or organism) rolls down a surface of branching valleys, with each valley representing a canalised developmental trajectory.1
In a landmark 1953 experiment, Waddington demonstrated genetic assimilation in Drosophila melanogaster. He exposed fly pupae to heat shock, which induced a crossveinless wing phenotype in a fraction of the population. By selecting for the crossveinless phenotype over multiple generations while continuing to apply the heat shock, he produced lines in which an increasing proportion of flies showed the phenotype. Eventually, the crossveinless phenotype appeared even without any heat-shock treatment — an environmentally induced trait had become genetically fixed.17 Waddington interpreted this result as evidence that selection had acted on the existing genetic variation underlying the threshold for environmental induction, shifting it until the phenotype was produced constitutively. Modern reinterpretations suggest that epigenetic mechanisms may facilitate this process: environmental stress could reveal cryptic epigenetic variation, which selection then stabilises, and the progressive fixation of the phenotype may involve both genetic and epigenetic changes.17, 23
West-Eberhard extended this line of reasoning in 2003 with her concept of phenotypic accommodation, arguing that developmental systems can self-adjust to produce integrated, functional phenotypes in response to novel environmental challenges without requiring any genetic change. Genes, in her view, are "more often followers than leaders in evolutionary change" — environmental induction of a novel phenotype occurs first, and genetic accommodation (including but not limited to genetic assimilation) follows, fixing the novel phenotype in the population.23 Epigenetic mechanisms provide a plausible molecular substrate for this process, offering a way for environmental signals to produce heritable phenotypic variation that can persist long enough for genetic changes to catch up.
Niche construction and ecological inheritance
A further connection between epigenetics and evolutionary theory arises through the concept of niche construction, formalised by Odling-Smee, Laland, and Feldman in 2003. Niche construction is the process by which organisms modify their own environment and, in doing so, alter the selection pressures acting on themselves and on other species sharing that environment. These environmental modifications can persist across generations, creating what Odling-Smee and colleagues termed ecological inheritance — a form of inheritance in which organisms inherit not only genes from their parents but also a modified set of environmental conditions.22
Epigenetic inheritance can interact with niche construction in a feedback loop that may accelerate adaptation. When organisms modify their environment through niche construction, the altered environmental conditions may induce epigenetic changes in the organisms exposed to them. If these epigenetic changes are heritable, the descendants of the niche-constructing organisms inherit both the modified environment and an epigenetic predisposition to thrive in it.16, 22 For example, organisms that construct environments with particular dietary resources may undergo epigenetic changes in metabolic gene regulation that are transmitted to offspring who will encounter those same resources. Whether this kind of feedback between niche construction and epigenetic inheritance occurs at a scale that significantly influences evolution in natural populations remains to be empirically demonstrated, but it represents one of the most intriguing predictions of the Extended Evolutionary Synthesis.
Phenotypic plasticity and epigenetic variation
Phenotypic plasticity — the ability of a single genotype to produce different phenotypes in response to different environmental conditions — is one of the most ubiquitous features of living organisms, and epigenetic mechanisms are among its primary molecular mediators. When an organism encounters a new environment, changes in DNA methylation, histone modification, and non-coding RNA expression can alter gene expression profiles across the genome, producing physiological, morphological, or behavioural adjustments without any change in DNA sequence.19, 23
The relationship between epigenetic variation and phenotypic plasticity has been particularly well studied in plants. Lira-Medeiros and colleagues examined two populations of the white mangrove (Laguncularia racemosa) growing in contrasting natural environments — a riverside habitat and a salt-marsh habitat — in Sepetiba Bay, Brazil. Despite occupying habitats separated by only a short distance, individuals from the two populations showed marked morphological differences in plant height, leaf size, and other traits. Genetic analysis using molecular markers revealed minimal genetic differentiation between the populations, but analysis of DNA methylation using methylation-sensitive amplified polymorphism (MSAP) revealed substantial epigenetic differentiation, suggesting that epigenetic variation rather than genetic variation was associated with the phenotypic divergence between habitats.21
This pattern — low genetic diversity accompanied by high epigenetic diversity — has also been observed in the context of biological invasions. Schrey and colleagues studied introduced populations of house sparrows (Passer domesticus) in Kenya and Florida, both of which had passed through population bottlenecks during colonisation that reduced their genetic diversity. They found that epigenetic (methylation) diversity was maintained or even elevated relative to native-range populations despite the reduction in genetic variation, suggesting that epigenetic variation may compensate for the loss of genetic diversity and facilitate the rapid adaptation of invasive species to novel environments.20 If confirmed by further studies, this finding would have significant implications for understanding how populations with limited genetic variation can nevertheless adapt rapidly to new conditions — a puzzle long recognised in invasion biology and conservation genetics.19, 20
Epigenetics and rapid adaptation
One of the most consequential potential roles of epigenetics in evolution is its capacity to facilitate rapid adaptation on timescales far shorter than those required for the accumulation of beneficial genetic mutations. Because epimutations arise at rates orders of magnitude higher than genetic mutations — Schmitz and colleagues found spontaneous CpG methylation change rates in Arabidopsis of approximately 2–5 × 10−4 per site per generation, compared with a genetic point mutation rate of approximately 7 × 10−9 per site per generation — epigenetic variation can in principle be generated far more quickly than genetic variation.7 Furthermore, because some epigenetic changes are environmentally induced, they may arise in direct response to the selective pressure itself, producing directed (or at least biased) variation rather than the undirected variation characteristic of random genetic mutation.13, 19
This capacity for rapid, environmentally responsive heritable change could be particularly important in the context of climate change, where organisms face the challenge of adapting to environmental shifts occurring over decades — timescales too fast for most genetic adaptation in organisms with long generation times. Bossdorf, Richards, and Pigliucci noted that epigenetic variation could provide a mechanism for ecological adaptation that operates on a different timescale than genetic evolution, potentially allowing populations to track rapidly changing environments through heritable adjustments in gene expression.19 Similarly, Skinner and colleagues argued that the ability of environmental factors to alter the epigenome of the germline creates a mechanism by which environmental exposures can influence the phenotypes of subsequent generations, with implications for understanding how populations respond to novel environmental stressors such as endocrine-disrupting chemicals.24
However, the evolutionary significance of rapid epigenetic change depends critically on the stability and fidelity of epigenetic inheritance over the number of generations relevant to adaptive evolution. If epigenetic marks are too unstable — reverting to their original state within a few generations — they may influence short-term phenotypic responses without contributing to lasting evolutionary change. Conversely, if they are highly stable, they become functionally equivalent to genetic mutations and may be subject to the same population-genetic principles. The intermediate case — epigenetic marks that persist for several to many generations but are ultimately reversible — is the most theoretically interesting, because such marks could allow populations to explore adaptive phenotypic space rapidly and reversibly before genetic assimilation consolidates successful adaptations into permanent changes.12, 14 Determining where on this continuum of stability natural epigenetic variation actually falls, and how this varies among taxa, environments, and genomic contexts, is one of the central empirical challenges facing the field today.
References
Transposable elements: targets for early nutritional effects on epigenetic gene regulation
Persistent epigenetic differences associated with prenatal exposure to famine in humans
Transgenerational epigenetic inheritance: prevalence, mechanisms, and implications for the study of heredity and evolution
Evolution in Four Dimensions: Genetic, Epigenetic, Behavioral, and Symbolic Variation in the History of Life
Epigenetic variation may compensate for decreased genetic variation with introductions: a case study using house sparrows (Passer domesticus) on two continents