Overview
- Homologous structures are anatomical, developmental, or molecular features in different species that share a common evolutionary origin, and their existence across the tree of life constitutes one of the most powerful lines of evidence for common descent.
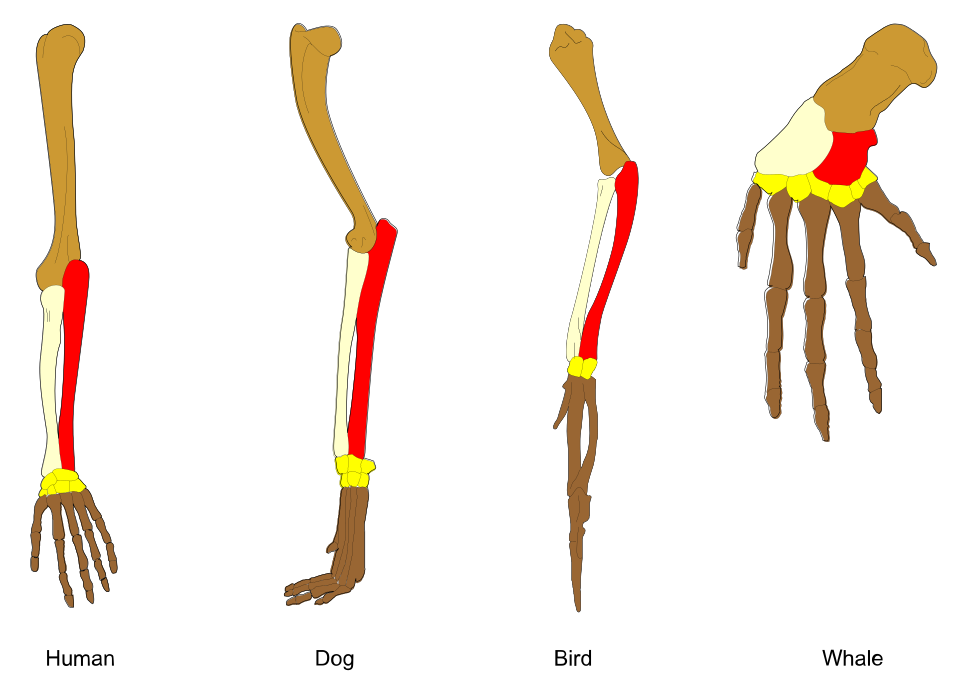
- The vertebrate forelimb exemplifies structural homology: the same arrangement of humerus, radius, ulna, carpals, metacarpals, and phalanges appears in the human arm, bat wing, whale flipper, and horse leg, modified for radically different functions but built from the same inherited blueprint.
- Molecular homologies, including conserved Hox gene clusters that pattern body axes from insects to humans, the universal cytochrome c protein whose sequence mirrors phylogenetic relationships, and the shared Pax6 gene that controls eye development across phyla, extend the evidence for common ancestry to the deepest levels of the genome.
Homologous structures are features in different species that share a common evolutionary origin, even when those features have been modified to serve very different functions. The human arm, the bat wing, the whale flipper, and the horse leg are all built from the same skeletal elements, inherited from a shared ancestor hundreds of millions of years ago and reshaped by natural selection to meet the demands of grasping, flying, swimming, and running.2, 3 This pattern of underlying structural similarity amid functional diversity is called homology, and it is one of the most compelling lines of evidence for common descent. Homology is found at every level of biological organization, from the gross anatomy of limbs and organs, to the developmental pathways that build embryos, to the molecular sequences of genes and proteins. The pervasive presence of homologous structures across the tree of life was central to Darwin's argument for evolution in 1859, and the subsequent discovery of molecular and developmental homologies in the twentieth and twenty-first centuries has only deepened the case.2, 4
Defining homology
The concept of homology predates evolutionary theory. In 1848, the British anatomist Richard Owen defined a homologue as "the same organ in different animals under every variety of form and function." Owen distinguished homology from analogy, which he defined as correspondence of function without correspondence of structure. Under Owen's framework, the wing of a bird and the arm of a human are homologous because they are the same organ modified; the wing of a bird and the wing of an insect are analogous because they serve the same function (flight) but are built from entirely different structures and have no shared structural ancestry.1 Owen was not an evolutionist; he attributed homology to an underlying ideal archetype, a divine blueprint from which all vertebrate forms were derived. His anatomical observations were precise, but his explanation was pre-Darwinian.
Darwin transformed the concept. In Chapter 13 of On the Origin of Species, titled "Mutual Affinities of Organic Beings: Morphology: Embryology: Rudimentary Organs," Darwin argued that the pattern Owen had documented was exactly what one would expect if species descended with modification from common ancestors. Homologous structures are similar not because they conform to an abstract archetype, but because they are inherited from a shared ancestor in which the structure first evolved.2 This reformulation made homology an explicitly historical concept: two structures are homologous if and only if they can be traced back to a single structure in the most recent common ancestor of the organisms being compared. Modern biologists retain this definition, though they have extended it beyond morphology to encompass developmental processes, gene sequences, and regulatory networks.4, 5
The evolutionary biologist Gunter Wagner has refined the concept further by proposing that homology, or character identity, is maintained over evolutionary time by the persistence of character identity networks: the gene regulatory networks that enable differential gene expression and give structures their distinct developmental identities. Under this framework, two structures in different species are homologous when the same underlying regulatory network specifies their development, even if downstream gene expression has diverged.4 This mechanistic definition provides a principled way to distinguish homology from superficial resemblance and has become increasingly influential in evolutionary developmental biology.
The vertebrate forelimb
The classic textbook example of homology is the vertebrate forelimb. All tetrapods, the four-limbed land vertebrates and their descendants, possess forelimbs built from the same set of bones arranged in the same basic pattern: a single proximal bone (the humerus) articulating with two bones of the forearm (the radius and ulna), followed by a cluster of small wrist bones (the carpals), a set of palm bones (the metacarpals), and finally the bones of the digits (the phalanges).3, 24 This one-two-many pattern is conserved across species whose forelimbs perform wildly different functions.
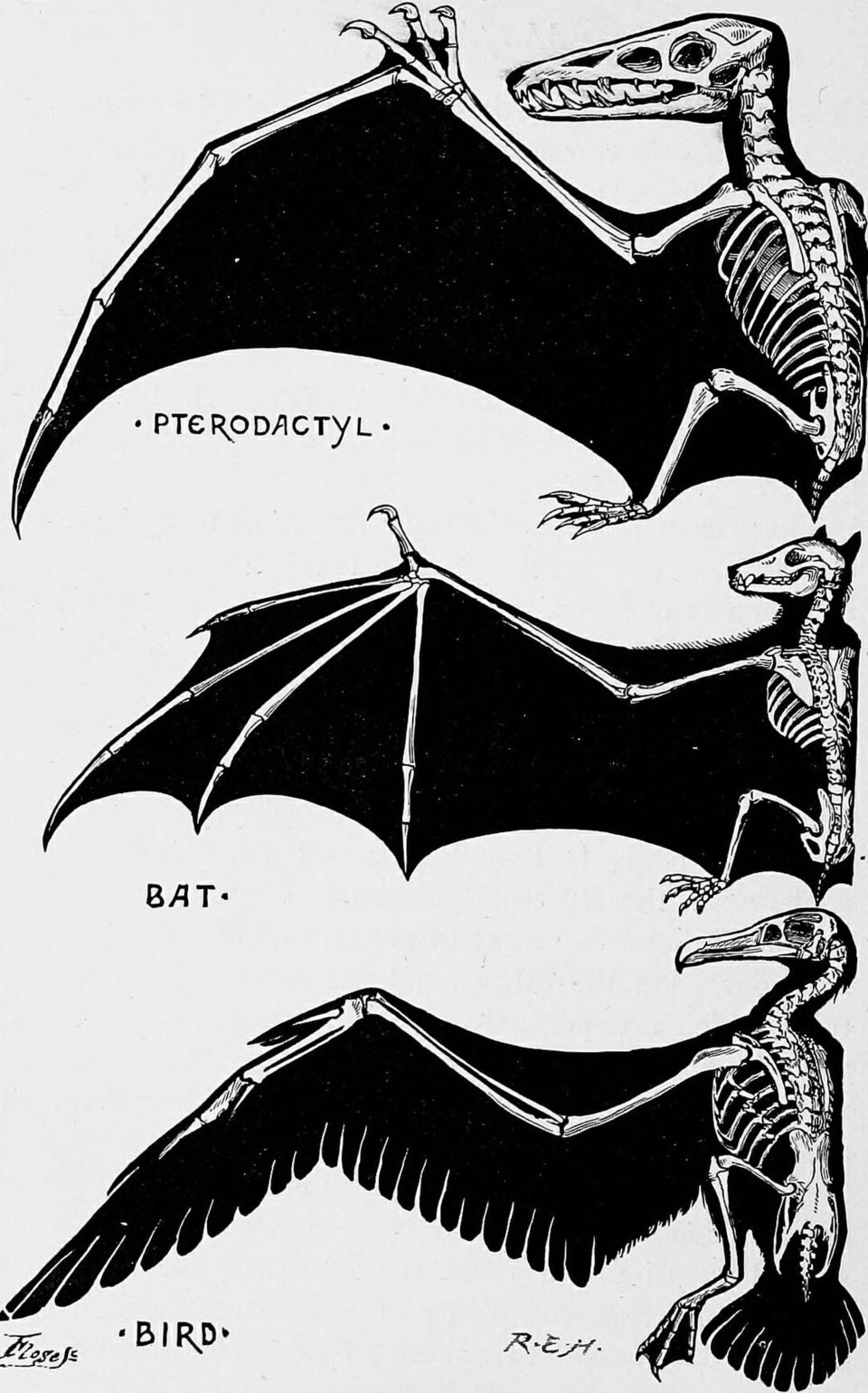

In the human arm, the forelimb is adapted for fine manipulation: the radius and ulna rotate to allow the hand to supinate and pronate, and the fingers are elongated and dexterous. In the bat wing, the same bones are present, but the metacarpals and phalanges of digits two through five are enormously elongated to support a thin membrane of skin used for powered flight. In the whale flipper, the humerus, radius, and ulna are shortened and flattened, the wrist bones are reduced, and the phalanges are multiplied (a condition called hyperphalangy) to form a broad, rigid paddle for swimming. In the horse leg, the ulna is reduced and fused to the radius, the wrist bones are compacted, and the animal stands on a single enlarged digit (the third), its hoof equivalent to a single fingernail, the entire limb elongated into a lever optimized for running.3, 23
If each of these limbs had been designed independently for its particular function, there would be no reason to expect them to share the same underlying skeletal architecture. An engineer designing a wing from scratch would not begin with the blueprint for a grasping hand. But evolution does not design from scratch; it modifies what already exists. The forelimb of every tetrapod is a modification of the forelimb that first evolved in the lobe-finned fish that gave rise to the earliest land vertebrates, approximately 375 million years ago. The discovery of Tiktaalik roseae, a Late Devonian transitional form between lobe-finned fish and early tetrapods, dramatically confirmed this. The pectoral fin of Tiktaalik contains a humerus, radius, and ulna homologous to those of tetrapods, along with a flexible wrist joint capable of supporting the animal's weight, a structure morphologically and functionally intermediate between a fin and a limb.6 Additional transitional forms such as Ventastega curonica have filled further gaps in this sequence, illustrating the step-by-step modification of the ancestral fin into the tetrapod forelimb.21
Homologous bones of the vertebrate forelimb across taxa3, 24
| Bone element | Human (grasping) | Bat (flying) | Whale (swimming) | Horse (running) |
|---|---|---|---|---|
| Humerus | Present, moderate length | Present, short | Present, short and flat | Present, stout |
| Radius & ulna | Separate, rotate | Separate, elongated | Short, flattened | Fused, elongated |
| Carpals | 8 wrist bones | Reduced | Reduced | Compacted |
| Metacarpals | 5, moderate | 4 greatly elongated | 5, within flipper | 1 (third), elongated |
| Phalanges | 5 digits, dexterous | 4 digits, support wing membrane | 5 digits, hyperphalangy | 1 digit (hoof) |
Developmental homologies
Homology is not limited to the finished anatomy of adults. Some of the most striking evidence for common ancestry comes from the study of embryonic development, where structures that will become very different in the adult appear remarkably similar in the embryo. The most widely cited example is the pharyngeal arches, a series of paired, arch-shaped swellings that appear in the throat region of all vertebrate embryos during early development. In fish, these arches develop into the gills and their supporting skeletal elements. In reptiles, birds, and mammals, which have no gills, the same embryonic arches are repurposed during development to form the jaw, the bones of the middle ear, the hyoid bone of the throat, and portions of the larynx.15
The genetic and developmental basis of pharyngeal arch patterning is deeply conserved across vertebrates. The Dlx family of transcription factors is deployed in a nested pattern along the dorsoventral axis of the arches in species as distant as mammals and cartilaginous fishes (skates and sharks), establishing a molecular "code" that specifies the identity of the upper and lower skeletal elements of each arch. Gillis, Modrell, and Baker demonstrated in 2013 that this Dlx code is a primitive feature of the mandibular, hyoid, and gill arches of jawed vertebrates, providing developmental genetic evidence for the classical hypothesis that the vertebrate jaw is serially homologous to the gill arch skeleton.14 The fact that the jaw of a mammal, the gill arch of a shark, and the hyoid bone of a bird are all patterned by the same ancient regulatory system is powerful evidence that these structures derive from a common ancestral arrangement.
Another dramatic developmental homology concerns the limbs of vertebrates and the appendages of arthropods. Despite the enormous morphological difference between a mouse leg and an insect leg, both are patterned along their proximodistal axis by homologous genes. The Distal-less gene in insects and its vertebrate homologues (the Dlx genes) are expressed in the distal (outermost) portions of developing appendages in both lineages, and signaling molecules of the Hedgehog family pattern the anterior-posterior axis of both vertebrate and arthropod limbs.7, 8 Shubin, Tabin, and Carroll coined the term deep homology to describe this phenomenon: the conservation of genetic regulatory circuits across vast evolutionary distances, even when the structures they specify are morphologically dissimilar. Deep homology suggests that the common ancestor of vertebrates and arthropods, which lived more than 500 million years ago, already possessed a genetic toolkit for building appendages, and that this toolkit was independently recruited and elaborated in different lineages.7, 8
Molecular homologies
The advent of molecular biology in the twentieth century revealed that homology extends far below the level of visible anatomy, into the genes and proteins that underlie all of life. Organisms that share common ancestry share homologous DNA sequences, and the degree of sequence similarity between species generally mirrors their phylogenetic relatedness, a pattern first articulated by Zuckerkandl and Pauling in 1965 when they proposed that molecules could serve as "documents of evolutionary history."12
One of the earliest and most influential demonstrations of molecular homology came from the protein cytochrome c, a small electron-transport protein essential to aerobic respiration in virtually all eukaryotes. Human cytochrome c consists of 104 amino acids. The cytochrome c of the rhesus monkey differs from the human sequence at just one amino acid position; that of the dog differs at 11 positions; that of the tuna at 21 positions; and that of baker's yeast at 44 positions. In 1967, Fitch and Margoliash constructed a phylogenetic tree based solely on cytochrome c sequence differences across 20 species. The tree they produced was "remarkably like that constructed in accord with classical zoological comparisons" based on anatomy and the fossil record, providing striking independent confirmation of evolutionary relationships derived from an entirely different class of data.11 This congruence between molecular and morphological phylogenies is precisely what common descent predicts and would be inexplicable if species were unrelated.
Perhaps the most dramatic molecular homologies are found in the Hox genes, a family of transcription factors that specify body-segment identity along the head-to-tail axis of virtually all bilaterian animals. In the fruit fly Drosophila, the Hox genes are arranged in two clusters on a single chromosome, and their linear order along the chromosome corresponds to the order in which they are expressed along the body axis, a property called collinearity. In vertebrates, an ancestral Hox cluster has been duplicated twice during whole-genome duplication events early in vertebrate evolution, producing four paralogous clusters (HoxA, HoxB, HoxC, and HoxD), each containing a subset of up to 13 paralog groups. Despite more than 500 million years of independent evolution, the gene order, sequence, and collinear expression pattern of the Hox clusters are conserved from flies to fish to humans.9, 10
Comparative genomic studies have revealed that the conservation of Hox clusters extends beyond the coding sequences of the genes themselves to the regulatory elements that control their expression. Santini and colleagues aligned the HoxA clusters of species separated by approximately 500 million years of evolution, including tilapia, pufferfish, zebrafish, horn shark, mouse, and human, and identified short regulatory fragments that are nearly identical across all of them, many of which correspond to known binding sites for regulatory proteins.20 This level of conservation is a direct consequence of the critical role these genes play in body-plan specification: mutations that disrupt Hox gene regulation tend to produce catastrophic developmental abnormalities and are thus purged by natural selection, preserving the ancestral regulatory architecture over hundreds of millions of years.10
A further striking example of molecular homology concerns the Pax6 gene and eye development. Despite the enormous morphological difference between the compound eye of a fly and the camera eye of a vertebrate, both are initiated and controlled by homologous Pax6 transcription factors. In 1995, Halder, Callaerts, and Gehring demonstrated that the Drosophila eyeless gene (the fly homologue of Pax6) could induce the formation of ectopic eyes when expressed in other tissues such as the legs, wings, and antennae of the fly. Even more remarkably, the mouse Pax6 gene, when introduced into Drosophila, could trigger the development of fly-type compound eyes, not mouse eyes, demonstrating that the gene's role as a master regulator of eye development has been conserved across a vast evolutionary distance, even though the downstream developmental programs it activates have diverged enormously.13
Vestigial structures as a subtype of homology
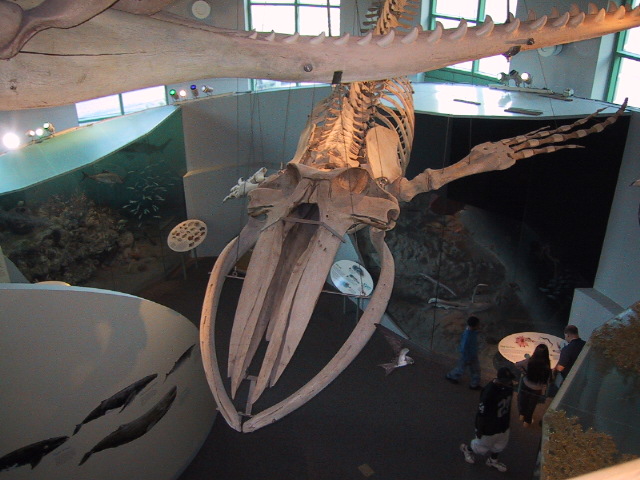

Vestigial structures are a special case of homology in which a feature that served an important function in an ancestor persists in a descendant in a reduced or nonfunctional form. The human coccyx (tailbone) is homologous to the tails of other primates and other mammals; it is the remnant of the ancestral tail, reduced to a small, fused set of vertebrae that no longer support an external tail. Similarly, the tiny, nonfunctional hind-limb bones embedded in the body wall of some whale species are homologous to the fully developed hind limbs of their terrestrial ancestors.3, 23
Vestigial structures make sense only in the context of evolutionary history. A structure that serves no current function, or a greatly diminished one, is difficult to explain as the product of independent design but follows naturally from descent with modification: the structure was inherited from an ancestor in which it was functional, and it has been progressively reduced because natural selection no longer maintains it. The presence of vestigial structures thus provides a particular kind of evidence for common ancestry, one that simultaneously reveals the historical trajectory of a lineage and the constraints that inheritance imposes on evolutionary change.2, 3 Molecular vestigia, such as pseudogenes (genes that have been disabled by mutation and no longer produce functional protein), extend this principle to the genomic level, revealing ancestral gene functions that have been lost over evolutionary time.23
Homology versus homoplasy
Not all similarity between organisms is due to common ancestry. When two species independently evolve similar features in response to similar environmental pressures, the result is called homoplasy (also referred to as convergence, parallelism, or reversal, depending on the specific pattern). The wings of bats and the wings of birds are homologous as forelimbs (both are modified tetrapod forelimbs built from the same bones), but their adaptation for flight evolved independently; bat wings are supported by elongated finger bones with a skin membrane, while bird wings are supported by fused hand bones with feathers. The wings of insects, in contrast, are not homologous to vertebrate forelimbs at all; they are entirely different structures that convergently evolved the function of flight.3, 16
Distinguishing homology from homoplasy is a central challenge in phylogenetic reconstruction. Homologous characters, when correctly identified, provide reliable evidence of evolutionary relationships, whereas homoplastic characters can mislead phylogenetic analysis by grouping unrelated organisms together. Colin Patterson identified three classical tests for distinguishing homology from homoplasy: similarity (detailed structural correspondence), conjunction (the structures do not occur simultaneously in the same organism, ruling out serial repetition), and congruence (the proposed homology is consistent with other characters in a phylogenetic analysis).5 Of these, congruence is the most powerful, because it evaluates a proposed homology against the totality of available evidence: if a character consistently groups the same species together across independent analyses using different data sets, it is almost certainly homologous.5, 22
David Wake's influential 1991 analysis demonstrated that homoplasy is far more common than early systematists had assumed, arising through convergent evolution, parallelism, and evolutionary reversal at rates that can be difficult to detect without rigorous phylogenetic analysis.16 Wake and colleagues later showed that homoplasy itself is informative, because the tendency of certain traits to evolve repeatedly reveals underlying developmental and genetic constraints that channel evolution along limited pathways. Repeated evolution of the same phenotype in related lineages, for instance, often reflects shared developmental biases inherited from a common ancestor, a phenomenon that blurs the boundary between homology and homoplasy and has led to a more nuanced understanding of both concepts.17
Cytochrome c amino acid differences from human sequence11
Homology before and after Darwin
The history of the homology concept illustrates how the same empirical observations can receive fundamentally different explanations depending on the theoretical framework in which they are interpreted. Before Darwin, naturalists such as Owen, Geoffroy Saint-Hilaire, and Goethe recognized the striking structural correspondences among vertebrates, but they attributed these patterns to a transcendental archetype, an ideal form from which all actual organisms were derived as variations. Owen's archetype was explicitly non-evolutionary; it was a Platonic ideal, a plan in the mind of the Creator that imposed a common structural theme on the diversity of life.1, 19
Darwin's theory of descent with modification replaced the archetype with ancestry. The reason all vertebrate forelimbs share the same bones is not that they conform to a metaphysical ideal, but that they inherited those bones from a single ancestral species that possessed them. This shift, from explaining homology by idealist philosophy to explaining it by historical causation, was one of the most consequential intellectual transformations in the history of biology. It converted homology from a descriptive observation into an explanatory principle: common structures exist because of common descent.2, 19
In the twentieth century, the development of cladistic methodology, pioneered by the entomologist Willi Hennig, formalized the relationship between homology and phylogenetic inference. Hennig argued that only shared derived characters (synapomorphies), features inherited from a common ancestor that are unique to a particular clade, provide evidence of evolutionary relationships. Shared primitive characters (symplesiomorphies), which are retained from a more distant ancestor, do not distinguish one group from another. Under this framework, the identification of homologous characters and the reconstruction of phylogenetic trees became inextricably linked: homology is both the evidence for phylogeny and a conclusion confirmed by it.3, 22
The nested hierarchical pattern
One of the most powerful predictions of common descent is that the distribution of homologous characters across species should form a nested hierarchy: a pattern in which groups of organisms defined by shared homologies nest within larger groups defined by more widely shared homologies, and so on, producing a branching, tree-like arrangement. This is precisely the pattern that Linnaeus recognized in the eighteenth century when he organized life into his hierarchical classification of kingdom, phylum, class, order, family, genus, and species, though he did so without an evolutionary explanation.2, 3
Under common descent, the nested hierarchy arises naturally because evolution is a branching process. When a lineage splits, the daughter lineages inherit the homologies of the parent and may subsequently acquire new ones. All mammals share mammary glands, hair, and three middle-ear bones because these features evolved in the common ancestor of mammals. Within mammals, all primates additionally share forward-facing eyes, grasping hands, and nails instead of claws. Within primates, all great apes additionally share the loss of a functional tail, broad chests, and particular dental formulae. Each level of the hierarchy corresponds to a branching event in evolutionary history, and the homologies that define each group are the inherited traces of that history.3, 23
The power of this pattern as evidence for evolution lies in its consistency across independent data sets. The same nested hierarchy emerges whether one classifies organisms by skeletal anatomy, by soft-tissue morphology, by developmental pathways, by protein sequences, or by DNA sequences. Phylogenomic studies that reconstruct evolutionary relationships using thousands of genes simultaneously produce trees that are broadly congruent with those derived from morphological data, confirming that the hierarchical pattern is real and reflects a genuine historical process of descent with modification.22 If species were not related by common ancestry, there would be no reason for homologous characters to distribute themselves in a nested, hierarchical pattern; traits could be distributed randomly, or in overlapping, non-hierarchical arrangements. The fact that they consistently form a tree is among the strongest evidence that the tree of life is real.3, 22, 23
Homology as a unifying concept
Homology stands as one of the central organizing principles of biology. It connects comparative anatomy, embryology, molecular biology, and phylogenetics into a unified framework: the observation that organisms share features because they share ancestors. The anatomical homologies documented by Owen and Darwin in the nineteenth century, the developmental homologies revealed by experimental embryology in the twentieth century, and the molecular homologies uncovered by genomics in the twenty-first century are all manifestations of the same underlying reality: the continuity of life through descent with modification.4, 18
The concept of deep homology, introduced by Shubin, Tabin, and Carroll, has extended this framework to its most sweeping scale. The discovery that the same regulatory genes control the development of eyes in flies and humans (Pax6), pattern limbs in vertebrates and arthropods (Distal-less and Hedgehog family members), and specify body-segment identity across the bilaterians (Hox genes) reveals that the genetic toolkit underlying animal body plans is extraordinarily ancient, predating the divergence of major animal phyla more than 500 million years ago.7, 8 The diversity of animal forms, from insects to fish to mammals, has been generated largely by the modification and redeployment of this shared ancestral toolkit rather than by the invention of entirely new genes. Evolution, as Sean Carroll has put it, is "tinkering with the toolkit," rewiring ancient regulatory circuits to produce new structures from old parts.18
From Owen's careful anatomical dissections to the sequencing of whole genomes, the study of homology has consistently revealed the same message: the extraordinary diversity of life is built on a foundation of shared ancestry. Every vertebrate forelimb, every pharyngeal arch, every Hox gene cluster, and every cytochrome c molecule is a testament to the deep historical connections that unite all living organisms on a single tree of life.4, 8, 22
References
Developmental evidence for serial homology of the vertebrate jaw and gill arch skeleton