Overview
- Horizontal gene transfer — the movement of genetic material between organisms by means other than vertical inheritance from parent to offspring — is a pervasive force in prokaryotic evolution, with comparative genomic analyses indicating that at least 75–81% of genes in sequenced bacterial and archaeal genomes have been laterally transferred at some point in their evolutionary history.
- Three primary mechanisms drive horizontal gene transfer in bacteria: transformation (uptake of free DNA from the environment), transduction (bacteriophage-mediated transfer), and conjugation (direct cell-to-cell transfer via pili or pores), all facilitated by mobile genetic elements including plasmids, transposons, integrons, and phages.
- The prevalence of horizontal gene transfer has fundamentally challenged the concept of a single universal tree of life, leading evolutionary biologists to reconceptualize prokaryotic evolutionary history as a complex web or network of gene sharing rather than a strictly bifurcating tree.
Horizontal gene transfer (HGT), also called lateral gene transfer, is the transmission of genetic material between organisms by mechanisms other than the vertical passage of DNA from parent to offspring. In contrast to the familiar pattern of heredity in which genes flow down through generations along lineages, horizontal transfer moves genes across lineage boundaries — between unrelated species, between different genera, and even between organisms belonging to different domains of life. First recognized through the study of bacterial transformation in the 1920s and 1930s, HGT is now understood to be one of the most important forces shaping prokaryotic genomes, responsible for the rapid spread of antibiotic resistance, the acquisition of new metabolic capabilities, and the emergence of pathogenic strains from non-pathogenic ancestors.1, 3
The prevalence of HGT in bacteria and archaea has fundamentally altered how biologists conceptualize evolutionary history. The traditional model of a single, bifurcating tree of life, in which all organisms are connected by a branching pattern of ancestor-descendant relationships, must be supplemented — and in the prokaryotic realm largely replaced — by a reticulate network or web in which genes have followed many different evolutionary paths through many different organisms, complicating the reconstruction of the tree of life.2, 14 Understanding horizontal gene transfer is therefore essential not only to microbiology and infectious disease but to the broader question of how evolution builds biological complexity and diversity.
Discovery and historical context
The phenomenon of horizontal gene transfer was discovered well before the term was coined, through a series of landmark experiments in bacterial genetics. In 1928, the British bacteriologist Frederick Griffith observed that heat-killed virulent (smooth-coated) pneumococci could transform living non-virulent (rough-coated) pneumococci into virulent forms, a result he attributed to a "transforming principle" of unknown chemical nature.4 Sixteen years later, Oswald Avery, Colin MacLeod, and Maclyn McCarty identified this transforming principle as deoxyribonucleic acid (DNA), demonstrating that purified DNA extracted from virulent pneumococci could confer virulence on non-virulent strains and that the transformation was heritable.5 This experiment was among the first to establish DNA as the material of heredity, and it simultaneously provided the first molecular evidence that genetic information could be transferred horizontally between bacterial cells.
A second mechanism of horizontal transfer was uncovered in 1946, when Joshua Lederberg and Edward Tatum demonstrated that strains of Escherichia coli with different nutritional requirements could exchange genetic material through direct cell-to-cell contact, a process they termed conjugation.6 In 1952, Norton Zinder and Lederberg discovered a third mechanism: transduction, in which bacteriophages (viruses that infect bacteria) inadvertently package segments of the host chromosome and deliver them to new bacterial cells upon subsequent infection, as demonstrated in Salmonella.7 Together, these three discoveries — transformation, conjugation, and transduction — established the mechanistic framework for understanding how genes move between bacterial cells, a framework that has been greatly elaborated but not fundamentally altered in the seven decades since.
Mechanisms of transfer
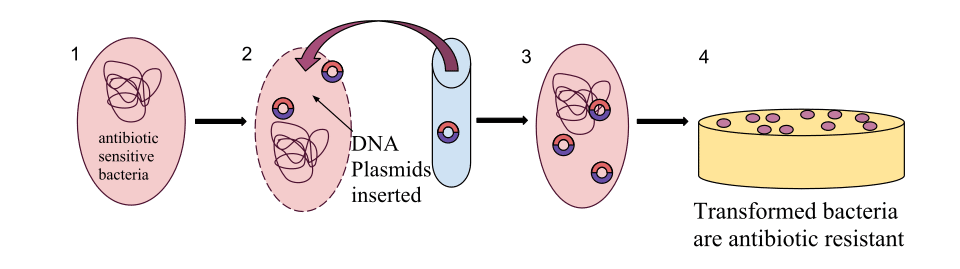
Transformation is the uptake and incorporation of free DNA from the extracellular environment by a competent bacterial cell. DNA released by lysed cells persists transiently in soil, water, and biofilms, and certain bacterial species have evolved regulated genetic programs that render them competent to import this exogenous DNA through specialized membrane-spanning transport machinery. Once inside the cell, the single-stranded DNA can be integrated into the recipient chromosome by homologous recombination if there is sufficient sequence similarity, or it can be maintained if it carries its own replication origin. The range of transformation is broad: because the DNA is naked and the uptake machinery in many species does not discriminate by sequence, transformation can in principle introduce DNA from any source, although the probability of stable integration declines sharply with increasing sequence divergence between donor and recipient.8 Species such as Streptococcus pneumoniae, Haemophilus influenzae, Bacillus subtilis, and Neisseria gonorrhoeae are naturally competent and undergo transformation at high frequencies under appropriate environmental conditions.8
Transduction is the transfer of host DNA from one bacterium to another via a bacteriophage vector. During the lytic cycle of phage replication, the host chromosome is fragmented and occasionally a segment of bacterial DNA is mistakenly packaged into a phage particle instead of phage DNA. When this defective phage infects a new host, it injects the bacterial DNA rather than its own genome, and the recipient cell may incorporate the transferred genes by recombination. In generalized transduction, any region of the donor chromosome may be transferred; in specialized transduction, only genes adjacent to the prophage integration site are mobilized when an integrated temperate phage excises imprecisely.7, 8 Because phages are extraordinarily abundant in natural environments — estimated to outnumber bacteria by a factor of ten or more in most aquatic habitats — transduction is thought to be a quantitatively significant mechanism of gene transfer in nature.9
Conjugation is the direct transfer of DNA from a donor cell to a recipient cell through a physical bridge, typically a pilus or a pore complex encoded by a conjugative plasmid or by a conjugative element integrated into the chromosome (an integrative and conjugative element, or ICE). The process requires cell-to-cell contact and is driven by a type IV secretion system that translocates a single strand of the plasmid or chromosomal DNA into the recipient, where the complementary strand is synthesized. Conjugation is the primary mechanism by which large, self-transmissible plasmids — including those carrying multiple antibiotic resistance genes — spread through bacterial populations.6, 8 Unlike transformation, conjugation can transfer DNA between phylogenetically distant organisms because the process does not depend on homologous recombination; the transferred plasmid can replicate autonomously in the new host if its replication origin is recognized by the recipient's cellular machinery.8, 9
Mobile genetic elements
The agents of horizontal gene transfer are not merely the three mechanisms described above but also the mobile genetic elements that carry genes between chromosomes, plasmids, and phage genomes. These elements — including plasmids, transposons, insertion sequences, integrons, and bacteriophages — have been described as the "agents of open source evolution," assembling, rearranging, and disseminating genetic modules across the prokaryotic world.9
Plasmids are extrachromosomal DNA molecules that replicate independently of the host chromosome. They range in size from a few kilobases to several hundred kilobases and may carry genes conferring antibiotic resistance, heavy metal tolerance, virulence factors, or novel metabolic functions. Conjugative plasmids carry the full set of genes required for their own transfer; mobilizable plasmids lack a complete transfer system but can be transferred if the conjugative machinery is supplied in trans by a co-resident conjugative element.9 The broad-host-range plasmids of incompatibility groups such as IncP, IncQ, and IncW can replicate in a wide diversity of Gram-negative bacteria, enabling gene transfer across genus and even phylum boundaries.8, 9
Transposons are DNA sequences that can move from one position to another within a genome or between different replicons (chromosome, plasmid, or phage) by a cut-and-paste or copy-and-paste mechanism mediated by a transposase enzyme. Composite transposons, flanked by insertion sequences, frequently carry accessory genes such as antibiotic resistance determinants, and their transposition onto conjugative plasmids enables the lateral spread of these genes across species boundaries.9 Integrons are genetic platforms that capture, excise, and rearrange small mobile elements called gene cassettes using a site-specific recombinase called an integrase. First identified in the context of antibiotic resistance, integrons have been found in the chromosomes of hundreds of bacterial species, where they may contain dozens to hundreds of gene cassettes encoding functions ranging from resistance and virulence to metabolic enzymes of unknown function.10 The Vibrio cholerae chromosome, for example, harbours a superintegron containing approximately 216 gene cassettes spanning over 125 kilobases, constituting a substantial fraction of the genome's accessory gene content.10
Prevalence in prokaryotes
Comparative genomic analyses have revealed that horizontal gene transfer is not an occasional curiosity in prokaryotic evolution but a dominant force. In a landmark review, Ochman, Lawrence, and Groisman argued that bacteria have obtained a significant proportion of their genetic diversity through the acquisition of sequences from distantly related organisms, and that HGT produces "extremely dynamic genomes in which substantial amounts of DNA are introduced into and deleted from the chromosome."1 Genomic analyses of sequenced bacterial genomes have consistently identified between 1.6% and 32% of open reading frames in any given genome as probable recent horizontal acquisitions, detected by atypical nucleotide composition, aberrant codon usage, or phylogenetic incongruence with the species tree.1, 13
These single-genome snapshots substantially underestimate the cumulative impact of HGT over evolutionary time, because horizontally transferred genes progressively ameliorate — their nucleotide composition converges with that of the recipient genome through mutational drift — making ancient transfers undetectable by compositional methods. Phylogenetic network analyses that reconstruct the history of individual gene families across large sets of genomes have produced far higher estimates. Dagan, Artzy-Randrup, and Martin analysed 539,723 genes distributed across 181 sequenced prokaryotic genomes and concluded that, on average, at least 81% (with a standard deviation of 15%) of the genes in each genome had been involved in at least one lateral transfer event at some point in their history.15 A subsequent analysis of 329 proteobacterial genomes found that at least 75% of protein families showed evidence of at least one HGT event, with the average number of inferred transfers per gene family increasing from approximately one at the species level to approximately two at the phylum level.25
One of the most striking early examples of massive inter-domain transfer came from the genome of Thermotoga maritima, a hyperthermophilic bacterium whose complete genome sequence, published in 1999, revealed that approximately 24% of its predicted open reading frames had their closest homologues in archaeal species rather than in other bacteria. Many of these archaeal-like genes were clustered in genomic islands with atypical composition, suggesting they had been acquired en bloc through lateral transfer from thermophilic archaea sharing the same extreme environment.16
Estimated prevalence of horizontally transferred genes in selected prokaryotic genomes1, 13, 15, 16
| Organism | Domain | Recent HGT (compositional methods) | Cumulative HGT (phylogenetic networks) |
|---|---|---|---|
| Escherichia coli K-12 | Bacteria | ~17% | ~81% |
| Thermotoga maritima | Bacteria | ~24% | >80% |
| Bacillus subtilis | Bacteria | ~7% | ~75% |
| Synechocystis sp. | Bacteria | ~16% | ~80% |
| Methanocaldococcus jannaschii | Archaea | ~5% | ~70% |
Antibiotic resistance and pathogenicity islands
The most consequential practical manifestation of horizontal gene transfer is the spread of antibiotic resistance among pathogenic bacteria. Resistance genes — encoding enzymes that degrade antibiotics, efflux pumps that expel them from the cell, or modified targets that are no longer inhibited by them — are frequently carried on mobile genetic elements such as conjugative plasmids, transposons, and integron gene cassettes. A single conjugative plasmid may carry resistance determinants against multiple drug classes, enabling the simultaneous acquisition of multidrug resistance in a single transfer event.11 The clinical impact is enormous: the World Health Organization has identified antibiotic resistance as one of the greatest threats to global health, with an estimated 1.27 million deaths directly attributable to drug-resistant infections in 2019.11
The rapidity with which resistance spreads through bacterial populations is a direct consequence of the efficiency of horizontal gene transfer. Resistance genes that arose in environmental bacteria — many of which produce natural antibiotics and have long possessed the corresponding resistance mechanisms — have been recruited onto mobile elements and disseminated into clinically relevant pathogens over mere decades, a timescale far too short to be explained by de novo mutation alone.11 The sulfonamide resistance gene sul1, for example, is found on class 1 integrons in hundreds of Gram-negative species worldwide, and its global distribution reflects the horizontal dissemination of integron-bearing plasmids rather than independent origins of resistance in each species.10, 11
Pathogenicity islands represent another major category of horizontally acquired genetic material. These are large (10 to 200 kilobases), discrete chromosomal regions that encode virulence factors such as adhesins, toxins, secretion systems, and iron uptake systems, and that differ from the surrounding genome in GC content and codon usage, indicating foreign origin. Pathogenicity islands are typically flanked by direct repeats, associated with tRNA genes (which serve as preferred integration sites), and often carry remnants of mobile genetic elements such as phage integrases or transposase genes.12 The locus of enterocyte effacement (LEE) pathogenicity island of enteropathogenic E. coli and the SPI-1 and SPI-2 pathogenicity islands of Salmonella enterica are well-characterized examples. The horizontal acquisition of pathogenicity islands is one of the principal mechanisms by which non-pathogenic bacterial species become pathogenic, a process that can occur in a single evolutionary step through the integration of a large block of virulence genes.12
Impact on phylogenetics and the tree of life
The recognition that horizontal gene transfer is pervasive in prokaryotes has provoked a fundamental reassessment of the concept of a universal tree of life. The traditional tree, rooted in the ribosomal RNA phylogeny pioneered by Carl Woese and widely accepted from the 1970s onward, depicts the history of life as a single bifurcating pattern in which all organisms are connected by a branching series of ancestor-descendant relationships. In a landmark 1999 paper, W. Ford Doolittle argued that if different genes in a genome have different evolutionary histories — as extensive HGT implies — then no single tree can accurately represent the evolutionary relationships among prokaryotes.2 Instead, the history of prokaryotic life is better represented as a web or network of gene sharing, in which vertical descent and horizontal transfer are both prominent threads.
Dagan and Martin formalized this argument quantitatively, demonstrating that the proportion of genes in a typical prokaryotic genome whose history is congruent with any single reference tree (such as the ribosomal RNA tree) is remarkably small — on the order of one percent or less of the genome. They coined the phrase "the tree of one percent" to capture this finding, arguing that the tree-like signal in prokaryotic genomes is a minority pattern superimposed on a much larger background of reticulate gene transfer.14 Subsequent phylogenomic analyses have confirmed that the degree of phylogenetic incongruence among gene families is high, that different genes support different topologies, and that the fraction of the genome with a strictly tree-like evolutionary history is small in most prokaryotic lineages.15, 25
This does not mean that vertical descent is unimportant. Highly conserved informational genes — those involved in transcription, translation, and DNA replication — tend to be transferred less frequently than operational genes involved in metabolism and transport, a pattern sometimes called the "complexity hypothesis," because the products of informational genes participate in large, co-evolved molecular complexes whose function would be disrupted by the substitution of a divergent homologue.3, 13 The ribosomal RNA gene tree therefore remains a useful, if incomplete, guide to the major clades of prokaryotic diversity. But the overall pattern of prokaryotic genome evolution is far more network-like than tree-like, and evolutionary biologists have increasingly adopted network-based methods — phylogenetic networks, gene-sharing graphs, and reconciliation methods that explicitly model HGT — to reconstruct the evolutionary history of microbial life.22, 23
Proportion of gene families with at least one inferred horizontal transfer event across phylogenetic depths in Proteobacteria25
Horizontal gene transfer in eukaryotes
Although HGT is most prevalent and best characterized in prokaryotes, it also occurs in eukaryotes, albeit at generally lower frequencies and through different mechanisms. The most ancient and consequential form of eukaryotic HGT is endosymbiotic gene transfer (EGT), closely related to the broader process of endosymbiosis — the movement of genes from the genomes of mitochondria and chloroplasts (which originated as bacterial endosymbionts) to the nuclear genome of the host cell. This process has been ongoing since the initial endosymbiotic events and has transferred hundreds to thousands of genes from the organelle genomes to the nucleus over more than a billion years of eukaryotic evolution.18 The result is that a substantial fraction of eukaryotic nuclear genes are ultimately of bacterial origin, inherited not by vertical descent from the proto-eukaryotic ancestor but by horizontal transfer from the endosymbiont.17, 18
Beyond endosymbiotic gene transfer, HGT from bacteria and other organisms to eukaryotic nuclear genomes has been documented in a growing number of lineages. Among the most remarkable cases are the bdelloid rotifers, microscopic freshwater invertebrates that have apparently reproduced asexually for tens of millions of years. Genomic analyses have revealed that bdelloid rotifers harbour an unusually high proportion of genes of non-metazoan origin, concentrated in telomeric regions of their chromosomes. These genes appear to have been acquired from bacteria, fungi, and plants, and some are intact and transcriptionally active, producing functional enzymes when expressed in E. coli.19 The mechanism of acquisition may involve the uptake of environmental DNA during the desiccation-rehydration cycles that bdelloids routinely undergo, during which their cell membranes become transiently permeable.19, 17
In plants, horizontal transfer of mitochondrial genes between distantly related species has been documented at surprisingly high frequency. Bergthorsson and colleagues demonstrated that standard mitochondrial genes encoding ribosomal and respiratory proteins have been transferred horizontally between flowering plant species, likely through direct physical contact between parasitic plants and their hosts or through illegitimate pollination and grafting.20 The plant pathogen Agrobacterium tumefaciens provides a particularly well-understood mechanism of bacterium-to-eukaryote gene transfer: during infection, Agrobacterium delivers a segment of its tumour-inducing (Ti) plasmid — the T-DNA — into the plant cell nucleus, where it integrates into the plant genome and directs the synthesis of opines that nourish the bacterium. This natural genetic engineering system has been co-opted as the primary tool for generating transgenic plants in biotechnology, but it also represents a naturally occurring pathway of cross-kingdom gene transfer that has left detectable signatures in the genomes of some plant species.21
More broadly, functional HGT from bacteria to eukaryotes has been documented in insects (including the transfer of genes from Wolbachia endosymbionts to their arthropod hosts), in nematodes (where cellulase genes of bacterial origin enable plant parasitism), and in fungi.24 Husnik and McCutcheon classified these events into two categories: additive HGT, in which a gene with a new function is gained, and replacing HGT, in which a transferred gene replaces an existing homologue, and noted that the number of well-supported cases of functional HGT from bacteria to eukaryotes has expanded rapidly with the growth of genome sequencing.24
Evolutionary significance
.jpg)
Horizontal gene transfer has emerged as one of the most important processes in microbial evolution, rivalling mutation and recombination in its capacity to generate genetic novelty. Whereas point mutations typically produce incremental modifications of existing genes, and recombination reshuffles alleles within a population, HGT can introduce entirely new genes and gene clusters in a single event, enabling quantum leaps in metabolic capability, ecological range, or pathogenic potential.1, 3 The acquisition of photosynthesis genes by non-photosynthetic bacteria, the transfer of nitrogen fixation genes between distantly related soil organisms, and the spread of antibiotic resistance across genus boundaries are all examples of evolutionary innovations that would be impossible or vanishingly improbable through mutation alone but that have occurred repeatedly through horizontal transfer.1, 22
The evolutionary dynamics of HGT also complicate the concept of the bacterial species. In sexually reproducing eukaryotes, species boundaries are maintained by reproductive isolation: members of different species cannot interbreed to produce fertile offspring. In bacteria, where reproduction is clonal and gene transfer is promiscuous, the boundaries between species are inherently fuzzy. A given bacterial cell may carry a mosaic genome in which different genes have different evolutionary origins, and the gene content of closely related strains of the same named species may vary by 20% or more — a phenomenon termed the pangenome.1, 23 The pangenome concept distinguishes between the core genome (genes shared by all strains of a species) and the accessory genome (genes present in some strains but absent in others), and the accessory genome is largely populated by horizontally acquired genes. For species such as E. coli, the pangenome is far larger than the genome of any individual strain, reflecting the cumulative history of gene acquisition and loss across the species as a whole.1, 22
The recognition that HGT is a central rather than peripheral process in evolution has led to a broader reconceptualization of how biological diversity is generated. Soucy, Huang, and Gogarten described HGT as "building the web of life," emphasizing that the evolutionary history of genes and the evolutionary history of organisms are not the same thing, and that a complete account of evolution must incorporate the lateral movement of genetic information alongside the vertical flow.22 This perspective does not diminish the importance of natural selection, genetic drift, or other established evolutionary mechanisms; rather, it expands the set of processes that generate the raw material — heritable genetic variation — upon which those mechanisms act. In prokaryotes especially, HGT is the primary source of new genes and new functions, and any account of bacterial evolution that neglects it is fundamentally incomplete.1, 3, 23
References
Studies on the chemical nature of the substance inducing transformation of pneumococcal types
Modular networks and cumulative impact of lateral transfer in prokaryote genome evolution
Evidence for lateral gene transfer between Archaea and Bacteria from genome sequence of Thermotoga maritima
Networks of gene sharing among 329 proteobacterial genomes reveal differences in lateral gene transfer frequency at different phylogenetic depths