Overview
- The RNA world hypothesis proposes that self-replicating RNA molecules preceded both DNA and proteins in the history of life, serving simultaneously as genetic material and as catalytic enzymes (ribozymes), thereby resolving the chicken-and-egg paradox of modern molecular biology.
- The discovery that RNA can catalyze chemical reactions, including peptide bond formation in the ribosome, provides the strongest evidence that an RNA-based stage of life preceded the current DNA-and-protein world.
- Laboratory experiments have demonstrated that RNA can be evolved to perform new catalytic functions, that RNA enzymes can replicate themselves, and that RNA nucleotides can be synthesized under plausible prebiotic conditions, though significant challenges remain in explaining how the first RNA molecules arose on the early Earth.
Modern life presents a fundamental molecular paradox. DNA stores genetic information but cannot replicate without protein enzymes; proteins perform the catalytic work of the cell but cannot be produced without the instructions encoded in DNA. This chicken-and-egg problem was long recognised as one of the deepest obstacles to understanding how life began. The RNA world hypothesis offers an elegant resolution: it proposes that RNA once served as both the genetic material and the principal catalyst of early life, before the emergence of either DNA or proteins.1, 5 In this view, the earliest living systems were composed primarily of RNA molecules capable of storing hereditary information in their nucleotide sequences and of folding into three-dimensional structures that could catalyze chemical reactions, including their own replication.
The term "RNA world" was coined by the molecular biologist Walter Gilbert in a brief 1986 commentary in Nature, in which he argued that the discovery of catalytic RNA made it plausible that RNA alone could have sustained the first self-replicating systems on the early Earth.1 The hypothesis rests on three pillars: the demonstrated catalytic capacity of RNA, the central role of RNA in the most ancient molecular machinery of the cell, and the results of laboratory experiments showing that RNA can evolve new functions under selective pressure. Together, these lines of evidence have made the RNA world hypothesis the leading framework for understanding the transition from prebiotic chemistry to the first living systems, though important questions about the prebiotic origin of RNA itself remain unresolved.5, 15
The discovery of catalytic RNA
Until the early 1980s, the catalytic work of the cell was thought to be the exclusive province of proteins.5
RNA was understood to play important roles in information transfer — as messenger RNA, transfer RNA, and ribosomal RNA — but not as an enzyme. This assumption was overturned by two independent discoveries that revealed RNA's capacity for catalysis.2, 3
In 1982, Thomas Cech and colleagues at the University of Colorado reported that the ribosomal RNA precursor of the ciliate Tetrahymena thermophila contained an intervening sequence (intron) that could excise itself from the precursor transcript and ligate the flanking exons without the assistance of any protein. The reaction required only monovalent and divalent metal cations and a free guanosine nucleoside as a cofactor.2 This was the first demonstration that RNA could catalyze a specific chemical reaction — the precise cleavage and re-ligation of phosphodiester bonds — and Cech called such catalytically active RNA molecules ribozymes. The discovery was initially met with scepticism, since the prevailing dogma held that only proteins could function as enzymes, but the results were rapidly confirmed and extended.
The following year, Sidney Altman's laboratory at Yale University demonstrated that the RNA component of ribonuclease P (RNase P), an enzyme responsible for processing precursor transfer RNA molecules in Escherichia coli, could catalyze the cleavage of its substrate in vitro in the absence of the protein subunit, provided that the concentration of magnesium ions was sufficiently elevated.3 Unlike the Tetrahymena intron, which acts on itself (and is therefore technically an autocatalyst rather than a true enzyme), the RNA subunit of RNase P acts on separate substrate molecules in a catalytic cycle with genuine enzymatic turnover. Cech and Altman shared the 1989 Nobel Prize in Chemistry for their independent discoveries that RNA could serve as a biological catalyst.2, 3
The implications for the origin of life were immediately recognised. In 1986, Cech proposed a detailed model in which an RNA molecule could serve as a template-directed RNA polymerase, catalyzing the replication of other RNA molecules and potentially of itself.4 In the same year, Gilbert published his influential commentary arguing that an entire world of self-replicating RNA organisms could have preceded the evolution of DNA and protein, and that the present-day reliance of life on DNA for information storage and protein for catalysis represented a later, more efficient arrangement that displaced the ancestral RNA-based system.1
Evidence from modern biology
The strongest evidence for a historical RNA world comes not from laboratory reconstructions of ancient chemistry but from the molecular architecture of modern cells, which retains what appear to be unmistakable vestiges of an earlier RNA-dominated era.5
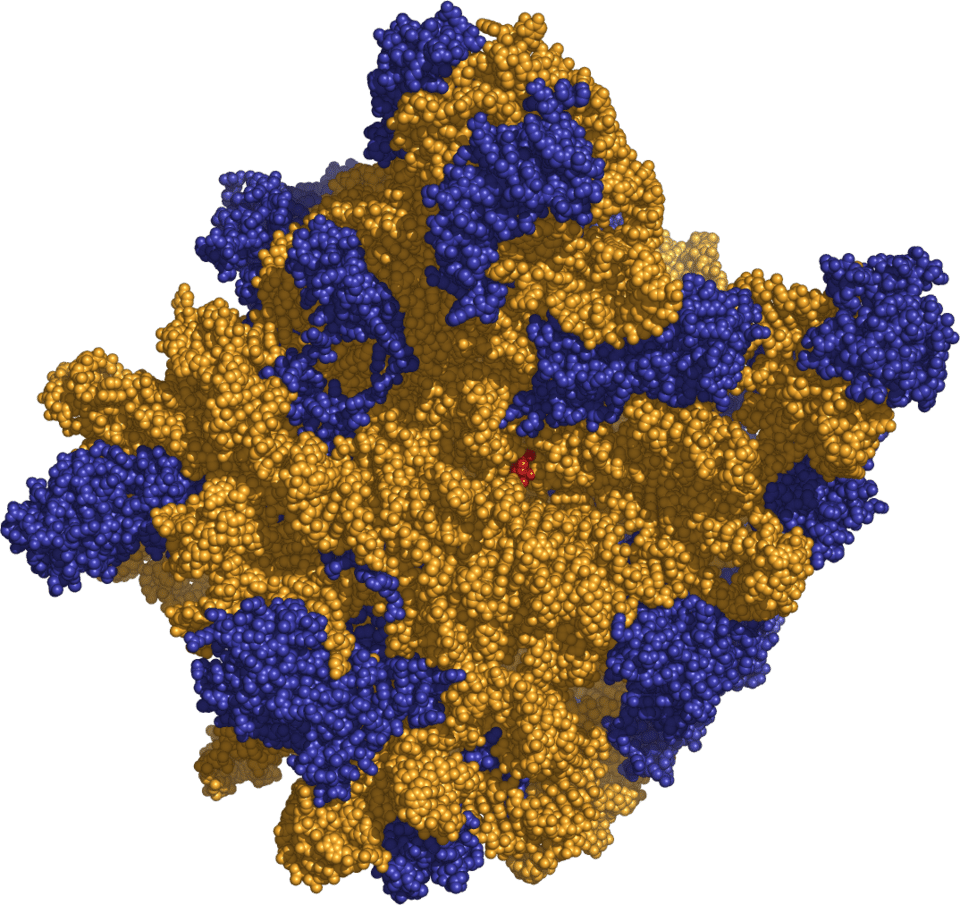
The most compelling of these vestiges is the ribosome, the molecular machine responsible for translating genetic information into protein in all living organisms. When the atomic structure of the large ribosomal subunit was determined at 2.4 angstrom resolution by Steitz, Ban, and colleagues in 2000, it revealed that the catalytic core of the ribosome — the peptidyl transferase centre where peptide bonds are formed — is composed entirely of RNA.6 No protein side chain comes within approximately 18 angstroms of the site of peptide bond formation; the reaction is catalyzed exclusively by ribosomal RNA.7 The ribosome is, at its heart, a ribozyme. This discovery was recognised by the 2009 Nobel Prize in Chemistry, awarded to Venkatraman Ramakrishnan, Thomas Steitz, and Ada Yonath for their structural studies of the ribosome.19 The fact that the universal mechanism of protein synthesis is catalyzed by RNA, not protein, strongly suggests that the ribosome originated in an RNA world and that protein synthesis itself is an invention of RNA-based life.5, 7
A second line of evidence comes from the nucleotide coenzymes that participate in metabolism across all domains of life. Many of the most ancient and essential coenzymes — including adenosine triphosphate (ATP), nicotinamide adenine dinucleotide (NAD), flavin adenine dinucleotide (FAD), and coenzyme A (CoA) — are nucleotides or contain nucleotide moieties that appear to serve no catalytic purpose in their current context. In 1976, Harold White proposed that these nucleotide coenzymes are molecular fossils of an earlier metabolic state in which nucleic acid enzymes performed catalytic functions that were later assumed by proteins, with the nucleotide portions of the coenzymes representing vestiges of the original ribozyme active sites.8 The ubiquity of nucleotide-based cofactors in the most fundamental metabolic pathways across all life is difficult to explain unless these molecules originated in a world where RNA was the dominant macromolecule.5, 8
Additional biological evidence comes from viroids, small circular RNA molecules of 246 to 401 nucleotides that infect plants and replicate autonomously without encoding any protein.17 Viroids are the smallest known agents of infectious disease and consist of nothing more than naked, single-stranded RNA. Their existence demonstrates that RNA alone can contain sufficient information to direct its own replication, albeit by co-opting host cell RNA polymerases. Viroids have been cited as possible molecular relics of an RNA world, or at the very least as demonstrations that autonomous, protein-free RNA replication is biologically feasible.5, 17
Laboratory RNA evolution
A critical prediction of the RNA world hypothesis is that RNA molecules should be capable of evolving new functions under selective pressure, just as protein enzymes do in living organisms. Beginning in 1990, a series of landmark experiments confirmed this prediction through the development of in vitro selection, a technique that subjects large populations of random-sequence RNA molecules to rounds of selection and amplification, mimicking natural selection in a test tube.9, 10

Two foundational methods were published independently in 1990. Tuerk and Gold developed SELEX (Systematic Evolution of Ligands by Exponential Enrichment), in which pools of random RNA sequences were screened for the ability to bind bacteriophage T4 DNA polymerase, with functional sequences amplified by reverse transcription and PCR to enrich the population over successive rounds.9 Simultaneously, Ellington and Szostak demonstrated that RNA molecules capable of binding specific small-molecule ligands — which they named aptamers — could be isolated from pools of approximately 1013 random-sequence RNA molecules, finding that roughly one in 1010 random sequences folded into a structure capable of specific molecular recognition.10
In 1993, Bartel and Szostak extended in vitro selection from binding to catalysis. Starting with a pool of approximately 1015 random RNA sequences, each 220 nucleotides in length, they isolated ribozymes capable of catalyzing the ligation of two RNA molecules aligned on a template — a reaction chemically analogous to the polymerization step performed by modern RNA polymerases. The best ribozymes accelerated the ligation rate by a factor of seven million over the uncatalyzed reaction.11 This experiment demonstrated that catalytic RNA molecules with polymerase-like activity could emerge from random sequences without any intelligent design, providing direct support for the plausibility of an RNA world origin.
Subsequent work progressively improved RNA polymerase ribozymes. In 2011, Wochner and colleagues evolved an RNA polymerase ribozyme capable of synthesizing RNA sequences up to 95 nucleotides in length on an RNA template, including the accurate synthesis of an enzymatically active hammerhead ribozyme — demonstrating the RNA-catalyzed synthesis of a functional RNA molecule from a template for the first time.18 By 2016, Horning and Joyce had evolved an RNA polymerase ribozyme capable of amplifying short RNA sequences in a protein-free form of the polymerase chain reaction and synthesizing structured functional RNAs including aptamers and ribozymes.20 In 2020, Tjhung and colleagues achieved another milestone by evolving a class I RNA polymerase ribozyme that could synthesize its own evolutionary ancestor — an RNA ligase ribozyme — in the form of three fragments that assembled to give a functional complex, as well as the complementary strands of each fragment.25
Perhaps the most striking demonstration came in 2009, when Lincoln and Joyce constructed a pair of RNA enzymes that catalyze each other's synthesis from oligonucleotide substrates. This cross-replicating system underwent self-sustained exponential amplification in the absence of any protein, with a doubling time of approximately one hour, and the amplification could be continued indefinitely.12 When populations of variant cross-replicating enzymes were allowed to compete for a common substrate pool, recombinant replicators arose and grew to dominate the population — an open-ended Darwinian evolution of RNA molecules occurring entirely without proteins or DNA.12
Prebiotic RNA synthesis
While the evidence that RNA can store information, catalyze reactions, and evolve new functions is now overwhelming, the question of how the first RNA molecules arose on the prebiotic Earth remains one of the most challenging problems in origin-of-life research. RNA nucleotides are chemically complex molecules, each composed of three components — a nitrogenous base (a purine or pyrimidine), a five-carbon sugar (ribose), and a phosphate group — and their abiotic synthesis under plausible early Earth conditions was long considered implausibly difficult.14
The central obstacle was what Leslie Orgel and others called the problem of prebiotic nucleotide synthesis. Classical approaches attempted to synthesize nucleotides by first forming the sugar and the base separately and then joining them, but the condensation of a free base with free ribose to form a nucleoside proceeds in very low yield and produces a complex mixture of stereoisomers and regioisomers, most of which are biologically irrelevant.14 Additionally, ribose itself is chemically unstable and is only a minor product of the formose reaction, the most commonly invoked prebiotic pathway for sugar synthesis. This cluster of difficulties was collectively termed the ribose problem.14, 15
A breakthrough came in 2009, when Matthew Powner, Béatrice Gerland, and John Sutherland at the University of Manchester demonstrated a plausible prebiotic synthesis of activated pyrimidine ribonucleotides that bypassed the classical approach entirely. Rather than attempting to join pre-formed sugar and base components, their synthetic route proceeded through a series of intermediates — including 2-aminooxazole and anhydronucleoside compounds — in which the sugar and base were assembled together in a stepwise fashion, never existing as free, separate molecules.13 The reactions required only simple, prebiotically plausible starting materials (cyanamide, cyanoacetylene, glycolaldehyde, glyceraldehyde, and inorganic phosphate) and proceeded in water under mild conditions with ultraviolet light providing a key selectivity step. This work was widely regarded as a major advance because it demonstrated, for the first time, that the chemical building blocks of RNA could plausibly have formed on the early Earth without enzymatic assistance.13, 15
In 2015, Sutherland's group extended this approach dramatically by demonstrating that precursors of ribonucleotides, amino acids, and lipids could all be derived from a common network of reactions driven by hydrogen cyanide and its derivatives under the action of ultraviolet light and hydrogen sulfide — a scenario they termed cyanosulfidic protometabolism.24 This result suggested that the building blocks of RNA, proteins, and cell membranes need not have arisen independently but could have emerged simultaneously from the same simple geochemistry, potentially alleviating one of the central difficulties of the RNA world hypothesis by placing RNA synthesis within a broader, more plausible prebiotic chemical context.24
The environmental settings proposed for prebiotic RNA synthesis include warm, shallow ponds subject to wet-dry cycling (which concentrates reactants through evaporation), volcanic hot springs, and impact-heated environments where meteoritic delivery of reactive molecules could have supplemented terrestrial synthesis.15 Ice has also been proposed as a surprisingly effective medium for prebiotic RNA chemistry: the eutectic phase in ice concentrates solutes by several orders of magnitude and stabilises RNA against hydrolytic degradation, and RNA polymerase ribozymes have been shown to function with enhanced fidelity and processivity in frozen conditions.23
Challenges and alternatives
Despite its explanatory power, the RNA world hypothesis faces several unresolved challenges that have prompted the development of alternative and complementary proposals.14, 15
The most fundamental challenge is the chemical fragility of RNA itself. The 2′-hydroxyl group on ribose renders RNA susceptible to hydrolysis of its phosphodiester backbone, particularly under the warm, alkaline conditions that are often invoked in prebiotic scenarios. The half-life of an RNA phosphodiester bond at neutral pH and 25°C is on the order of several years, but at higher temperatures and higher pH values relevant to some proposed prebiotic environments, degradation can be far more rapid.14 This instability raises the question of whether RNA molecules could have accumulated to sufficient concentrations and lengths to sustain a self-replicating system before being destroyed by hydrolysis.
Template-directed replication of RNA without enzymes also presents difficulties. Non-enzymatic copying of RNA templates using activated nucleotides tends to be slow, error-prone, and strongly sequence-dependent, with certain template sequences being copied far more efficiently than others. The faithful copying of cytosine-rich templates is particularly problematic because of the tendency of the product strand to form stable secondary structures that block further extension.14, 15 Additionally, the strand separation problem — how to denature a double-stranded RNA product to allow the next round of replication — remains largely unsolved in a prebiotic context without the sophisticated helicase enzymes available to modern cells.
These difficulties have led some researchers to propose that RNA was preceded by one or more simpler genetic polymers in a pre-RNA world. Candidate molecules include peptide nucleic acid (PNA), in which the sugar-phosphate backbone is replaced by a polyamide backbone; threose nucleic acid (TNA), which uses a simpler four-carbon sugar; and glycol nucleic acid (GNA), which has an acyclic backbone. These alternative polymers are generally more chemically stable than RNA and easier to synthesize abiotically, and they can form base-paired duplexes with each other and with RNA, potentially allowing a gradual transition from a simpler polymer to RNA over time.22
An entirely different approach to the origin of life bypasses genetics-first scenarios altogether. The metabolism-first hypothesis proposes that self-sustaining networks of chemical reactions arose before any information-carrying polymer. The most developed version of this idea is Günter Wächtershäuser's iron-sulfur world hypothesis, which proposes that the first metabolic cycles were driven by the chemical energy released during the formation of pyrite (FeS2) from iron monosulfide and hydrogen sulfide on the surfaces of mineral particles in deep-sea hydrothermal environments.16 In this model, organic molecules were synthesised and retained on positively charged mineral surfaces, and an autocatalytic version of the reductive citric acid cycle preceded the emergence of any genetic system. The metabolism-first and RNA world hypotheses are not necessarily mutually exclusive; it is possible that simple metabolic chemistry generated the building blocks from which the first RNA molecules were assembled.5, 16
The transition to DNA and proteins
If the RNA world hypothesis is correct, then the contemporary molecular architecture of life — in which DNA serves as the genetic material, proteins perform most catalysis, and RNA acts primarily as an intermediary — must have evolved from an ancestral state in which RNA performed all three functions. Understanding the selective pressures and molecular mechanisms behind this transition is a central challenge of the hypothesis.5, 15
The replacement of RNA by DNA as the primary repository of genetic information likely conferred a significant selective advantage because DNA is substantially more chemically stable than RNA. The absence of the 2′-hydroxyl group on deoxyribose eliminates the intramolecular cleavage reaction that makes RNA susceptible to hydrolysis, and the use of thymine in place of uracil protects against the mutational consequences of cytosine deamination (which produces uracil in DNA, a lesion that can be recognised and repaired, but which would be indistinguishable from a legitimate base in an RNA genome).5 The synthesis of deoxyribonucleotides from ribonucleotides by ribonucleotide reductases, enzymes found in all domains of life, may preserve the ancestral pathway by which DNA building blocks were first produced from the RNA world's nucleotide pool.15
The emergence of coded protein synthesis was arguably the most consequential innovation in the history of life. The ribosome's RNA-catalyzed mechanism of peptide bond formation strongly suggests that the translation apparatus originated within the RNA world, and that the earliest proteins were synthesised by ribozymes.7, 19 Once even rudimentary proteins could be produced under RNA direction, natural selection would have rapidly favoured those RNA molecules whose encoded proteins enhanced the replication or survival of the RNA system. Proteins, with their twenty chemically diverse amino acid side chains, are far more versatile catalysts than RNA, and their progressive recruitment to perform catalytic functions previously carried out by ribozymes would have gradually transformed the RNA world into the ribonucleoprotein world that characterises all modern life.5, 21
The genetic code itself may be an artefact of this transition. The assignment of codons to amino acids likely began as a set of direct chemical interactions between RNA sequences and amino acids — a hypothesis supported by the observation that some aptamers selected in vitro to bind specific amino acids contain codons or anticodons for those amino acids in their binding sites — and was subsequently frozen in place by the interdependence of the translational machinery.5 The universal conservation of the genetic code across all life on Earth is consistent with its origin in a single ancestral RNA-based lineage.
Key milestones in RNA world research
The development of the RNA world hypothesis has proceeded through a series of experimental and theoretical advances spanning more than four decades, from the first discovery of catalytic RNA to the demonstration of self-sustaining RNA replication systems.
Major discoveries and experiments supporting the RNA world hypothesis1, 2, 3, 5, 12, 13
| Year | Discovery | Researchers | Significance |
|---|---|---|---|
| 1976 | Nucleotide coenzymes as molecular fossils | White | Proposed that coenzymes are vestiges of an RNA-based metabolism |
| 1982 | Self-splicing RNA intron in Tetrahymena | Cech et al. | First demonstration that RNA can catalyze a chemical reaction |
| 1983 | RNase P RNA catalyzes tRNA processing | Guerrier-Takada et al. | First true RNA enzyme with catalytic turnover |
| 1986 | "RNA world" hypothesis articulated | Gilbert | Coined the term and proposed RNA preceded DNA and protein |
| 1990 | SELEX and in vitro RNA selection | Tuerk & Gold; Ellington & Szostak | Demonstrated RNA can evolve new binding and functional capacities |
| 1993 | RNA ligase ribozyme from random sequences | Bartel & Szostak | Catalytic RNA evolved de novo; 7-million-fold rate enhancement |
| 2000 | Ribosome structure at atomic resolution | Ban et al.; Nissen et al. | Proved the ribosome is a ribozyme; peptide bond synthesis is RNA-catalyzed |
| 2009 | Prebiotic pyrimidine nucleotide synthesis | Powner, Gerland & Sutherland | First plausible abiotic route to RNA building blocks |
| 2009 | Self-sustained cross-replicating RNA enzymes | Lincoln & Joyce | Indefinite exponential RNA replication without proteins |
| 2015 | Cyanosulfidic protometabolism | Patel et al. | Common prebiotic origins for RNA, protein, and lipid precursors |
| 2016 | RNA polymerase ribozyme amplifies RNA | Horning & Joyce | Protein-free RNA amplification of functional RNA molecules |
| 2020 | Ribozyme synthesizes its own ancestor | Tjhung et al. | RNA polymerase ribozyme copies an RNA ligase from which it evolved |
The trajectory of these discoveries illustrates a progressive strengthening of the RNA world hypothesis over time. The initial theoretical proposal by Gilbert in 1986 was grounded in the newly discovered catalytic abilities of RNA, but the hypothesis could have been falsified at many points: if the ribosome's catalytic centre had turned out to be protein-based, if random RNA sequences had proved incapable of evolving new catalytic functions, or if no plausible prebiotic route to RNA nucleotides could be found. Instead, each of these critical tests has yielded results consistent with the hypothesis, lending it increasing empirical support even as significant gaps — particularly regarding the abiotic origin of the first functional RNA polymers — remain to be closed.5, 15
Current status and ongoing research
The RNA world hypothesis occupies a central position in contemporary origin-of-life research. It is not a complete account of how life began — no hypothesis yet provides such an account — but it offers the most coherent framework for understanding the transition from prebiotic chemistry to the first systems capable of Darwinian evolution. The hypothesis is supported by the catalytic properties of RNA, the RNA-based mechanism of the ribosome, the molecular fossil record of nucleotide coenzymes, the demonstrated evolvability of RNA in the laboratory, and the increasing success of prebiotic chemists in synthesising RNA components under plausible early Earth conditions.5, 13, 15
Active areas of research include the continuing improvement of RNA polymerase ribozymes toward the goal of a fully self-replicating RNA system, the development of more complete prebiotic synthesis routes for both pyrimidine and purine ribonucleotides, the investigation of mineral surfaces, lipid vesicles, and ice as environments that could have concentrated and protected early RNA molecules, and the exploration of how simple protocells — compartments bounded by fatty acid membranes and containing replicating RNA — could have provided the physical basis for natural selection at the cellular level.20, 21, 23, 25
The ultimate goal of this research programme, as articulated by Szostak, Bartel, and Luisi, is the laboratory construction of a synthetic protocell: a self-replicating vesicle containing a self-replicating RNA genome, capable of open-ended Darwinian evolution.21 Such a system would not merely support the RNA world hypothesis but would constitute a demonstration, in principle, of how life could emerge from chemistry. While that goal has not yet been achieved, the progress of the past four decades — from the first discovery that RNA could catalyze a single reaction to the construction of self-sustaining RNA replication systems and the identification of plausible prebiotic pathways to RNA's building blocks — suggests that the remaining obstacles, though formidable, are not insurmountable.5, 12, 15, 24
References
Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena
Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase
Synthesis of activated pyrimidine ribonucleotides in prebiotically plausible conditions
Common origins of RNA, protein and lipid precursors in a cyanosulfidic protometabolism