Overview
- Selfish genetic elements are stretches of DNA that enhance their own transmission to the next generation at the expense of the rest of the genome or the organism, and they include transposable elements, meiotic drivers, segregation distorters, B chromosomes, homing endonucleases, and cytoplasmic endosymbionts such as Wolbachia.
- Transposable elements alone constitute approximately 45 percent of the human genome and over 85 percent of the maize genome, making them the single largest category of DNA in most eukaryotic genomes and demonstrating that a substantial fraction of genomic DNA originated not through organismal adaptation but through the self-propagation of parasitic sequences.
- The evolutionary arms races between selfish elements and host genomes have driven major innovations including new gene regulatory networks, novel genes through molecular domestication, epigenetic silencing systems, sex determination mechanisms, and even the CRISPR-Cas defense system of prokaryotes, revealing that intragenomic conflict is a powerful engine of evolutionary change.
Selfish genetic elements are stretches of DNA that enhance their own transmission to the next generation relative to the rest of the genome, often at a cost to the organism that carries them. Unlike conventional genes, whose evolutionary success is tightly linked to the reproductive success of their host, selfish elements exploit the molecular machinery of the cell to copy, spread, or bias their own inheritance, sometimes producing harmful effects on fertility, viability, or genome integrity in the process. The concept was introduced in two landmark papers published back-to-back in Nature in 1980: Doolittle and Sapienza argued that natural selection operating within genomes would inevitably produce DNA sequences whose only "function" is their own survival, and Orgel and Crick independently proposed that much of the repetitive DNA in eukaryotic genomes is best understood as a parasitic entity rather than as an adaptation of the organism.1, 2 In the decades since, research has revealed that selfish genetic elements are ubiquitous features of virtually all genomes, from bacteria to humans, and that the evolutionary arms races they provoke between parasitic DNA and host defenses have been a major engine of genomic innovation.3, 4
Selfish elements take many forms. Transposable elements copy themselves throughout the genome. Meiotic drivers distort the segregation of chromosomes during gamete formation to ensure their own preferential transmission. B chromosomes are dispensable extra chromosomes that accumulate through non-Mendelian inheritance mechanisms. Homing endonucleases cut their way into new genomic locations. Cytoplasmic elements such as the bacterium Wolbachia manipulate host reproduction to favor infected females. What unites these diverse entities is a common evolutionary logic: they persist and spread not because they benefit the organism but because they have evolved mechanisms to promote their own replication or transmission.3, 4, 20
Conceptual foundations
The theoretical framework for understanding selfish genetic elements emerged from the gene-centred view of evolution articulated by George Williams and Richard Dawkins, which holds that the gene, not the individual organism, is the fundamental unit of natural selection. Under this view, a gene's evolutionary success is measured by its frequency in the gene pool, and any mechanism that increases a gene's transmission relative to its allelic competitors will be favoured, even if it reduces the fitness of the organism carrying it. Doolittle and Sapienza formalised this logic for genomic parasites in 1980, arguing that "when a given DNA, or class of DNAs, of unproven phenotypic function can be shown to have evolved a strategy (such as transposition) which ensures its genomic survival, then no other explanation for its existence is necessary."1 Orgel and Crick arrived at the same conclusion independently, proposing that the large quantities of repetitive DNA in eukaryotic genomes — long dismissed as "junk" — were best understood as the accumulated debris of self-replicating parasitic sequences.2
The population genetics of selfish elements was clarified by Hickey in 1982, who demonstrated mathematically that a transposable element capable of increasing its copy number within a genome can spread through a sexually reproducing population even if it imposes a substantial fitness cost on its host, provided that the rate of transposition exceeds the rate at which selection eliminates element-bearing chromosomes.16 This result established a fundamental principle: the dynamics of selfish elements cannot be understood by considering organismal fitness alone. Instead, selection operates simultaneously at two levels — the level of the element, which is selected for increased replication, and the level of the organism, which is selected for reduced transposition and damage. The resulting tension between these two levels of selection creates an evolutionary arms race that has profoundly shaped genome architecture and function.3, 17
Werren proposed a comprehensive classification of selfish genetic elements based on their mechanism of selfish action, distinguishing five broad categories: transposable elements, which increase in copy number within genomes; biased gene converters, which overwrite alternative alleles through molecular mechanisms; meiotic drivers, which manipulate chromosome segregation during gamete formation; postsegregation killers, which destroy competing gametes or offspring that do not inherit the element; and cytoplasmic drivers, which are maternally inherited elements that manipulate host reproduction to favour their own transmission.3 This classification scheme emphasises that selfishness in genetic elements is not a single phenomenon but a family of distinct strategies, each exploiting a different aspect of the cellular machinery of heredity.
Transposable elements
Transposable elements (TEs) are the most abundant and widespread category of selfish genetic elements. These are DNA sequences that can move or copy themselves from one position to another within a genome, and their cumulative activity over evolutionary time has made them the dominant component of most eukaryotic genomes. The human genome, for example, is composed of approximately 45 percent transposable element-derived sequences, a proportion that dwarfs the roughly 1.5 percent that encodes proteins.6 In plants, TE content varies even more dramatically: the compact genome of Arabidopsis thaliana is approximately 14 percent TEs, while the maize genome is over 85 percent transposable element-derived DNA, with much of this consisting of nested layers of retrotransposon insertions.7

Transposable elements were first discovered by the American geneticist Barbara McClintock, who in the 1940s and 1950s identified "controlling elements" in maize — genetic loci she named Activator (Ac) and Dissociation (Ds) — that could change their chromosomal position and alter the expression of nearby genes, producing the mosaic kernel colour patterns that were her experimental system.5 McClintock's discovery was initially met with scepticism, in part because it challenged the prevailing view of the genome as a stable, linear array of genes. Her work was vindicated decades later when molecular biology confirmed that transposable elements are universal features of genomes across all domains of life, and she was awarded the Nobel Prize in Physiology or Medicine in 1983.
TEs are classified into two major classes based on their mechanism of transposition. Class I elements (retrotransposons) transpose through an RNA intermediate: the element is transcribed into RNA, which is then reverse-transcribed into DNA by a reverse transcriptase encoded by the element itself, and the new DNA copy is inserted at a new genomic location. Because the original copy remains in place, retrotransposons amplify through a "copy-and-paste" mechanism, and their numbers can increase rapidly over evolutionary time. Retrotransposons include long interspersed nuclear elements (LINEs), short interspersed nuclear elements (SINEs), and long terminal repeat (LTR) retrotransposons. In the human genome, the LINE-1 (L1) family alone accounts for approximately 17 percent of genomic DNA, with over 500,000 copies, although only 80 to 100 of these retain the capacity for autonomous transposition.6, 7
Class II elements (DNA transposons) move by a "cut-and-paste" mechanism mediated by a transposase enzyme that excises the element from one location and inserts it at another. Because the element is physically relocated rather than copied, DNA transposons do not inherently increase in copy number through transposition. However, they can amplify through replication-dependent mechanisms, such as transposing from a replicated to an unreplicated region of the genome during S phase, and some DNA transposon families have achieved high copy numbers in certain lineages.8 DNA transposons constitute approximately 3 percent of the human genome, a modest proportion compared with retrotransposons, and none are currently active in the human lineage, although they remain active in many other organisms including insects, plants, and fish.6, 8
Transposable element content as a percentage of genome size in selected organisms6, 7
Meiotic drive systems
Meiotic drive refers to the biased transmission of a genetic element through the manipulation of chromosome segregation during gamete formation, so that the driving element is inherited by more than the expected Mendelian 50 percent of offspring. Meiotic drivers are selfish genetic elements that exploit the asymmetries of meiosis or post-meiotic development to ensure their preferential inclusion in functional gametes, often at the expense of the alternative allele, which is destroyed or rendered dysfunctional.11 The phenomenon was first described by Sandler and Novitski in 1957, and it has since been documented in fungi, plants, insects, and mammals.
Meiotic drive systems typically involve a linked complex of genes: a driver locus that encodes the distorting function, a target or responder locus whose allelic state determines sensitivity to the driver, and often enhancer or suppressor loci that modulate the strength of drive. Because the driver and its enhancers benefit from being inherited together, meiotic drive systems are characteristically associated with inversions or other chromosomal rearrangements that suppress recombination, binding the components into a co-adapted supergene complex.11, 4
The evolutionary consequences of meiotic drive are far-reaching. By distorting transmission ratios, drivers can spread rapidly through populations even if they reduce organismal fitness, creating a conflict between the interests of the driving element and the rest of the genome. This conflict selects for suppressors of drive — unlinked genes that restore Mendelian segregation — and for insensitive responder alleles that are resistant to the driver's action. The result is an ongoing arms race between drivers and suppressors that can drive rapid evolutionary change in the genes involved in chromosome segregation, chromatin organisation, and gametogenesis.3, 11
Segregation Distorter and the t-haplotype
The two best-characterised meiotic drive systems are the Segregation Distorter (SD) complex in Drosophila melanogaster and the t-haplotype in the house mouse (Mus musculus). Both operate during male gametogenesis by destroying or incapacitating sperm that do not carry the driving element, ensuring that virtually all functional sperm transmit the driver.

The SD system of Drosophila was discovered in the 1950s and has been the subject of intensive genetic and molecular analysis. SD is a multigenic complex on the second chromosome consisting of three principal loci: Sd (Segregation Distorter), the primary driving gene; E(SD) (Enhancer of SD), which increases the strength of distortion; and Rsp (Responder), the target of drive. The Sd gene is a truncated duplication of the gene RanGAP, which encodes a GTPase-activating protein involved in nucleocytoplasmic transport. The truncated Sd-RanGAP protein retains enzymatic activity but is mislocalised within the cell, and it somehow disrupts the normal histone-to-protamine transition during spermatid maturation in spermatids bearing a sensitive Rsp allele. The Rsp locus is a block of 120-base-pair satellite DNA repeats in the pericentromeric heterochromatin; spermatids with many copies of the repeat (sensitive allele) are destroyed, while spermatids with few or no copies (insensitive allele) are spared. In heterozygous SD/SD+ males, the result is that over 95 percent of functional sperm carry the SD chromosome.10
The mouse t-haplotype is a variant form of approximately 30 to 40 megabases of proximal chromosome 17 that has been maintained in wild mouse populations for over a million years. Like SD, the t-haplotype achieves its selfish transmission through a poison-antidote mechanism operating during spermatogenesis. Multiple "distorter" loci encoded within the t-haplotype produce factors that interfere with sperm motility or flagellar function in all spermatids, while a linked "responder" locus (Tcr, also called Smok1) encodes a rescue factor that protects t-bearing sperm from the distorter's effects. In heterozygous +/t males, t-bearing sperm are rescued while +-bearing sperm are rendered dysfunctional, resulting in transmission ratios of 90 percent or greater in favour of the t-haplotype.12 Four large, non-overlapping inversions on the t-haplotype suppress recombination with the wild-type chromosome, binding the distorter and responder loci into a non-recombining supergene complex and preventing the rescue factor from separating from the driving alleles.4, 12
Despite their transmission advantage, neither SD nor the t-haplotype has gone to fixation in natural populations. Both systems carry fitness costs that limit their spread: homozygous SD/SD Drosophila males are nearly sterile because almost all spermatids carry a sensitive Rsp allele and are destroyed, and homozygous t/t mice typically die during embryonic development due to recessive lethal alleles embedded within the t-haplotype. These balancing forces — the transmission advantage of the heterozygote versus the lethality or sterility of the homozygote — maintain both systems as stable polymorphisms in natural populations at intermediate frequencies.10, 11, 12
B chromosomes
B chromosomes (also called supernumerary chromosomes) are extra chromosomes found in some individuals of a species but not others. They are not required for normal growth and development, they do not pair or recombine with any of the standard (A) chromosomes during meiosis, and they are present in widely varying numbers among individuals, populations, and even cells within the same organism. B chromosomes have been identified in over 2,800 species of plants, animals, and fungi, and they are now understood to be a distinct class of selfish genetic element that persists through non-Mendelian mechanisms of accumulation.13
The selfish nature of B chromosomes lies in their drive mechanisms — the processes by which they achieve transmission rates exceeding the Mendelian expectation, compensating for the fitness costs they typically impose on their carriers. In plants, the most common drive mechanism is nondisjunction at the second pollen mitosis, in which the two copies of the B chromosome fail to separate during the mitotic division that produces the sperm cells within the pollen grain. Both copies move to the generative nucleus that will fertilise the egg cell, effectively doubling the B chromosome number in the next generation. In maize, this drive mechanism results in B chromosome transmission rates of approximately 70 percent through the pollen, well above the 50 percent Mendelian expectation.13 In animals, B chromosome accumulation mechanisms include directed nondisjunction during female meiosis (as in grasshoppers) and premeiotic accumulation through mitotic drive in the germline (as in the parasitoid wasp Nasonia vitripennis).4, 13
Molecular analyses have revealed that B chromosomes are mosaics composed primarily of sequences derived from the standard A chromosomes, including fragments of genes, repetitive sequences, ribosomal DNA, and even organellar DNA. Their sequence composition indicates that B chromosomes originate from A chromosomes, probably through a process involving centromere-containing fragments that acquire drive mechanisms and begin to accumulate independently of the regular genome.13 Despite carrying gene-derived sequences, most B chromosomes appear to be largely transcriptionally inert, consistent with a parasitic element whose presence is tolerated rather than beneficial. At low copy numbers, B chromosomes may be selectively neutral, but at high copy numbers they typically reduce fertility and vigour, creating a dynamic equilibrium between drive and selection that maintains B chromosome frequencies within populations.4, 13
Homing endonucleases and gene drives
Homing endonucleases are a class of selfish genetic element found primarily in the organellar genomes of fungi and in certain bacteriophages. These elements encode a nuclease enzyme that recognises and cleaves a specific DNA sequence of 15 to 30 base pairs, which typically occurs at the precise chromosomal position where the homing endonuclease gene would be inserted if it were present. In a heterozygous cell containing one chromosome with the element and one without, the endonuclease cuts the element-free chromosome at the recognition site, and the cell's DNA repair machinery uses the element-bearing chromosome as a template for homologous recombination repair, copying the endonuclease gene into the break site. The result is that the heterozygote is converted to a homozygote — a process called homing — and the element achieves nearly 100 percent transmission.4, 14
Homing endonuclease genes (HEGs) are the quintessential site-specific selfish elements: they exploit the host's own DNA repair pathway to copy themselves into a defined target sequence. Because the homing process is so efficient, a newly introduced HEG can spread from a single individual through an entire population in a relatively small number of generations, even if it confers no benefit to the organism. Burt recognised this property and proposed in 2003 that synthetic homing endonucleases could be engineered as gene drives — genetic constructs designed to spread through wild populations of target species, potentially enabling the control of disease vectors such as malaria-transmitting mosquitoes or the eradication of invasive species.14
The development of CRISPR-Cas9 genome editing technology has since made the construction of synthetic gene drives practical. A CRISPR-based gene drive encodes both the Cas9 nuclease and a guide RNA targeting a specific genomic site; when the drive construct is present on one chromosome, it cuts the corresponding site on the homologous chromosome, and the break is repaired using the drive-bearing chromosome as a template, converting the heterozygote to a drive homozygote. Laboratory demonstrations have shown that CRISPR gene drives can achieve inheritance rates exceeding 99 percent in yeast and Drosophila, confirming the theoretical predictions made for natural homing endonucleases.14 The technology raises profound ecological and ethical questions, as a released gene drive could potentially spread through and alter an entire wild species, with consequences that may be difficult to predict or reverse.
Cytoplasmic elements and reproductive manipulation
Selfish genetic elements are not confined to nuclear DNA. Cytoplasmically inherited elements — including mitochondria, chloroplasts, and intracellular bacterial endosymbionts — are transmitted exclusively or preferentially through the maternal lineage via the egg cytoplasm. This mode of inheritance creates a conflict between cytoplasmic elements and nuclear genes, because cytoplasmic elements benefit from any manipulation of host reproduction that increases the number or proportion of females (which transmit the cytoplasm), even if such manipulation is costly to male fitness or to the organism as a whole.3, 18
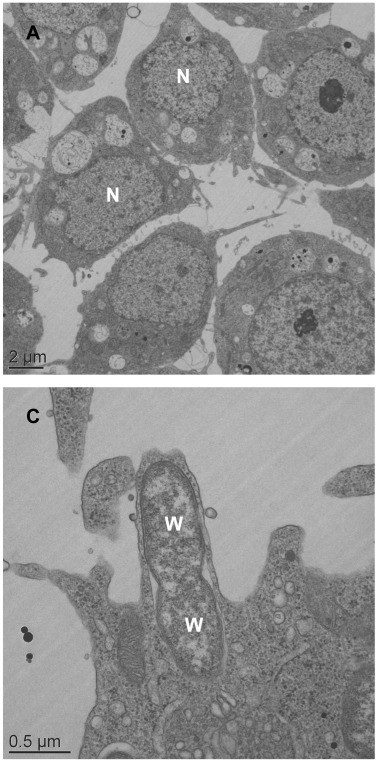
The most extensively studied cytoplasmic selfish element is Wolbachia, an alphaproteobacterium that infects an estimated 40 to 65 percent of all insect species as well as many other arthropods and filarial nematodes. Wolbachia is inherited vertically through the egg cytoplasm and has evolved at least four distinct strategies for manipulating host reproduction to enhance its own transmission: cytoplasmic incompatibility (CI), in which sperm from Wolbachia-infected males are unable to successfully fertilise uninfected eggs, giving infected females a reproductive advantage; male killing, in which infected male embryos are selectively killed, freeing resources for their infected sisters; feminization, in which genetic males are converted to functional females that transmit the bacterium; and parthenogenesis induction, in which unfertilised eggs develop into females, bypassing the need for males entirely.15
Cytoplasmic incompatibility is the most common of these manipulations. In CI, Wolbachia modifies sperm in infected males through a set of CI-inducing factors encoded by bicistronic operons in the Wolbachia genome. When modified sperm fertilise an uninfected egg, the paternal chromosomes fail to condense properly and are lost during early mitotic divisions, resulting in embryonic lethality. When the same modified sperm fertilise an egg from a female infected with the same Wolbachia strain, a rescue factor expressed in the egg cytoplasm neutralises the modification, and development proceeds normally.15, 18 The net effect is that infected females can mate successfully with any male, while uninfected females are unable to mate successfully with infected males, creating a frequency-dependent advantage for infected females that drives Wolbachia to high frequencies in host populations. This reproductive manipulation makes Wolbachia one of the most successful selfish genetic elements in the biosphere, having invaded a substantial fraction of all insect species.3, 15
Another form of cytoplasmic conflict is cytoplasmic male sterility (CMS) in flowering plants, in which mitochondrial genes suppress pollen production, redirecting resources toward seed production and thereby enhancing the transmission of the maternally inherited mitochondrial genome. CMS has been documented in over 150 plant species and is typically counteracted by nuclear restorer-of-fertility genes that suppress the CMS phenotype, reflecting the evolutionary arms race between cytoplasmic and nuclear genomes.4, 18
Host defense mechanisms
The ubiquity of selfish genetic elements has selected for a diverse array of host defense mechanisms that suppress, silence, or eliminate parasitic DNA. These defenses operate at multiple levels, from the chromatin structure of the genome to the post-transcriptional degradation of element-derived transcripts, and they represent some of the most elaborate molecular systems in biology.
Epigenetic silencing is the primary defense against transposable elements in eukaryotic genomes. DNA methylation of cytosine residues, directed by small RNA pathways, represses transcription of transposable elements by converting element-containing regions of the genome into condensed, transcriptionally inactive heterochromatin. In plants, the RNA-directed DNA methylation (RdDM) pathway uses small interfering RNAs (siRNAs) derived from TE transcripts to guide de novo methylation of homologous TE sequences, establishing and maintaining their silenced state across cell divisions. In animals, the piRNA (PIWI-interacting RNA) pathway performs an analogous function in the germline: piRNAs complementary to transposable element sequences guide PIWI-clade Argonaute proteins to TE transcripts, triggering their degradation and directing chromatin modifications that silence TEs at the transcriptional level.7, 9
In prokaryotes, the CRISPR-Cas system functions as an adaptive immune defense against mobile genetic elements, including phages and plasmids. Bacteria incorporate short sequences of invading DNA into CRISPR arrays in their chromosomes, and these stored sequences are transcribed and processed into guide RNAs that direct Cas nucleases to cleave matching foreign DNA upon reinfection. The parallel between CRISPR-Cas in bacteria and the piRNA pathway in animals is striking: both are small RNA-guided defense systems that use sequence information from previous encounters with selfish elements to recognise and destroy current invaders.7
Beyond silencing, host genomes have evolved suppressor genes that counteract specific selfish elements. As noted above, nuclear restorer-of-fertility genes suppress cytoplasmic male sterility in plants, and unlinked suppressor loci can reduce or eliminate the transmission advantage of meiotic drivers such as SD and the t-haplotype. The rapid evolution of these suppressor systems, driven by the fitness costs imposed by selfish elements, contributes to the pattern of evolutionary arms races that characterises the relationship between parasitic DNA and its hosts.3, 11
Selfish elements as engines of innovation
Although selfish genetic elements are parasitic in origin, the evolutionary arms races they provoke have been a surprisingly productive source of biological innovation. In many cases, sequences or functions that originated in selfish elements have been domesticated — co-opted by the host genome for new cellular functions — in a process sometimes called molecular exaptation. The result is that much of the regulatory and structural complexity of modern genomes has its ultimate origin in the activity of selfish DNA.3, 9
One of the most consequential examples of domestication is the vertebrate adaptive immune system. The RAG1 and RAG2 proteins that catalyse V(D)J recombination — the somatic rearrangement of immunoglobulin and T-cell receptor gene segments that generates antibody diversity — are derived from a transposase enzyme encoded by an ancient DNA transposon of the Transib superfamily. The mechanism of V(D)J recombination, including the recognition of recombination signal sequences flanking the gene segments, is structurally and mechanistically homologous to cut-and-paste transposition. The adaptive immune system, one of the most sophisticated molecular systems in vertebrate biology, thus originated from the molecular machinery of a selfish genetic element.8, 9
Transposable elements have also been a major source of new gene regulatory sequences. Because TEs carry their own promoters, enhancers, and transcription factor binding sites, their insertion near host genes can alter gene expression patterns. While most such insertions are neutral or deleterious, some have been co-opted to serve as regulatory elements controlling the expression of nearby genes. A genome-wide analysis found that transposable element-derived sequences contribute a substantial fraction of the regulatory elements active in human cells, including enhancers, promoters, and insulator elements.9 Chuong, Elde, and Feschotte demonstrated that endogenous retroviruses — remnants of ancient retroviral infections that are a type of retrotransposon — have been co-opted to serve as interferon-inducible enhancers in the mammalian innate immune system, showing that selfish elements have contributed directly to the evolution of immune gene regulation.9
The centromeres of eukaryotic chromosomes, the chromosomal regions essential for proper segregation during cell division, are themselves embedded in dense arrays of transposable elements and satellite DNA. There is growing evidence that the rapid evolution of centromeric sequences is driven by centromere drive, a form of meiotic drive in which centromeric variants compete for preferential inclusion in the egg cell during female meiosis (since only one of the four meiotic products becomes the egg in most animals and plants). This competition may explain why centromeric DNA evolves far more rapidly than expected for a functionally essential chromosomal region, and why centromere-associated proteins coevolve with their rapidly changing DNA substrates in a pattern consistent with an arms race.3, 19
Selfish genetic elements have also been implicated in the evolution of sex determination systems. In many organisms, sex-determining loci are located in regions of the genome characterised by suppressed recombination, degenerated gene content, and accumulation of repetitive DNA — precisely the features associated with selfish genetic elements and their consequences. Werren has argued that conflicts between sex-ratio distorters (selfish elements that bias the offspring sex ratio in favour of one sex) and their suppressors can drive turnover of sex-determining mechanisms, potentially explaining the remarkable diversity of sex determination systems observed across the animal kingdom.3
Examples of domesticated selfish genetic elements8, 9
| Domesticated element | Origin | Current function | Organism |
|---|---|---|---|
| RAG1/RAG2 | Transib DNA transposon | V(D)J recombination (adaptive immunity) | Jawed vertebrates |
| Syncytin-1/Syncytin-2 | Endogenous retrovirus envelope genes | Placental cell fusion (syncytiotrophoblast) | Mammals |
| Telomerase | Retrotransposon reverse transcriptase | Chromosome end maintenance | Most eukaryotes |
| CENP-B | pogo-like DNA transposase | Centromere function | Mammals |
| MER41 enhancers | Endogenous retrovirus LTRs | Interferon-inducible immune gene regulation | Primates |
Impact on genome size and structure
The accumulation and deletion of transposable elements is the single most important determinant of genome size variation among eukaryotes. The roughly 200-fold variation in genome size among flowering plants, for example, is driven almost entirely by differences in the abundance of retrotransposons, with large-genome species such as maize and wheat containing enormous quantities of nested TE insertions and small-genome species such as Arabidopsis having experienced extensive TE removal through recombination-based deletion mechanisms.7, 17 Even within mammalian lineages, genome size variation is largely attributable to differences in TE content: the smallest mammalian genomes, found in certain bats, have undergone lineage-specific contraction of TE families, while the larger genomes of some marsupials and rodents reflect ongoing retrotransposon amplification.7
Transposable elements also shape genome structure through the chromosomal rearrangements they catalyse. Recombination between non-allelic copies of the same TE family located at different positions on the same or different chromosomes can generate deletions, duplications, inversions, and translocations. Such TE-mediated rearrangements have been documented as the cause of numerous human genetic diseases, including cases of haemophilia A (caused by an inversion mediated by LINE-1 elements within the factor VIII gene), certain forms of Charcot-Marie-Tooth disease, and a variety of cancers in which somatic transposition or TE-mediated rearrangement disrupts tumour suppressor genes.7, 20
The conflict between selfish elements and their hosts has also left deep marks on the regulatory landscape of the genome. The epigenetic silencing machinery that evolved to repress transposable elements — including DNA methylation, histone modification, and small RNA pathways — has been co-opted for broader roles in gene regulation, X-chromosome inactivation, and genomic imprinting. The evolution of these epigenetic systems may have been driven, at least in part, by the selective pressure imposed by transposable elements, making the defense against selfish DNA a catalyst for the evolution of the complex regulatory architecture that characterises eukaryotic gene expression.9, 17
Evolutionary significance
The study of selfish genetic elements has fundamentally altered the understanding of genome evolution. The traditional view that genomes are streamlined repositories of adaptive information, optimised by natural selection for the benefit of the organism, has been replaced by a more complex picture in which genomes are ecological communities of genetic elements with divergent evolutionary interests. In this view, the genome is not a harmonious whole but an arena of conflict, shaped by the interplay between selfish elements seeking to propagate themselves and host defenses seeking to suppress or domesticate them.3, 4, 20
Agren and Clark have argued that the recognition of selfish genetic elements provides the strongest empirical support for the gene's-eye view of evolution, because selfish elements represent the clearest cases of genes that increase in frequency despite being harmful to the organisms that carry them.20 The existence of such elements demonstrates that natural selection operates at the level of the gene, not only at the level of the organism, and that the evolutionary trajectory of a genome is the resultant of selection pressures acting at multiple levels simultaneously. This multilevel perspective has implications far beyond the study of parasitic DNA, informing the understanding of evolutionary transitions, the origin of multicellularity, the evolution of sex, and the dynamics of intragenomic conflict that accompany every fusion of formerly independent genetic systems.3, 19
The practical implications of selfish genetic elements are equally significant. The development of gene drive technology, inspired directly by the natural biology of homing endonucleases and meiotic drivers, has opened the prospect of engineering entire wild populations for conservation or public health purposes. The understanding of transposable element biology underlies modern tools for insertional mutagenesis, gene tagging, and genome engineering. And the ongoing discovery of new selfish elements and their interactions with host genomes continues to reveal unexpected dimensions of genome biology, from the role of TE-derived regulatory sequences in species-specific gene expression to the contribution of intragenomic conflict to speciation and reproductive isolation.3, 14, 20
References
Site-specific selfish genes as tools for the control and genetic engineering of natural populations