Overview
- All non-African modern human populations carry approximately 1 to 4 percent Neanderthal DNA as a result of interbreeding events that occurred after the out-of-Africa dispersal, while some Melanesian and Aboriginal Australian populations carry an additional 3 to 6 percent Denisovan ancestry.
- Many archaic alleles have been retained by natural selection because they conferred adaptive advantages, including immune system genes from Neanderthals and Denisovans, the EPAS1 altitude-adaptation allele from Denisovans in Tibetans, and variants influencing skin pigmentation and fat metabolism.
- The human genome also contains introgression deserts, regions where archaic DNA has been systematically purged by purifying selection, suggesting that many archaic alleles were deleterious in the hybrid genetic background, and emerging evidence points to even deeper superarchaic introgression into Denisovans themselves.
Archaic introgression is the incorporation of genetic material from one species or population into the gene pool of another through hybridisation and backcrossing. In the context of human evolution, the term refers specifically to the transfer of DNA from archaic hominin lineages — principally Neanderthals and Denisovans — into the genome of anatomically modern Homo sapiens. The landmark sequencing of the Neanderthal genome in 2010 demonstrated that all non-African human populations carry approximately 1 to 4 percent Neanderthal-derived DNA, the legacy of interbreeding events that occurred after modern humans dispersed from Africa.1 Subsequent work revealed an additional contribution from Denisovans, a sister group to Neanderthals known primarily from fragmentary fossils and ancient DNA recovered from Denisova Cave in Siberia, with present-day Melanesian and Aboriginal Australian populations carrying approximately 3 to 6 percent Denisovan ancestry.2, 16 These discoveries fundamentally transformed the understanding of modern human origins, replacing a strict out-of-Africa replacement model with a more complex picture in which gene flow among divergent hominin populations was a recurrent feature of the Late Pleistocene.
Discovery of Neanderthal admixture
The question of whether Neanderthals and modern humans interbred was debated for decades before genomic evidence settled the matter. Early mitochondrial DNA studies had found no trace of Neanderthal lineages in living humans, which was initially interpreted as evidence against interbreeding. However, mitochondrial DNA represents only a single maternal lineage and can be lost through genetic drift even if hybridisation occurred. The decisive evidence came in 2010, when a team led by Svante Paabo at the Max Planck Institute for Evolutionary Anthropology published a draft sequence of the Neanderthal genome, assembled from DNA extracted from three bones found in Vindija Cave, Croatia.1
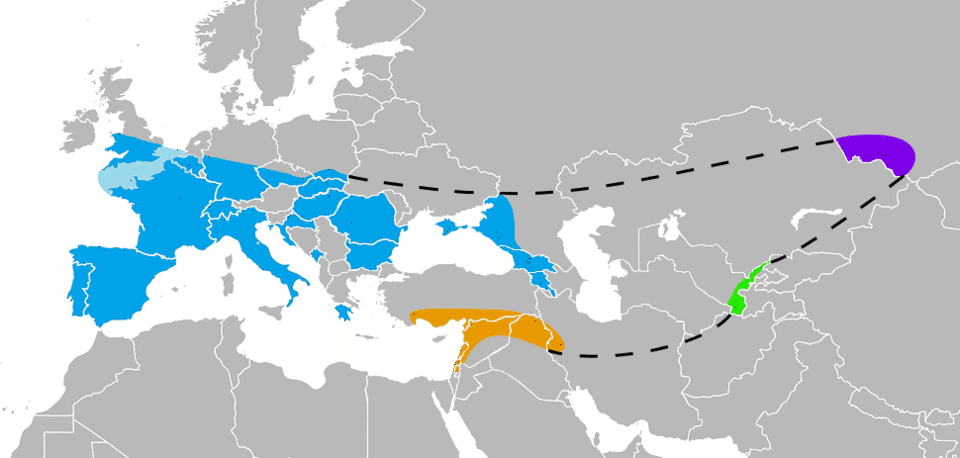
Green and colleagues compared the Neanderthal genome to the genomes of five present-day humans from different continents and found that non-African individuals — from Europe, East Asia, and Oceania — shared significantly more derived alleles with Neanderthals than did West African individuals. This pattern, quantified using the D-statistic (a measure of allele-sharing asymmetry), was most parsimoniously explained by gene flow from Neanderthals into the ancestors of non-Africans after their departure from Africa but before the divergence of European and East Asian lineages.1 The estimated proportion of Neanderthal ancestry in non-African genomes was approximately 1 to 4 percent, a figure that has been refined by subsequent studies using higher-coverage Neanderthal genomes to approximately 1.5 to 2.1 percent in most Eurasian populations.9, 21
The timing of this admixture has been estimated by examining the lengths of Neanderthal-derived haplotype blocks in present-day genomes. Because recombination progressively breaks down introgressed segments over generations, the average length of archaic haplotypes serves as a molecular clock. Analyses of linkage disequilibrium patterns suggest that the primary episode of Neanderthal-modern human interbreeding occurred approximately 47,000 to 65,000 years ago, consistent with the archaeological evidence for overlap between the two populations in the Near East.10
Denisovan introgression
The Denisovans were identified as a distinct archaic hominin lineage in 2010, when DNA extracted from a finger bone discovered in Denisova Cave, Siberia, revealed a mitochondrial genome deeply divergent from both modern humans and Neanderthals.2 Nuclear genome sequencing confirmed that Denisovans were a sister group to Neanderthals, having diverged from them approximately 390,000 to 440,000 years ago, and that they had contributed genetic material to certain modern human populations. Specifically, present-day Melanesians, Aboriginal Australians, and some Southeast Asian populations were found to carry approximately 3 to 6 percent Denisovan-derived DNA, a proportion substantially higher than the Neanderthal contribution.2, 3, 16
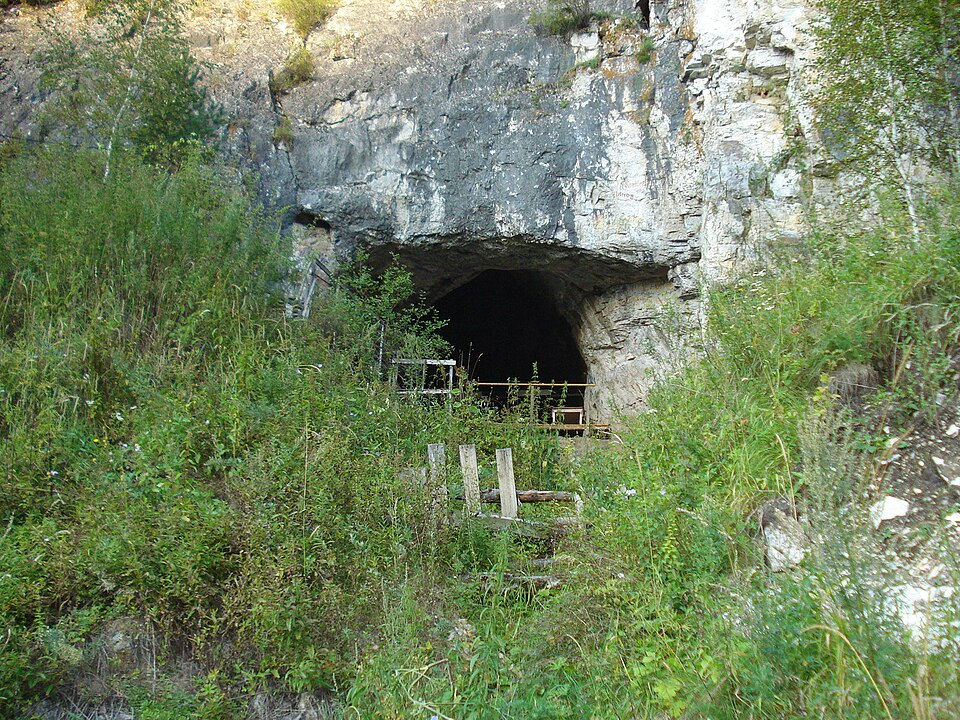

A high-coverage Denisovan genome published in 2012 sharpened these estimates and revealed additional complexity. The Denisovan contribution to mainland East Asian and Native American populations is considerably smaller, on the order of 0.2 percent, suggesting that the primary admixture event involved ancestors of present-day Oceanian populations rather than a broadly shared admixture predating the diversification of non-African populations.3, 5 Further analysis of modern genomes from Island Southeast Asia and Oceania revealed that the Denisovan ancestry in these regions derives from at least two, and possibly three, genetically distinct Denisovan populations, indicating multiple independent admixture events at different times and places.12 This finding implies that Denisovans were not a single, homogeneous population but rather a diverse group that occupied a vast geographic range across Asia.
Adaptive introgression
While most archaic DNA in modern human genomes appears to be selectively neutral or mildly deleterious, a subset of introgressed alleles has been retained at high frequencies because it conferred adaptive advantages in specific environments. This phenomenon, known as adaptive introgression, represents a form of evolutionary shortcut: rather than waiting for beneficial mutations to arise de novo, modern humans acquired pre-adapted genetic variants from archaic populations that had evolved in Eurasia for hundreds of thousands of years.15
One of the most striking examples involves the EPAS1 gene, which encodes a transcription factor in the hypoxia-inducible factor (HIF) pathway that regulates the physiological response to low oxygen levels. A haplotype of EPAS1 found at high frequency in Tibetan populations, which is associated with reduced haemoglobin concentration at high altitude and appears to protect against chronic mountain sickness, was shown to derive from Denisovan introgression. The introgressed haplotype is essentially absent in Han Chinese lowland populations but reaches frequencies exceeding 80 percent in Tibetans, a pattern consistent with strong positive selection at high altitude.6 This allele thus appears to have facilitated the permanent settlement of the Tibetan Plateau, one of the most physiologically challenging environments inhabited by humans.
The innate immune system has been another major beneficiary of archaic introgression. Genes encoding human leukocyte antigen (HLA) class I molecules, which present pathogen-derived peptides to T cells and are critical for adaptive immune responses, show exceptionally high levels of archaic ancestry. Abi-Rached and colleagues demonstrated that certain HLA alleles in modern Eurasian populations were acquired from Neanderthals and Denisovans, and that some of these alleles now represent the majority variants in certain populations. The HLA-B*73 allele, for instance, appears to have been reintroduced into modern humans from Denisovans and is most common in West Asian populations.7 Similarly, Toll-like receptor genes TLR1, TLR6, and TLR10, which encode pattern-recognition receptors of the innate immune system, carry Neanderthal- and Denisovan-derived haplotypes at high frequencies in Eurasian populations, suggesting that archaic introgression provided modern humans with a ready-made toolkit for combating novel pathogens encountered outside Africa.8
Archaic alleles have also been implicated in variation in skin and hair phenotypes, fat storage and metabolism, and response to ultraviolet radiation. Neanderthal-derived variants affecting keratin filament genes are found at elevated frequencies in European and East Asian populations and may have facilitated adaptation to cold, dry climates by modifying skin and hair properties.15, 19 The TBX15/WARS2 locus, which influences body fat distribution and is associated with adaptation to cold environments, carries a Denisovan-derived haplotype at high frequency in Inuit and other high-latitude populations.17
Selected examples of adaptive archaic introgression6, 7, 8, 17
| Gene or region | Archaic source | Function | Population with highest frequency |
|---|---|---|---|
| EPAS1 | Denisovan | Altitude adaptation (HIF pathway) | Tibetans (~80%) |
| HLA class I alleles | Neanderthal & Denisovan | Immune response (antigen presentation) | Eurasians (up to ~50% of HLA-A alleles) |
| TLR1, TLR6, TLR10 | Neanderthal & Denisovan | Innate immunity (pathogen recognition) | Europeans and East Asians (~50–60%) |
| Keratin gene cluster | Neanderthal | Skin and hair adaptation | Europeans and East Asians (variable) |
| TBX15/WARS2 | Denisovan | Body fat distribution, cold adaptation | Inuit (~70%) |
Introgression deserts
Although archaic DNA is distributed broadly across modern human genomes, its distribution is far from uniform. Comprehensive genome-wide surveys have identified large regions that are essentially devoid of Neanderthal or Denisovan ancestry, termed introgression deserts. These regions are thought to represent portions of the genome where archaic alleles were systematically removed by purifying selection because they were incompatible with the modern human genetic background.4, 5
Sankararaman and colleagues mapped the landscape of Neanderthal ancestry across the genomes of over 1,000 present-day individuals and found that regions depleted of archaic DNA are enriched for genes expressed in the brain and in the male germline, particularly the testes.4 The depletion near testis-expressed genes is consistent with a role for hybrid male sterility in limiting introgression, a pattern well documented in other hybridising animal species. The X chromosome is particularly depleted of Neanderthal ancestry — approximately fivefold lower than the autosomal average — which is consistent with Haldane's rule, the observation that the heterogametic sex (males, in mammals) is more likely to suffer reduced fertility in hybrids.4, 5
Analysis of regulatory regions has added further resolution to this picture. Petr and colleagues demonstrated that Neanderthal-derived variants in gene regulatory elements, particularly enhancers and promoters active in the brain, have been preferentially removed from modern human genomes over the past 45,000 years, suggesting ongoing purifying selection against archaic regulatory variants even as coding-sequence variants persist.18 This pattern implies that regulatory incompatibilities, rather than protein-coding differences alone, were a major source of fitness costs in Neanderthal-modern human hybrids.
Phenotypic legacy in modern humans
Beyond the clear cases of adaptive introgression, archaic DNA has been linked to a wide range of phenotypic variation in present-day humans through genome-wide association studies and analyses of biobank datasets. Dannemann and Kelso systematically examined the phenotypic consequences of Neanderthal-derived alleles using data from the UK Biobank and found associations with traits including skin tone, hair colour, sleep patterns, mood, and pain sensitivity.19 Some of these associations may reflect adaptation to high-latitude environments, while others may be incidental consequences of genetic drift.
Neanderthal-derived variants have also been associated with cranial morphology. Gunz and colleagues identified two Neanderthal-introgressed loci that influence endocranial shape, with carriers of the archaic alleles tending toward a slightly more elongated braincase shape reminiscent of Neanderthals, as opposed to the globular cranial form characteristic of modern humans. These alleles affect genes expressed during early brain development, suggesting that archaic introgression has left subtle traces in the neurodevelopmental programmes of living people.20
The phenotypic effects of Denisovan introgression have been less thoroughly characterised, in part because fewer modern populations carry substantial Denisovan ancestry and because large biobank datasets are not yet available for the most Denisovan-enriched populations. However, beyond the well-established case of EPAS1 in Tibetans and TBX15/WARS2 in Inuit populations, Denisovan-derived variants have been implicated in immune function across Melanesian populations and in variation in tooth morphology, consistent with the distinctively large and robust dental remains recovered from Denisova Cave.16, 17
Multiple admixture events
The initial picture of a single admixture episode between Neanderthals and the ancestors of all non-Africans has given way to a considerably more complex model involving multiple gene flow events in different directions and at different times. Evidence now supports at least two pulses of Neanderthal introgression into modern human lineages. East Asian populations carry slightly more Neanderthal ancestry (approximately 2.3 to 2.6 percent) than European populations (approximately 1.8 to 2.0 percent), a difference that may reflect an additional admixture event or differential dilution by unadmixed African-ancestry populations.21, 5
Gene flow was not unidirectional. Analysis of the genome of a Neanderthal from the Altai Mountains revealed segments of modern human DNA, indicating that early modern humans interbred with Neanderthals and contributed genetic material to Neanderthal populations as well. The timing of this reverse gene flow, estimated at roughly 100,000 years ago, predates the main out-of-Africa dispersal and may correspond to an earlier, geographically limited expansion of modern humans into the Near East.11
Recent work has also detected low levels of Neanderthal-like ancestry in some African populations, which had previously been considered free of archaic admixture. Chen and colleagues developed improved statistical methods for identifying introgressed segments and found that several West African populations carry approximately 0.3 percent ancestry from a Neanderthal-related lineage. This signal may result from back-migration of Neanderthal-admixed Eurasian populations into Africa, rather than direct Neanderthal-African contact.13
Estimated archaic ancestry in modern human populations1, 5, 16, 21
Superarchaic introgression
The web of archaic gene flow extends even deeper than the interactions between modern humans, Neanderthals, and Denisovans. Genomic analyses have revealed that the Denisovan genome itself contains DNA from a more ancient, or "superarchaic," hominin lineage that diverged from the modern human-Neanderthal-Denisovan clade approximately 1 million years ago. Rogers, Harris, and Achenbach used a method based on the frequency of shared and derived mutations among modern human, Neanderthal, and Denisovan genomes to identify a signal of introgression from an unknown population into Denisovans, contributing roughly 1 percent of their genome.14
The identity of this superarchaic source remains unknown, though Homo erectus, which occupied parts of Asia for over a million years, is a plausible candidate based on geography and chronology. If some of this superarchaic DNA was subsequently passed from Denisovans to modern humans through the admixture events described above, then living humans may carry trace amounts of genetic material from an even more deeply divergent hominin lineage, adding yet another layer to the complex mosaic of ancestry that characterises the modern human genome.14
Broader implications
The discovery of widespread archaic introgression has reshaped the understanding of human evolution in several fundamental ways. First, it has demonstrated that species boundaries among Late Pleistocene hominins were permeable, and that hybridisation was not a rare accident but a recurrent process with lasting evolutionary consequences. The survival of archaic alleles in present-day populations — particularly those under positive selection — indicates that introgression served as a significant source of adaptive genetic variation during the global expansion of modern humans.7, 15
Second, the existence of introgression deserts and the progressive purging of archaic regulatory variants demonstrate that Neanderthal-modern human and Denisovan-modern human hybrids suffered reduced fitness in certain respects, consistent with partial reproductive isolation between these lineages. The concentration of depleted regions near genes involved in brain development and male fertility suggests that these were the functional domains most sensitive to interspecific genetic incompatibilities.4, 18
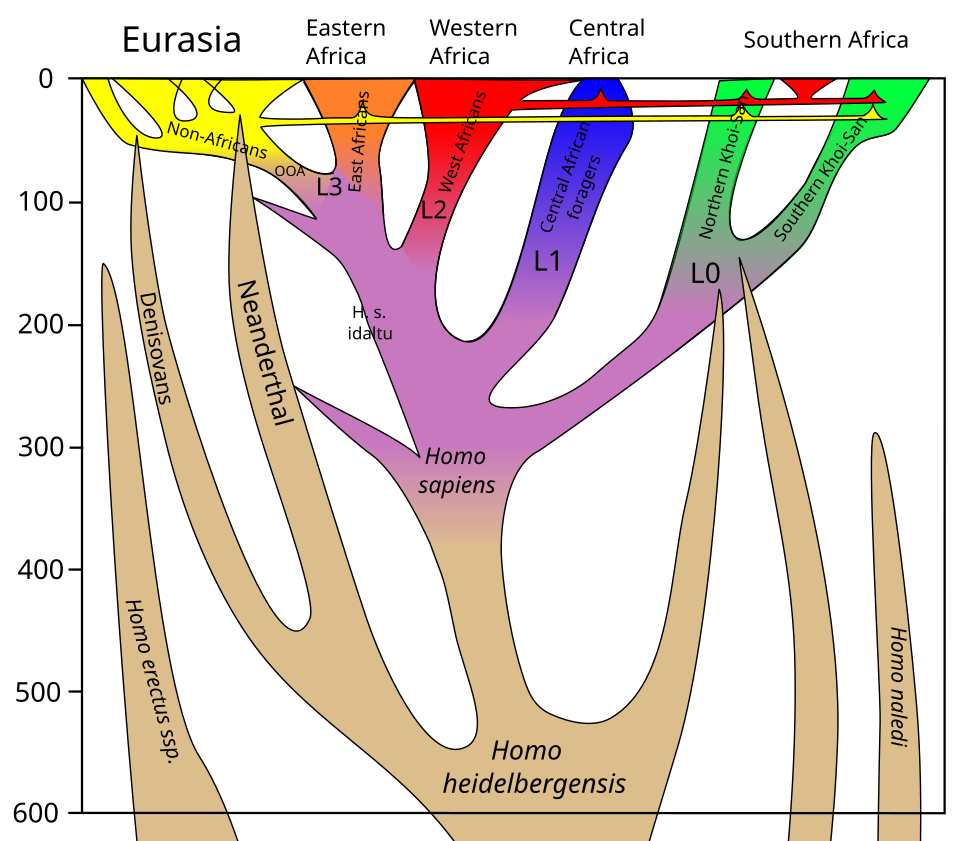
Third, the identification of superarchaic introgression into Denisovans, and possibly deeper gene flow events yet to be discovered, implies that the hominin lineage has been characterised by reticulate evolution — a branching pattern interwoven with gene flow — for at least the past million years. Rather than a cleanly bifurcating tree, the evolutionary history of the genus Homo increasingly resembles a braided stream, in which populations diverged, evolved in partial isolation, and periodically reconnected through hybridisation, leaving a complex mosaic of ancestries in every living human genome.11, 14
References
The shaping of modern human immune systems by multiregional admixture with archaic humans
Introgression of Neandertal- and Denisovan-like haplotypes contributes to adaptive variation in human Toll-like receptors