Overview
- The 2010 draft Neanderthal genome revealed that 1–4% of the DNA of all living non-African humans derives from interbreeding with Neanderthals, overturning earlier mitochondrial DNA studies that had found no evidence of admixture.
- Neanderthal-derived DNA is unevenly distributed across the human genome, depleted near genes on the X chromosome and in testes-expressed loci—suggesting reduced fertility in hybrid males—but enriched at immune and skin-pigmentation genes where archaic variants conferred adaptive advantages.
- Far from a simple branching tree, human evolution involved repeated gene flow among divergent hominin lineages, making admixture a central rather than incidental feature of our species' history.
For most of the twentieth century, the relationship between Neanderthals and modern humans was framed as a straightforward question: were they ancestors, or were they an evolutionary dead end replaced entirely by expanding Homo sapiens populations? The molecular revolution in paleoanthropology transformed this debate. Early mitochondrial DNA studies suggested complete replacement with no interbreeding, but the sequencing of the Neanderthal nuclear genome in 2010 revealed that all living non-African humans carry approximately 1 to 4 percent Neanderthal-derived DNA, the unmistakable genetic signature of ancient admixture.1, 3 Subsequent work has mapped the distribution of this archaic ancestry across the human genome, identified regions where Neanderthal variants were advantageous or deleterious, estimated the timing and number of interbreeding events, and fundamentally recast human evolution as a reticulate process — a braided stream of gene flow rather than a cleanly bifurcating tree.5, 6
Early molecular evidence against admixture
The first direct molecular comparison between Neanderthals and modern humans came in 1997, when Matthias Krings, Svante Pääbo, and colleagues extracted and sequenced a 379-base-pair segment of the hypervariable region I of mitochondrial DNA (mtDNA) from the original Neanderthal type specimen discovered in the Neander Valley in 1856. The Neanderthal sequence fell well outside the range of variation observed in living humans, and phylogenetic analyses placed the most recent common ancestor of the Neanderthal and modern human mitochondrial lineages at roughly 550,000 to 690,000 years ago — far older than the coalescence time of all modern human mtDNA lineages. Krings and colleagues concluded that Neanderthals had made no lasting contribution to the modern human mitochondrial gene pool.1
In 2004, David Serre, Pääbo, and colleagues extended this analysis by extracting mtDNA from four additional Neanderthal specimens from Germany, Russia, and Croatia, and from five early modern human fossils from sites across Europe. All four Neanderthal specimens yielded mtDNA sequences clustering with previously published Neanderthal sequences, while none of the early modern humans carried Neanderthal-type mtDNA. The authors concluded that this pattern excluded any large genetic contribution from Neanderthals to early modern humans in Europe, though they acknowledged that a small contribution — below the detection threshold of mtDNA analysis — could not be ruled out.2
These mtDNA results were widely interpreted as decisive evidence for the complete replacement of Neanderthals by modern humans, and they dominated the scientific consensus for over a decade. However, mitochondrial DNA represents only a tiny fraction of the genome and is inherited exclusively through the maternal line, meaning that it is blind to any admixture mediated through male hybrids or to nuclear gene flow that did not involve the mitochondrial lineage. The limitations of mtDNA as a tool for detecting low-level admixture would become starkly apparent once nuclear genome data became available.1, 2
The Neanderthal Genome Project
The decisive breakthrough came in 2010, when Richard Green, Pääbo, and a large international team published a draft sequence of the Neanderthal nuclear genome, assembled from DNA extracted from three Neanderthal bones from Vindija Cave in Croatia dated to approximately 38,000 to 44,000 years ago. By comparing the Neanderthal genome to the genomes of five present-day humans — a French European, a Han Chinese, a Papuan, a Yoruba from West Africa, and a San from southern Africa — the team discovered that the Neanderthal genome shared significantly more derived alleles with the three non-African individuals than with either of the two African individuals. The most parsimonious explanation was that Neanderthals had interbred with the ancestors of non-African populations after those ancestors left Africa but before they dispersed across Eurasia and Oceania. The estimated proportion of Neanderthal ancestry in non-Africans was 1 to 4 percent.3
This finding overturned the prevailing replacement model and established what has been called the leaky replacement or assimilation model: modern humans did replace Neanderthals across their range, but not before absorbing a measurable fraction of Neanderthal genetic material through interbreeding. The fact that Neanderthal ancestry was approximately equal in Europeans, East Asians, and Papuans suggested that the primary admixture event occurred in the Near East or western Asia, where expanding modern human populations first encountered Neanderthals, rather than later in Europe where the two groups coexisted for thousands of years.3
In 2014, Kay Prüfer and colleagues published a high-coverage (52-fold) Neanderthal genome from a toe bone found in Denisova Cave in the Altai Mountains of Siberia. This high-quality sequence confirmed the earlier admixture estimates and further revealed that the Altai Neanderthal's parents were closely related — equivalent to half-siblings — indicating that inbreeding was common among at least some Neanderthal populations. The Altai genome also provided evidence for multiple gene flow events among Neanderthals, Denisovans, and early modern humans, painting an increasingly complex picture of Late Pleistocene hominin interactions.4
Mapping introgressed segments
With the Neanderthal reference genome in hand, researchers turned to identifying exactly which segments of the modern human genome derive from Neanderthal ancestors. In two landmark 2014 studies published within weeks of each other, Sriram Sankararaman and colleagues and Benjamin Vernot and Joshua Akey independently mapped Neanderthal-introgressed haplotypes across large samples of present-day human genomes.5, 6
Sankararaman and colleagues analysed 1,004 present-day individuals and found that while the average Neanderthal ancestry per individual was approximately 1 to 3 percent, the introgressed segments were not uniformly distributed across the genome. By aggregating the Neanderthal ancestry across all individuals, the team estimated that roughly 20 percent of the Neanderthal genome could be recovered in fragments spread among present-day humans. Some genomic regions were significantly enriched in Neanderthal ancestry, while others were conspicuously depleted, forming so-called deserts of introgression where Neanderthal DNA had been systematically purged by natural selection.5
Vernot and Akey reached similar conclusions using an independent computational approach applied to 665 individuals of European and East Asian ancestry. Their method identified Neanderthal-derived haplotypes without requiring a Neanderthal reference genome, instead relying on patterns of linkage disequilibrium and allele frequency that distinguish introgressed segments from ancestral modern human variation. They recovered more than 15 gigabases of introgressed sequence spanning approximately 20 percent of the Neanderthal genome, confirming the Sankararaman results and demonstrating that Neanderthal ancestry in East Asians was slightly but significantly higher than in Europeans — approximately 1.5 to 2.1 percent versus 1.4 to 1.8 percent.6 Subsequent analyses have refined the Neanderthal ancestry fraction to approximately 2 percent in most Eurasian populations, with the East Asian excess now attributed to additional episodes of admixture or to differences in the strength of purifying selection between populations.16, 19
Timing and number of admixture events
The timing of Neanderthal–sapiens interbreeding has been estimated through several independent lines of evidence, including the length of introgressed haplotype blocks (which shorten over generations due to recombination), the degree of sequence divergence between Neanderthal and introgressed segments, and direct dating of ancient human individuals with elevated Neanderthal ancestry. Collectively, these approaches point to a primary admixture event occurring approximately 50,000 to 60,000 years ago, most likely in the Near East or western Asia as modern humans expanded out of Africa.3, 5
The discovery of individuals with unusually high Neanderthal ancestry has provided direct evidence that admixture was not limited to a single episode. In 2015, Qiaomei Fu and colleagues reported genome-wide data from Oase 1, a modern human mandible from Peștera cu Oase in Romania dated to approximately 37,000 to 42,000 years ago. Oase 1 carried 6 to 9 percent Neanderthal ancestry — substantially more than any present-day human — and three of its Neanderthal-derived chromosomal segments exceeded 50 centimorgans in length, indicating that the individual had a Neanderthal ancestor as recently as four to six generations back. Remarkably, Oase 1 did not contribute detectably to later European populations, suggesting that at least some early modern human lineages in Europe interbred with Neanderthals but subsequently went extinct.7
In 2021, Mateja Hajdinjak and colleagues reported genome-wide data from three individuals dated to 42,580 to 45,930 years ago from Bacho Kiro Cave in Bulgaria, associated with Initial Upper Palaeolithic artefacts. All three carried 3.4 to 3.8 percent Neanderthal ancestry with long introgressed segments, again indicating recent Neanderthal forebears within roughly six to seven generations. Unlike Oase 1, however, the Bacho Kiro individuals were more closely related to present-day East Asian and Native American populations than to later western Eurasians, demonstrating that some early admixed lineages did persist and contribute to living populations.22
Villanea and Schraiber (2019) applied maximum-likelihood and machine-learning analyses to the distribution of Neanderthal ancestry in European and East Asian populations and found that a model of a single admixture pulse could not explain the observed patterns. Instead, the data favoured multiple episodes of gene flow, with at least one additional pulse into the ancestors of East Asians after their separation from Europeans. This finding helps explain the slightly elevated Neanderthal ancestry in East Asian populations and underscores that Neanderthal–sapiens contact was not a singular event but a recurring phenomenon over thousands of years of coexistence.19
Deserts of introgression and hybrid incompatibility
Although Neanderthal DNA persists throughout the genomes of non-African humans, its distribution is strikingly non-random. Sankararaman and colleagues identified broad regions of the genome where Neanderthal ancestry is almost entirely absent — deserts of introgression that are far more depleted of archaic DNA than would be expected by chance. These deserts are concentrated in two categories of genomic regions: the X chromosome and autosomal regions enriched for genes expressed in the testes.5
The X chromosome carries roughly five times less Neanderthal ancestry than the autosomes, a pattern that is strikingly reminiscent of Haldane's rule — the empirical observation that in crosses between divergent species, the heterogametic sex (males, in mammals) is more likely to be sterile or inviable. The depletion of Neanderthal ancestry on the X chromosome and near testes-expressed genes strongly suggests that male hybrids between Neanderthals and modern humans suffered reduced fertility, and that natural selection progressively eliminated Neanderthal alleles from these reproductively sensitive regions over thousands of generations.5, 6
Beyond the X chromosome, Neanderthal ancestry is broadly depleted in the vicinity of genes across the genome, particularly near genes with high expression levels and in regions of low recombination. This genome-wide depletion reflects the action of purifying selection against mildly deleterious Neanderthal variants. Harris and Nielsen (2016) proposed that the Neanderthal population, which had a long-term effective size of only a few thousand individuals, had accumulated a substantial burden of weakly deleterious mutations through genetic drift. When these mutations entered the much larger modern human population through admixture, they were exposed to more efficient purifying selection, and the carriers of Neanderthal segments with deleterious alleles were gradually removed from the population. The authors estimated that Neanderthals had at least 40 percent lower fitness than modern humans on average, and that the residual Neanderthal mutational load reduces the fitness of present-day non-Africans by at least 0.5 percent.12
Juric, Aeschbacher, and Coop (2016) reached complementary conclusions using a different analytical framework, finding that selection against individual Neanderthal alleles is very weak on average but that the aggregate effect across thousands of loci is substantial. Most of the purifying selection against Neanderthal ancestry appears to act on many weakly deleterious alleles rather than on a few strongly harmful ones.13 Petr and colleagues (2019) further showed that the bulk of this purifying selection occurred rapidly after the initial admixture event and that Neanderthal ancestry has remained relatively stable in European populations over the past 45,000 years, with no evidence for a continued long-term decline.20
Distribution of Neanderthal ancestry across the human genome5, 16
Adaptive introgression
While much of the Neanderthal DNA retained in modern humans appears to be selectively neutral or mildly deleterious, a growing body of evidence demonstrates that some introgressed variants were positively selected because they conferred adaptive advantages in the environments that modern humans encountered outside Africa. This process, known as adaptive introgression, allowed expanding human populations to acquire locally adapted genetic variants from Neanderthals who had evolved in Eurasian environments for several hundred thousand years, rather than having to evolve these adaptations from scratch.8
Skin and keratin biology. Among the most compelling examples of adaptive introgression are Neanderthal-derived variants in genes involved in keratin filament formation and skin pigmentation. Sankararaman and colleagues (2014) found that genes in the keratin pathway are significantly enriched for Neanderthal ancestry in both Europeans and East Asians.5 A particularly striking case involves the BNC2 gene on chromosome 9, which encodes a zinc finger protein expressed in keratinocytes and associated with skin pigmentation and freckling. The Neanderthal-derived haplotype at BNC2 reaches frequencies of approximately 70 percent in European populations, far above the genome-wide average of 2 percent, providing strong evidence for positive selection on this introgressed variant.15 Dannemann and Kelso (2017), using phenotype data from 112,000 UK Biobank participants, showed that multiple Neanderthal alleles at different loci contribute to variation in skin tone and hair colour in present-day Europeans, and that Neanderthal alleles are associated with both lighter and darker pigmentation, suggesting that Neanderthals themselves were variable in these traits.17
Immune system genes. Innate and adaptive immune genes constitute perhaps the most important category of adaptive introgression from Neanderthals. Abi-Rached and colleagues (2011) demonstrated that archaic HLA class I haplotypes — the highly polymorphic molecules of the major histocompatibility complex that present pathogen-derived peptides to T cells — were introduced into modern human populations through admixture with both Neanderthals and Denisovans. These archaic HLA alleles now represent more than half of the HLA-A, HLA-B, and HLA-C variants in some Eurasian populations, a remarkable frequency that reflects strong pathogen-driven selection maintaining these variants over tens of thousands of years.9
Deschamps and colleagues (2016) showed that innate immunity genes as a class harbour higher Neanderthal ancestry than the remainder of the coding genome. Among the loci with the highest Neanderthal ancestry, the TLR6–TLR1–TLR10 cluster on chromosome 4 stands out. These Toll-like receptors recognise molecular patterns on the surfaces of bacteria and fungi and trigger pro-inflammatory immune responses.10 Dannemann, Andrés, and Kelso (2016) confirmed that archaic-like alleles at the TLR cluster are associated with increased microbial resistance and, as a trade-off, with increased susceptibility to allergic disease in present-day Europeans — a vivid demonstration that adaptive introgression can have both beneficial and detrimental phenotypic consequences.11
Antiviral defence. The OAS gene cluster on chromosome 12 encodes 2′-5′-oligoadenylate synthetases, enzymes that activate a ribonuclease pathway to degrade viral RNA inside infected cells. Mendez, Watkins, and Hammer (2013) identified a ~185-kilobase haplotype of Neanderthal origin spanning this cluster, nearly restricted to Eurasian populations, that diverged from the Neanderthal lineage approximately 125,000 years ago.23 Enard and Petrov (2018) provided broader evidence that RNA viruses drove adaptive introgression between Neanderthals and modern humans. They found that proteins interacting specifically with RNA viruses were significantly more likely to reside within introgressed segments, suggesting that Neanderthal-derived alleles provided resistance against novel viral pathogens encountered during the expansion into Eurasia.18
Phenotypic consequences in living humans
Beyond immune function and pigmentation, Neanderthal-derived variants influence a surprisingly wide range of traits in present-day humans. Simonti and colleagues (2016) systematically tested the associations between common Neanderthal variants and more than 1,000 clinical phenotypes recorded in the electronic health records of approximately 28,000 adults of European ancestry. They discovered and replicated significant associations between Neanderthal alleles and phenotypes spanning neurological, psychiatric, immunological, and dermatological categories. Neanderthal alleles collectively explained a significant fraction of the variation in risk for depression, in skin lesions caused by sun exposure (actinic keratosis), and in blood hypercoagulability.14
Dannemann and Kelso (2017) extended these analyses using 112,000 participants from the UK Biobank and identified Neanderthal alleles significantly associated with hair colour, skin tone, ease of tanning, frequency of childhood sunburn, body height, sleeping patterns, mood, and smoking status. Notably, the directions of these associations were not uniform: at some loci, the Neanderthal allele was associated with darker skin, while at others it was associated with lighter skin, reinforcing the view that phenotypic variation in Neanderthals was substantial and that introgressed variants can push modern human phenotypes in either direction depending on the specific allele and the genetic background into which it was introduced.17
Selected Neanderthal-introgressed variants and their phenotypic associations5, 14, 15, 17
| Gene or region | Chromosome | Phenotype | Frequency in Europeans |
|---|---|---|---|
| BNC2 | 9 | Skin pigmentation, freckling | ~70% |
| TLR6–TLR1–TLR10 | 4 | Innate immunity, allergy risk | ~50% |
| HLA class I (HLA-A, -B, -C) | 6 | Adaptive immunity, pathogen defence | Variable (up to >50% at some alleles) |
| OAS1–OAS2–OAS3 | 12 | Antiviral innate immunity | ~30–40% |
| Keratin pathway genes | Multiple | Hair and skin morphology | Variable (enriched) |
| Hypercoagulation loci | Multiple | Blood clotting phenotypes | Variable |
Archaic admixture in Africa
The discovery of Neanderthal admixture in non-African populations naturally raised the question of whether analogous gene flow from other archaic hominins occurred within Africa, where the fossil record documents the presence of multiple morphologically distinct hominin populations during the Middle and Late Pleistocene. In 2020, Arun Durvasula and Sriram Sankararaman developed statistical methods to detect introgression from an unsampled — or ghost — archaic population and applied them to whole-genome data from four West African populations. Their analyses indicated that 2 to 19 percent of the genetic ancestry of these populations traces to an archaic population that diverged from the modern human lineage before the split between modern humans and Neanderthals, with estimated split times ranging from 360,000 to over 1 million years ago and admixture times between 0 and 124,000 years ago.21
The identity of this ghost archaic population remains unknown, as no ancient DNA has been recovered from Middle Pleistocene fossils in sub-Saharan Africa due to the poor preservation conditions in tropical environments. Candidate taxa include late-surviving Homo heidelbergensis or Homo naledi populations, though these attributions are speculative in the absence of genomic data. Regardless of the source population's identity, the finding of archaic introgression in Africa demonstrates that gene flow between divergent hominin lineages was not unique to the encounter between Neanderthals and modern humans in Eurasia but was a widespread feature of hominin evolution across the globe.21
Reticulate evolution and the braided stream
The cumulative evidence from Neanderthal–sapiens admixture, Denisovan introgression into Oceanian and Asian populations, and archaic African gene flow has fundamentally transformed the conceptual framework through which scientists understand human evolution. The traditional model of a cleanly branching phylogenetic tree — with lineages splitting and evolving in isolation — has given way to a model of reticulate evolution, in which divergent lineages repeatedly come into contact and exchange genes before eventually becoming reproductively isolated or going extinct.4, 8
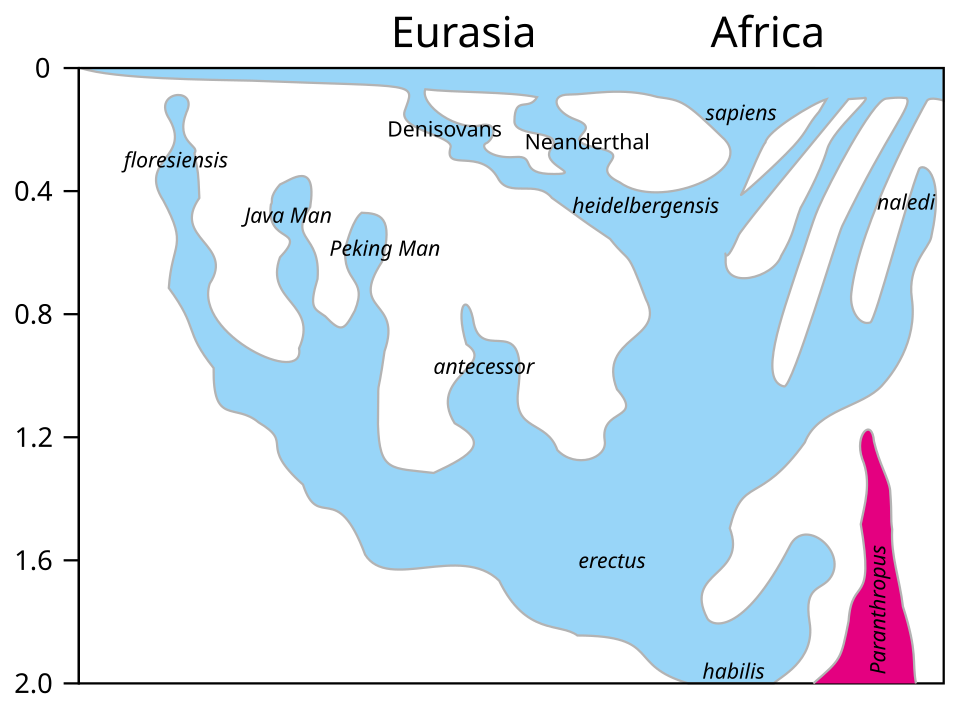
The metaphor of a braided stream has been proposed to capture this pattern: just as a braided river splits into multiple channels that diverge and reconverge as they flow downstream, hominin populations diverged from one another geographically and morphologically but periodically reconnected through migration and interbreeding. The Neanderthal genome was not a closed book upon the divergence of the Neanderthal and modern human lineages some 550,000 to 700,000 years ago; instead, gene flow continued episodically across that boundary for hundreds of thousands of years, with the most consequential pulses occurring as modern humans expanded into Neanderthal territory approximately 50,000 to 60,000 years ago and continued for at least another 10,000 to 15,000 years before Neanderthals disappeared from the fossil record.3, 7, 19, 22
The Neanderthal contribution to the modern human genome is neither trivial nor overwhelming. At approximately 2 percent of any individual's genome, it is modest in total quantity, yet the aggregate Neanderthal DNA recoverable across all living non-Africans covers roughly 20 percent of the Neanderthal genome, meaning that a substantial fraction of Neanderthal genetic diversity persists in fragmentary form in the collective genomes of living people.5, 6 Some of that legacy is deleterious — a residual mutational burden from a small, inbred population — but some has been unambiguously beneficial, providing modern humans with immune defences, pigmentation variants, and other adaptations that facilitated survival in Eurasian environments. The combined landscape of Denisovan and Neanderthal ancestry in present-day humans, mapped by Sankararaman and colleagues (2016), reveals shared patterns of depletion and enrichment across archaic sources, suggesting that similar evolutionary forces shaped the retention and loss of introgressed DNA from both archaic groups.16
The emerging picture is one in which admixture was a central, creative force in human evolution rather than an anomaly. The genomes of all living humans are mosaics, built from contributions of multiple populations that met, interbred, and in some cases subsequently vanished from the fossil record. Understanding the full scope and consequences of this admixture — identifying the specific genes and traits shaped by archaic contributions, disentangling the selective forces that retained or eliminated introgressed variants, and extending the analysis to the still-undersampled populations of Africa and southern Asia — remains one of the most active frontiers in human evolutionary genetics.8, 16, 21
References
The shaping of modern human immune systems by multiregional admixture with archaic humans
Genomic signatures of selective pressures and introgression from archaic hominins at human innate immunity genes
Introgression of Neandertal- and Denisovan-like haplotypes contributes to adaptive variation in human Toll-like receptors
Evidence that RNA viruses drove adaptive introgression between Neanderthals and modern humans