Overview
- Artificial selection is the deliberate breeding of organisms by humans to favour desired traits, and it has transformed wild species into the domesticated plants and animals that sustain modern civilization over roughly 10,000 years.
- Charles Darwin drew extensively on pigeon breeding and other examples of artificial selection to argue that natural selection could produce comparable changes given sufficient time, making domestication a cornerstone of the argument in On the Origin of Species.
- Modern artificial selection encompasses rapid-cycle laboratory evolution in organisms such as Drosophila and bacteria, quantitative trait locus mapping in crop improvement, and marker-assisted breeding, all of which confirm that heritable variation under sustained directional selection produces cumulative evolutionary change.
Artificial selection is the process by which humans breed plants and animals for particular traits by choosing which individuals are allowed to reproduce. Unlike natural selection, which operates through differential survival and reproduction in response to environmental pressures, artificial selection is driven by human preferences — for larger fruit, docile temperament, faster growth, or novel appearance. Charles Darwin devoted the entire first chapter of On the Origin of Species to artificial selection, arguing that if human breeders could produce dramatic changes in domesticated species within a few generations, then natural selection acting over millions of years could readily account for the diversity of life.1 The study of artificial selection remains central to evolutionary biology, providing direct, observable evidence that heritable variation, when subjected to sustained directional pressure, produces cumulative and sometimes radical evolutionary change.
Darwin and the pigeon analogy
Darwin was an avid pigeon fancier who joined two London pigeon clubs and maintained a loft of living birds at his home in Down House. He chose pigeon breeding as his opening argument for evolution because the results were so visually striking: from the ancestral rock pigeon (Columba livia), breeders had produced fantails with 42 tail feathers (the wild type has 12), pouters with enormously inflated crops, tumblers that somersault in flight, and jacobins with feathered hoods so extreme they can barely see.1, 6 Darwin noted that if these varieties had been found in the wild, any naturalist would classify them as separate species, yet all descended from a single ancestor through selective breeding spanning perhaps a few hundred years.
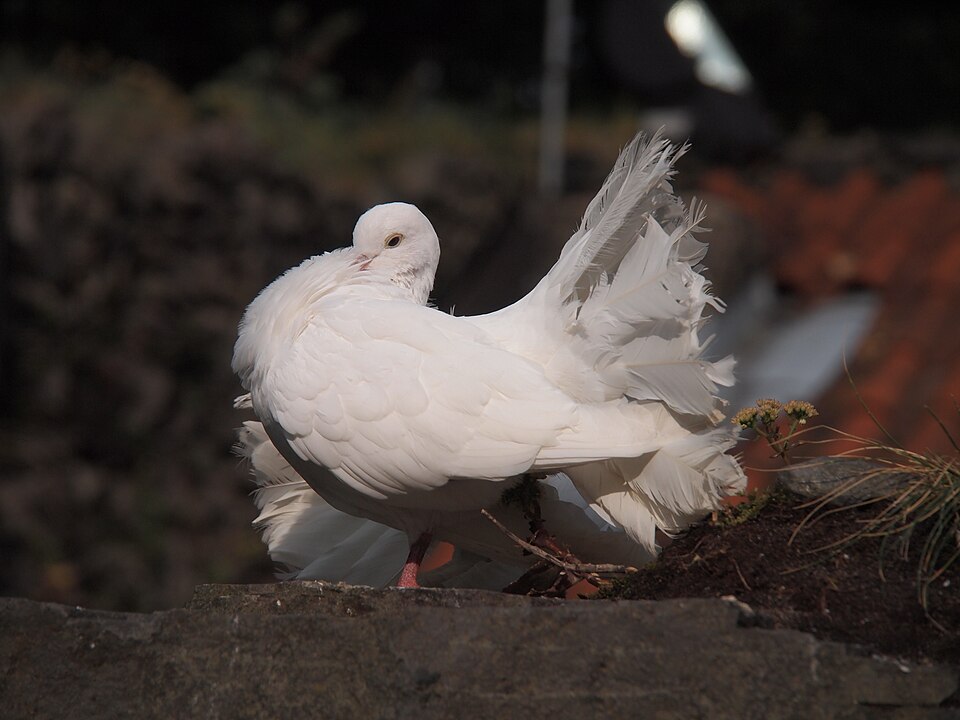

The power of this argument lay in its accessibility. Every reader of the Origin was familiar with domesticated animals and plants, and Darwin used that familiarity to establish two principles that would underpin his entire theory: first, that variation exists within every population; and second, that when certain variants are consistently favoured, the population shifts in the direction of the favoured trait over successive generations.1 Modern genomic studies have confirmed Darwin's inference that all domestic pigeon breeds derive from Columba livia, and have identified specific loci responsible for the dramatic morphological differences among breeds, including the EphB2 gene underlying the head crest phenotype.6
Mechanisms of artificial selection
The mechanism of artificial selection is conceptually straightforward: a breeder examines a population, selects individuals exhibiting the desired trait, allows only those individuals to mate, and repeats the process each generation. This imposes directional selection — a persistent bias in reproductive success toward one extreme of the trait distribution. Over generations, the population mean shifts in the direction of the breeder's preference, and alleles associated with the favoured trait increase in frequency while alternative alleles decline.1, 5
The response to selection depends on the heritability of the trait — the proportion of phenotypic variation attributable to additive genetic effects. Traits with high heritability respond rapidly to selection, while traits strongly influenced by environmental variation respond slowly or not at all. The breeder's equation, R = h2S, formalises this relationship: the response to selection (R) equals the heritability (h2) multiplied by the selection differential (S), the difference between the mean trait value of the selected parents and the overall population mean.15 This quantitative framework, developed in the early twentieth century with the rise of Mendelian genetics, transformed animal and plant breeding from an intuitive craft into a predictive science.
Artificial selection can also impose genetic bottlenecks. When only a small number of individuals are chosen as breeding stock, genetic diversity is reduced, and deleterious recessive alleles can become fixed through drift. Many purebred dog breeds suffer from high rates of inherited diseases — hip dysplasia in German shepherds, brachycephalic airway syndrome in bulldogs — as a direct consequence of intense selection from small founder populations.2, 13
Domestication of the dog
The domestic dog (Canis lupus familiaris) represents the oldest and most extensively studied case of animal domestication. Genomic analyses indicate that dogs diverged from wolves at least 15,000 years ago, and possibly as early as 30,000 years ago, making them the only animal domesticated before the advent of agriculture.4, 14 The initial phase of domestication may have involved a degree of self-selection, in which wolves with lower flight distances and reduced aggression were able to exploit food resources near human camps, but subsequent deliberate breeding rapidly diversified dogs into the more than 400 recognised breeds that exist today.2
The domestic dog genome, sequenced in 2005, revealed that breed-defining traits — body size, coat texture, skull shape — are often controlled by a remarkably small number of genes. Variation at the IGF1 locus accounts for a large proportion of the body-size difference between small and large breeds, and a single mutation in the FGF4 gene produces the chondrodysplasia (shortened limbs) characteristic of dachshunds and corgis.2 The speed with which artificial selection has reshaped dog morphology — most modern breeds were established within the past 200 years — demonstrates how rapidly phenotypic evolution can proceed when selection intensity is high and generation time is short.
Crop domestication
The domestication of crop plants, which began approximately 10,000 years ago in the Fertile Crescent and independently in several other regions, constitutes one of the most consequential episodes of artificial selection in human history.11 Early farmers, whether consciously or inadvertently, selected for traits that collectively define the domestication syndrome in plants: loss of natural seed dispersal (so seeds remain on the plant for harvest), increased seed size, reduced seed dormancy (allowing synchronous germination), and more determinate growth habit.5, 12
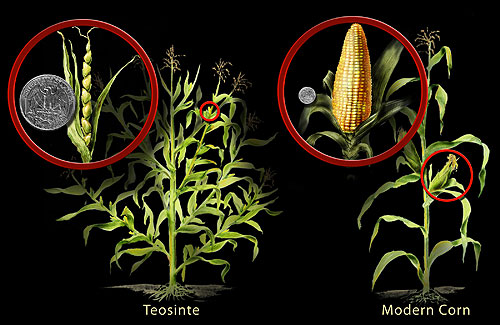
The transformation of teosinte into modern maize (Zea mays) is perhaps the most dramatic example. Teosinte, the wild ancestor of maize, is a Mexican grass with small ears bearing 5 to 12 hard-cased kernels arranged in two rows. Modern maize ears carry 500 or more exposed kernels in up to 20 rows. Genetic mapping has shown that this radical morphological shift was driven primarily by changes at a small number of regulatory loci, most notably teosinte branched1 (tb1), which controls plant architecture and apical dominance.3, 7 Similarly, the Waxy locus in rice underwent a selective sweep during domestication, fixing alleles that produce the glutinous starch preferred in many Asian cuisines.8
Selected domestication events and approximate dates5, 11, 16
| Species | Wild ancestor | Region | Approximate date |
|---|---|---|---|
| Dog | Grey wolf (Canis lupus) | Eurasia | ~15,000–30,000 BP |
| Wheat | Wild emmer (Triticum dicoccoides) | Fertile Crescent | ~10,000 BP |
| Maize | Teosinte (Zea mays ssp. parviglumis) | Mesoamerica | ~9,000 BP |
| Rice | Wild rice (Oryza rufipogon) | East Asia | ~8,000–9,000 BP |
| Pig | Wild boar (Sus scrofa) | Near East & China | ~9,000 BP |
| Cattle | Aurochs (Bos primigenius) | Near East | ~10,000 BP |
Laboratory selection experiments
Artificial selection in the laboratory provides some of the most controlled and rigorous demonstrations of evolutionary change. The fruit fly Drosophila melanogaster, with its short generation time of roughly two weeks, has been subjected to selection experiments for over a century.
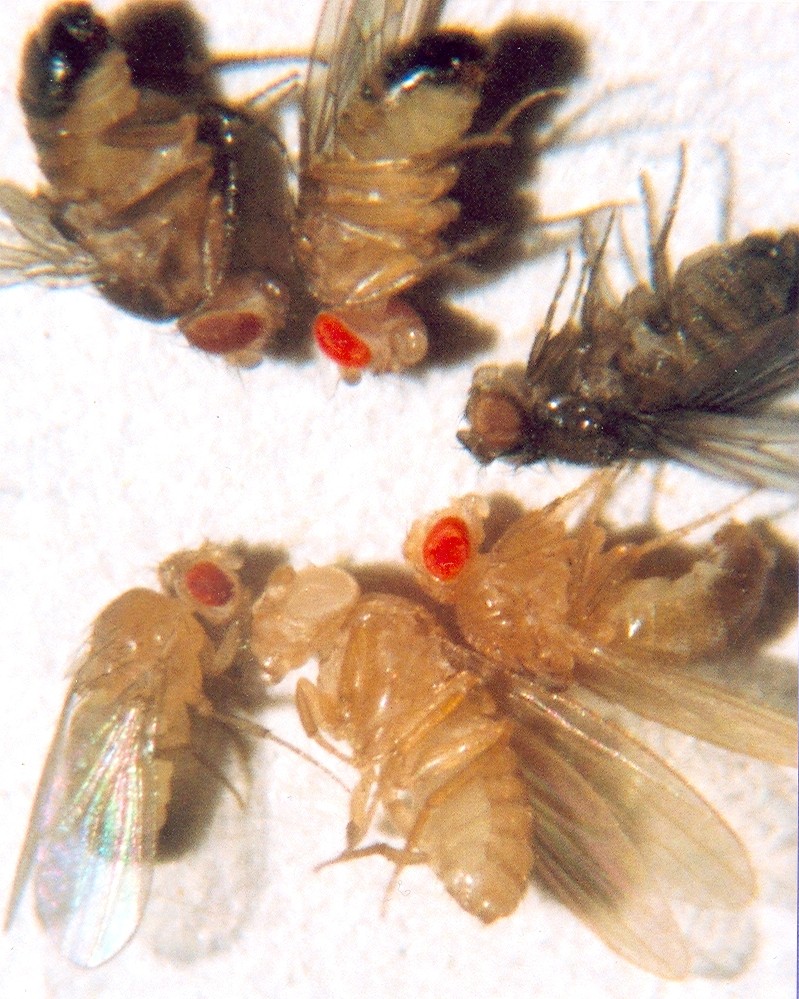

Researchers have successfully selected for increased and decreased bristle number, body size, longevity, geotaxis (preference for moving up or down), ethanol tolerance, and mating speed, confirming that virtually any measurable trait with heritable variation will respond to sustained directional selection.9
Richard Lenski's long-term evolution experiment with Escherichia coli, begun in 1988, extends the principle of artificial selection to microbial populations. Although this experiment tracks evolution under constant environmental conditions rather than imposing a breeder's choice, the serial transfer protocol imposes a form of selection for competitive fitness in glucose-limited medium. Over tens of thousands of generations, the bacterial populations have exhibited repeatable adaptive changes, including increased growth rate, altered cell size, and in one famous lineage, the evolution of a novel ability to metabolise citrate under aerobic conditions.10 These experiments demonstrate that the same processes that produce new dog breeds or crop varieties — heritable variation filtered by differential reproduction — operate across all domains of life.
Modern breeding and quantitative genetics
Contemporary plant and animal breeding has moved far beyond the intuitive selection practised by Darwin-era fanciers. Quantitative trait locus (QTL) mapping allows researchers to identify the specific chromosomal regions responsible for variation in complex traits such as grain yield, disease resistance, or milk production, enabling breeders to select for favourable alleles with precision.15 Marker-assisted selection (MAS) uses DNA markers linked to known QTLs to screen seedlings or embryos before the trait is even expressed, dramatically accelerating the breeding cycle and reducing the cost of field trials.17
These advances have not altered the fundamental logic of artificial selection — heritable variation filtered by a selection criterion — but they have compressed timescales and increased precision. The Green Revolution of the 1960s and 1970s, which doubled wheat and rice yields through the introduction of semi-dwarf varieties selected for short stature and high harvest index, was accomplished largely through conventional crossing and phenotypic selection.12 Modern genomics-assisted breeding builds on this foundation by allowing breeders to target specific genes and predict the outcome of crosses before they are made, further confirming that the evolutionary principles Darwin inferred from pigeon lofts operate at every scale of biological organisation.
Implications for understanding evolution
Artificial selection remains one of the most powerful lines of evidence for evolution by natural selection. It demonstrates three propositions that are necessary and sufficient for evolution to occur: that populations contain heritable variation; that variation in traits affects reproductive success (in this case, success determined by the breeder); and that the offspring of selected parents resemble their parents more than they resemble random members of the population.1 Every domesticated species is a living experiment in evolution, and the speed with which artificial selection reshapes organisms undermines any claim that natural selection lacks the power to produce large-scale change given geological time.
The parallel between artificial and natural selection is not perfect. Breeders can impose far stronger selection differentials than most natural environments, and they can maintain traits that would be disadvantageous in the wild — no bulldog would survive as a feral predator, and no domestic turkey could fly to escape a coyote.2 Nevertheless, the underlying mechanism is identical: differential reproduction based on heritable trait variation. Darwin's insight that the familiar transformations of the farmyard and the pigeon loft could be extrapolated to the whole of life remains one of the most important conceptual achievements in the history of science.1
References
Darwin's pigeons and the evolution of the columbiforms: recapitulation of ancient genes
Teosinte branched1 and the origin of maize: evidence for epistasis and the evolution of dominance
Long-term experimental evolution in Escherichia coli. I. Adaptation and divergence during 2,000 generations
Single nucleotide polymorphism (SNP) variation of wolves (Canis lupus) in Southeast Alaska and comparison with wolves, dogs, and coyotes in North America
Marker-assisted selection: an approach for precision plant breeding in the twenty-first century