Overview
- Chromosomes evolve through structural rearrangements including Robertsonian fusions, pericentric and paracentric inversions, reciprocal translocations, and duplications, which can alter gene linkage, suppress recombination, and create reproductive barriers between populations.
- Whole-genome duplication (polyploidy) has been a major force in the evolution of plants, fishes, and other lineages, with polyploid events often associated with bursts of speciation and the origin of major evolutionary innovations due to the creation of redundant gene copies free to diverge in function.
- The human karyotype of 46 chromosomes includes chromosome 2, which arose from the fusion of two ancestral chromosomes still present as separate entities in chimpanzees and other great apes, providing direct cytogenetic evidence for common ancestry between humans and the other apes.
Chromosomes are not static structures. Over evolutionary time, they undergo a variety of structural changes — fusions, fissions, inversions, translocations, duplications, and whole-genome duplications — that alter gene order, chromosome number, and genome architecture. These chromosomal rearrangements have played a central role in the evolution of species, both by creating the raw material for adaptive innovation through gene duplication and by establishing reproductive barriers that drive speciation.2, 7
Robertsonian fusions and fissions
Robertsonian translocations are among the most common chromosomal rearrangements in mammals. They involve the fusion of two acrocentric chromosomes (chromosomes with the centromere near one end) at their centromeres to form a single metacentric or submetacentric chromosome, reducing the diploid chromosome number by two without a net change in gene content, or the reverse process (fission) in which a metacentric chromosome splits into two acrocentric chromosomes.1
The most celebrated example of a Robertsonian fusion in human evolution is human chromosome 2. Humans have 46 chromosomes (23 pairs), while chimpanzees, gorillas, and orangutans all have 48 (24 pairs). IJdo and colleagues demonstrated in 1991 that human chromosome 2 contains a vestigial telomere-telomere fusion site at band 2q13, precisely where two ancestral chromosomes (corresponding to chimpanzee chromosomes 2p and 2q) fused end-to-end. The fusion site contains degenerate telomeric repeat sequences (TTAGGG)n oriented head-to-head, exactly as predicted by a telomere-telomere fusion, and the chromosome retains a vestigial second centromere (now inactive) corresponding to the centromere of one of the ancestral chromosomes.4 Chromosome painting studies confirmed that the banding pattern and gene content of human chromosome 2 correspond precisely to the combined content of the two separate great ape chromosomes.3, 12
Robertsonian fusions and fissions are also widespread in other mammalian lineages. The house mouse (Mus musculus) exhibits remarkable karyotypic variation across European populations, with diploid numbers ranging from 22 to 40 due to varying numbers of Robertsonian fusions. Different fusion combinations characterise different geographic populations, and hybrids between populations carrying incompatible fusions show reduced fertility due to meiotic segregation errors, suggesting that Robertsonian rearrangements can contribute to reproductive isolation and incipient speciation.1, 2
Chromosomal inversions and speciation
Inversions — the reversal of a segment of chromosome relative to its normal orientation — are among the most important chromosomal rearrangements for the evolution of species. Inversions are classified as pericentric (including the centromere) or paracentric (not including the centromere). Because an inversion reverses gene order, recombination within an inverted segment in heterozygotes (individuals carrying both the normal and inverted arrangements) produces unbalanced gametes with duplications and deletions, which are typically inviable. This effective suppression of recombination within inverted regions has profound evolutionary consequences.5
By suppressing recombination, inversions can hold together co-adapted gene complexes — groups of alleles at different loci that function well together but would be broken up by recombination. This property has led to the concept of "supergenes," in which an inversion locks together alleles controlling multiple components of a complex phenotype, such as mimicry patterns in butterflies or mating types in birds.13 The classic example involves the white-throated sparrow (Zonotrichia albicollis), in which a large pericentric inversion on chromosome 2 captures genes affecting plumage colour, song behaviour, and mating strategy, effectively creating two alternative "morphs" that are maintained by disassortative mating.13
In Drosophila, inversions have been implicated in speciation through the suppression-of-recombination model. Noor and colleagues proposed that inversions accumulate between incipient species because they capture and protect alleles that contribute to reproductive isolation (such as genes affecting mate choice or hybrid inviability) from being disrupted by gene flow between the diverging populations.5 Comparative cytogenetic studies of Drosophila species pairs have confirmed that genomic regions within inversions show greater genetic divergence than collinear regions, consistent with the prediction that inversions facilitate speciation by shielding divergent loci from homogenisation by gene flow.5, 6
Polyploidy: whole-genome duplication
Polyploidy — the possession of more than two complete sets of chromosomes — is one of the most dramatic forms of chromosomal change and has been a major force in the evolution of plants, fishes, amphibians, and other lineages. Polyploidy can arise through autopolyploidy (duplication of a single genome, typically by failure of meiotic reduction) or allopolyploidy (hybridisation between two species followed by genome doubling, combining two distinct genomes in a single organism).7, 11
In plants, polyploidy is extraordinarily common. Approximately 15 percent of angiosperm speciation events are estimated to involve polyploidy, and many of the world's most important crop species are polyploid, including wheat (hexaploid, 6x), cotton (tetraploid, 4x), potato (tetraploid, 4x), and oilseed rape (tetraploid, 4x).7, 14 The genus Tragopogon (goatsbeards) provides a natural laboratory for studying allopolyploid speciation in real time: two allopolyploid species (T. mirus and T. miscellus) formed within the past century through hybridisation between diploid species introduced to North America, and genomic studies of these young polyploids have documented rapid genomic restructuring, gene loss, and epigenetic changes occurring within just a few generations of polyploid formation.16
Allopolyploidy can produce instant reproductive isolation from both parental species, because the polyploid hybrid typically produces inviable or infertile offspring when crossed with either diploid parent due to mismatched chromosome numbers during meiosis. This immediate barrier to gene flow means that allopolyploidy can generate new species in a single generation, making it one of the few known mechanisms of sympatric speciation that does not require geographic isolation.11, 14
Genome duplications in vertebrate evolution
The vertebrate lineage experienced two rounds of whole-genome duplication early in its history, an event supported by the analysis of paralogous gene sets distributed across four chromosomal regions in the human and other vertebrate genomes (the 2R hypothesis). Dehal and Boore provided strong computational evidence for these duplications by analysing the pattern of four-fold paralogy blocks in the human genome, which is consistent with two successive tetraploidisation events followed by extensive gene loss and chromosomal rearrangement.8
These ancient genome duplications provided raw material for the evolution of vertebrate complexity. Duplicated genes that were retained (rather than being lost by pseudogenisation) diversified through subfunctionalisation (partitioning of the ancestral function) and neofunctionalisation (acquisition of a novel function), contributing to the expansion and elaboration of gene families involved in development, signalling, and neural function.8 The four paralogous Hox gene clusters of vertebrates, for example, arose through these duplications and have been linked to the increased morphological complexity of the vertebrate body plan.
Teleost fishes experienced an additional, third round of whole-genome duplication (the teleost-specific genome duplication, or TSGD) approximately 320 to 350 million years ago. The TSGD produced a dramatic increase in gene number, and the subsequent divergence of duplicated genes has been associated with the explosive diversification of teleosts, which comprise more than half of all living vertebrate species.9 In fungi, the yeast Saccharomyces cerevisiae experienced a whole-genome duplication approximately 100 million years ago, and comparison with pre-duplication species has illuminated the process of diploidisation, in which the doubled genome is restructured through reciprocal gene loss between duplicated chromosomal segments.10
Post-polyploidy genome restructuring
The immediate aftermath of a polyploidy event is a period of genomic instability during which the newly formed polyploid genome undergoes rapid structural and regulatory changes. Studies of newly synthesised allopolyploids in wheat (Triticum-Aegilops crosses) have documented the loss of specific DNA sequences, changes in transposable element activity, alterations in gene expression, and epigenetic modifications occurring within the first few generations after polyploid formation.15
Over longer evolutionary timescales, polyploid genomes undergo a process of diploidisation in which the genome progressively reverts toward a diploid-like state through differential gene loss from duplicated chromosomal segments, chromosomal rearrangements that break up synteny between homeologous (related by the polyploidy event) chromosomes, and the divergence of duplicated regulatory elements. This process has been documented in detail in yeast, maize, and Arabidopsis, all of which carry the signatures of ancient polyploidy events that have been largely obscured by subsequent genome restructuring.7, 10
Chromosomal change and the diversification of life
Chromosomal evolution operates at the interface of genome structure and population genetics, linking molecular-level changes in DNA organisation to organism-level processes of adaptation, reproductive isolation, and speciation. The fusion that created human chromosome 2 is a direct cytogenetic record of our shared ancestry with the great apes.4 The inversions that suppress recombination between incipient species facilitate the accumulation of genetic incompatibilities that complete the speciation process.5 And the polyploidy events that have punctuated the history of plants and vertebrates have generated the redundant genetic material from which new functions evolve, driving some of the most significant evolutionary transitions in the history of life.7, 8
Reciprocal translocations
Reciprocal translocations—the exchange of segments between non-homologous chromosomes—are another class of chromosomal rearrangement with significant evolutionary consequences. Unlike Robertsonian fusions, which involve whole-arm exchanges at centromeres, reciprocal translocations can break chromosomes at any point and rejoin segments in new combinations, creating derivative chromosomes with novel gene linkage arrangements. In heterozygous carriers, these rearrangements produce unbalanced gametes during meiosis due to adjacent segregation from the translocation quadrivalent, reducing fertility and creating a partial reproductive barrier between carriers and non-carriers of the translocation.17
Faria and Navarro analysed the relationship between chromosomal rearrangement rates and speciation rates across mammalian lineages and found a significant positive correlation: lineages with higher rates of chromosomal rearrangement tend to have higher rates of speciation. This correlation held after controlling for potential confounds such as body size and generation time, suggesting that chromosomal rearrangements play a causal role in promoting reproductive isolation and accelerating the speciation process.17 The pattern has been documented across diverse mammalian orders, from carnivores and primates to rodents and artiodactyls, suggesting that chromosomal change is a general facilitator of mammalian diversification.2, 17
Chromosomal evolution and sex determination
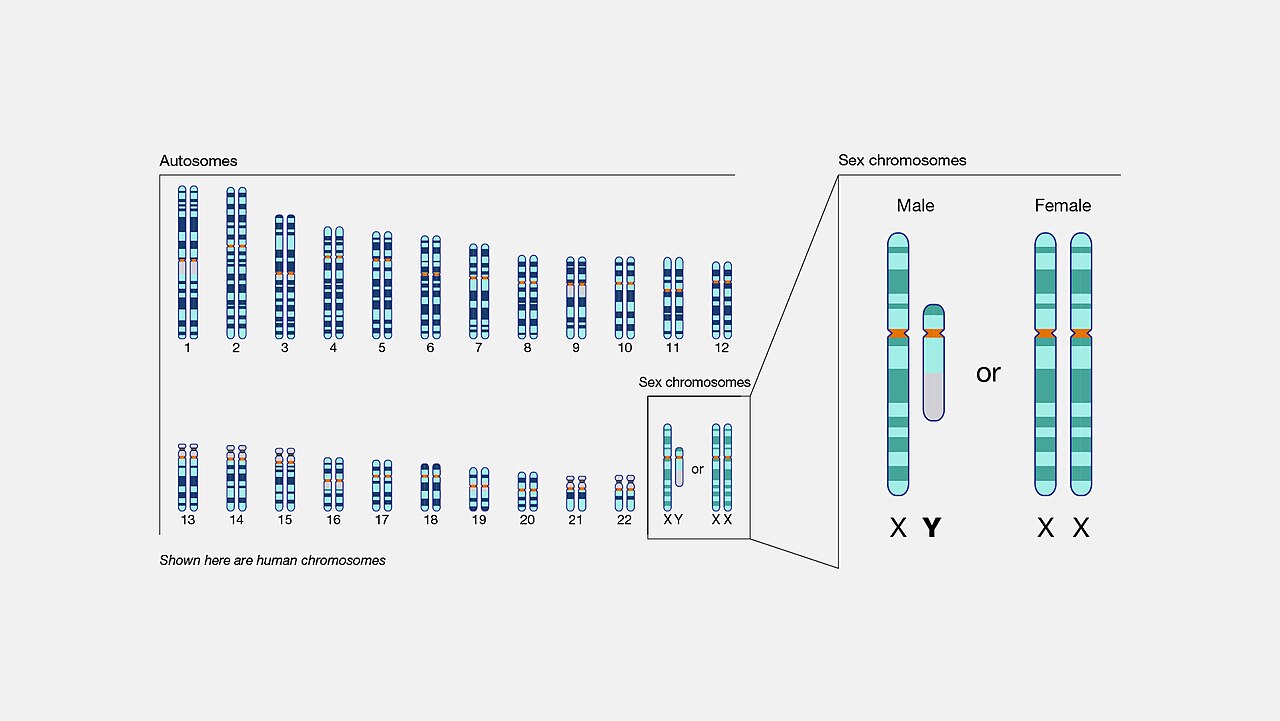
The evolution of sex chromosomes provides a particularly well-studied example of how chromosomal change shapes genome architecture over long timescales. The mammalian Y chromosome originated as an ordinary autosome that acquired a sex-determining gene (SRY) and then progressively degenerated through a series of inversions that suppressed recombination with the X chromosome. Once recombination was lost, the Y chromosome accumulated deleterious mutations, transposable element insertions, and gene losses, shrinking from an ancestral autosome of approximately 1,500 genes to the present human Y chromosome of fewer than 80 protein-coding genes.2, 18
Sex chromosome evolution has followed different trajectories in different lineages. In birds, the female is the heterogametic sex (ZW), and the W chromosome has degenerated in a manner analogous to the mammalian Y. In some plant species, such as papaya (Carica papaya), sex-determining regions are relatively young, showing early stages of recombination suppression and incipient degeneration that provide a window into the initial steps of sex chromosome evolution.7, 19 The comparative study of sex chromosomes across taxa illustrates how the same chromosomal mechanisms—inversions, suppressed recombination, and subsequent degeneration—independently produce convergent outcomes in unrelated lineages.
References
Genomic divergences between humans and other hominoids and the effective population size of the common ancestor of humans and chimpanzees
Chromosome painting shows that the three great ape chromosomes equivalent to human chromosome 2 are not only rearranged but also differentiated by structural variation
Allopolyploid speciation in Tragopogon: a model for revealing the dynamics of genome evolution
The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus)