Overview
- Deep homology refers to the discovery that vastly different animals share the same core developmental genes—a conserved genetic toolkit inherited from a common ancestor over 500 million years ago.
- The same master regulatory gene, Pax6, controls eye development in both insects and vertebrates; Hox genes pattern body segments in flies and humans; Tinman/Nkx2-5 directs heart formation across animal phyla—despite the enormous morphological differences between these structures.
- Deep homology reveals that evolution does not build new body plans from scratch but instead reuses and rewires ancient genetic programs, explaining both the fundamental unity and the extraordinary diversity of animal life.
One of the most surprising discoveries of modern biology is that vastly different animals—flies and fish, worms and whales, jellyfish and humans—share a remarkably conserved set of genes that control the fundamental processes of embryonic development. These genes, often called the developmental genetic toolkit, were inherited from a common ancestor that lived over 500 million years ago, before the major animal phyla diverged. The concept of deep homology, articulated by Shubin, Tabin, and Carroll, describes this phenomenon: structures that appear very different on the surface (such as insect eyes and vertebrate eyes, or fly legs and mouse limbs) are built using the same underlying genetic programs, revealing a shared developmental ancestry far deeper than their morphological differences would suggest.1, 2
Pax6 and the universal eye gene
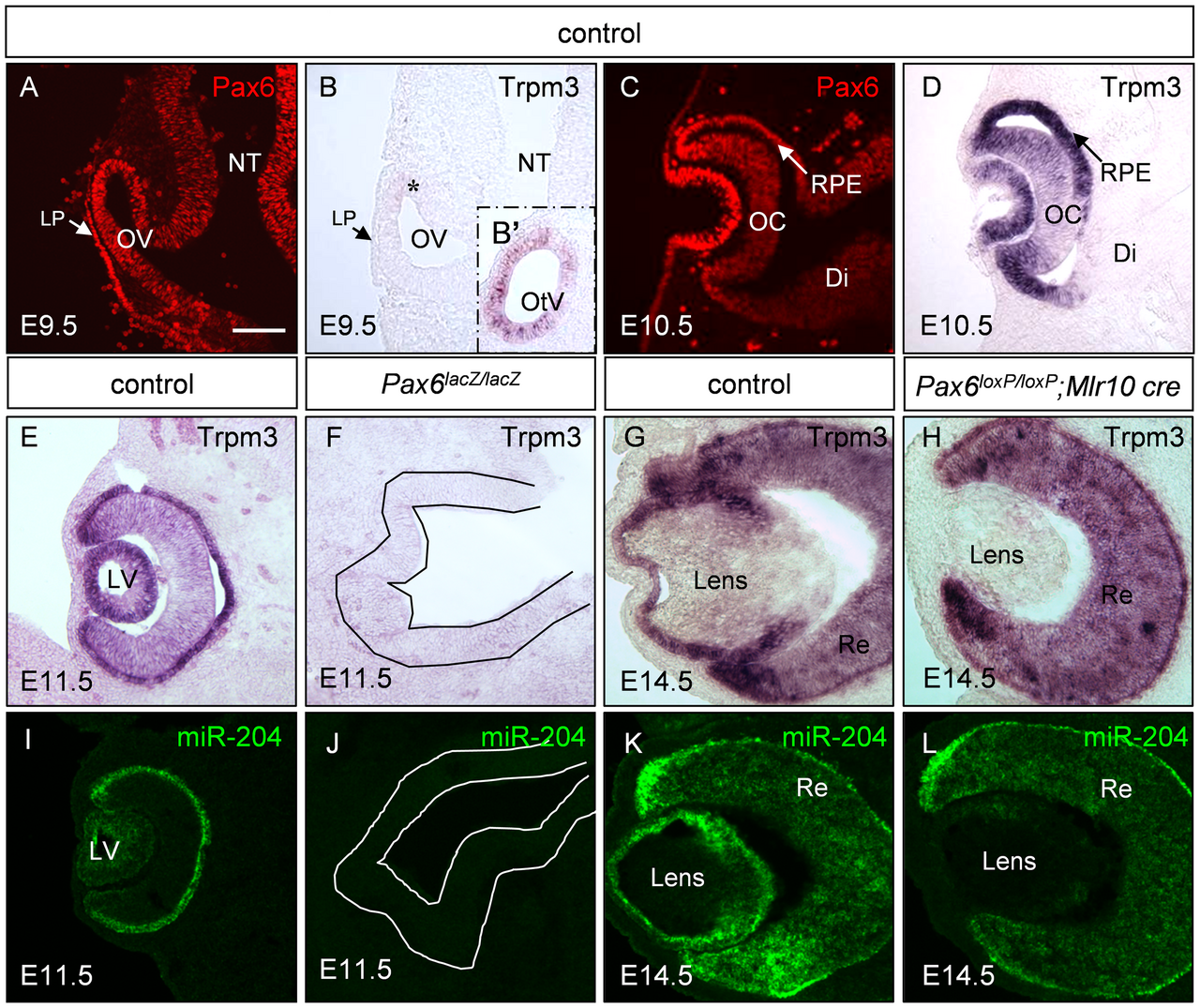
The gene Pax6 is arguably the most famous example of deep homology. In vertebrates, Pax6 is a master regulatory gene for eye development; mutations that inactivate one copy cause the condition aniridia (absence of the iris) in humans and the "Small eye" phenotype in mice. In the fruit fly Drosophila melanogaster, the homologous gene is called eyeless, and its loss produces flies without eyes.3, 4
The remarkable discovery came in 1995, when Walter Gehring's laboratory demonstrated that the mouse Pax6 gene, when experimentally expressed in various tissues of Drosophila, could induce the formation of ectopic compound eyes on the fly's wings, legs, and antennae.3 A mammalian gene, separated from the insect lineage by over 500 million years of independent evolution, could still activate the downstream eye-development program in a fly. This functional interchangeability indicated that Pax6 is not merely a gene that happens to be involved in eye development in both lineages—it is the ancestral master switch for eye formation, inherited from a common bilaterian ancestor and retained in both arthropod and vertebrate lineages despite the vast morphological differences between compound eyes and camera eyes.4, 5, 13
Eyes have evolved independently at least 40 times across the animal kingdom, producing structures as different as the pin-hole eyes of nautiluses, the compound eyes of insects, and the camera eyes of vertebrates and cephalopods. The discovery that these diverse structures are all initiated by the same master regulatory gene suggests that the ancestral bilaterian animal possessed a simple light-sensing organ controlled by Pax6, and that subsequent eye evolution in different lineages elaborated on this ancient genetic foundation rather than inventing eye development from scratch.1, 13
Hox genes and body patterning
Hox genes encode transcription factors that specify positional identity along the head-to-tail axis of animal embryos. They are arranged in clusters on the chromosome, and their physical order along the chromosome corresponds to the order in which they are expressed along the body axis—a property called collinearity. In Drosophila, Hox genes specify the identity of each body segment, determining, for instance, whether a segment bears wings, legs, or halteres. In vertebrates, Hox genes perform an analogous function, specifying the identity of vertebral regions (cervical, thoracic, lumbar, sacral) and patterning the limbs, hindbrain, and other structures.6, 7
The deep homology here is profound. The Hox genes of flies and mice are not merely similar in sequence; they are functionally interchangeable to a remarkable degree. Experimental replacement of a mouse Hox gene with its Drosophila counterpart can partially rescue the normal developmental program, demonstrating that the biochemical function of these proteins has been conserved over half a billion years of divergent evolution.6 The common ancestor of insects and vertebrates possessed a single Hox cluster, and this ancestral toolkit has been inherited, duplicated, and redeployed in different ways across the animal kingdom. Vertebrates expanded their Hox toolkit through whole-genome duplication, producing four Hox clusters that enabled finer patterning of increasingly complex body plans.6, 7
Tinman/Nkx2-5 and heart development
The tinman gene in Drosophila (named because flies lacking this gene develop without a heart, like the Tin Man from The Wizard of Oz) controls the specification of cardiac precursor cells in the fly embryo. Its vertebrate homolog, Nkx2-5, plays an equivalent role in mammalian heart development; mutations in Nkx2-5 cause congenital heart defects in humans.8, 9
The fly heart is a simple dorsal tube that pumps hemolymph through an open circulatory system. The mammalian heart is a four-chambered muscular organ with valves, septa, and a closed circulatory system. Despite these vast structural differences, the initial specification of cardiac tissue in both organisms is directed by homologous transcription factors, acting through conserved downstream regulatory networks.8, 9 This conservation indicates that the genetic program for heart specification predates the divergence of arthropods and vertebrates and was present in their last common ancestor, a simple organism that likely possessed a rudimentary contractile vessel. The subsequent elaboration of this vessel into the fly heart and the mammalian heart occurred through changes in gene regulation rather than through the invention of new genes.1, 9
Dorsal-ventral axis inversion
Another striking example of deep homology involves the genetic control of dorsal-ventral (back-to-belly) patterning. In vertebrates, the signaling molecule BMP4 specifies ventral (belly-side) identity, while its antagonist Chordin specifies dorsal (back-side) identity. In Drosophila, the homologous molecules Decapentaplegic (Dpp, homolog of BMP4) and Short gastrulation (Sog, homolog of Chordin) perform the same function but with inverted polarity: Dpp specifies dorsal structures in the fly, while Sog specifies ventral structures.12
This "inversion" of the dorsal-ventral axis between vertebrates and arthropods had been noted by the nineteenth-century anatomist Étienne Geoffroy Saint-Hilaire, who observed that arthropods appear to be built like "upside-down" vertebrates (with the nerve cord ventral rather than dorsal). The molecular discovery that the same signaling system controls dorsal-ventral patterning in both groups, but with opposite polarity, provided a genetic explanation for Geoffroy's observation and confirmed that the body plans of vertebrates and arthropods are deeply homologous, built from the same ancestral developmental program that was inverted in one lineage.12
Distal-less and the origin of appendages
The gene Distal-less (Dll) provides yet another example of deep homology. Panganiban et al. demonstrated that Dll is expressed in the developing appendages of organisms as diverse as insects, crustaceans, spiders, sea urchins, and vertebrates.14 The appendages of these animals are morphologically very different—insect legs, lobster claws, butterfly wings, sea urchin tube feet, and vertebrate limbs have little superficial resemblance. Yet in every case, Dll is expressed in the distal (outermost) portion of the developing appendage, suggesting that all animal appendages share a common developmental origin in an ancestral appendage-patterning program controlled by Dll.1, 14
Implications for evolutionary theory
Deep homology fundamentally changed how biologists understand the evolution of animal body plans. Before the molecular revolution in evolutionary developmental biology, the independent evolution of complex structures like eyes, hearts, and limbs in different animal lineages was seen as evidence that evolution could build similar solutions from scratch through convergent evolution. The discovery of deep homology revealed that these structures are not as independently derived as they appear. They are built from the same ancient genetic toolkit, inherited from a common ancestor and redeployed in different developmental contexts.1, 10, 11
This insight explains one of the most puzzling features of animal evolution: how can organisms so different in form be so similar at the genetic level? The answer is that the diversity of animal body plans arises not primarily from the evolution of new genes but from changes in when, where, and how much existing toolkit genes are expressed. The same Pax6 gene can build a compound eye or a camera eye; the same Hox genes can pattern an insect thorax or a vertebrate spine; the same Tinman/Nkx2-5 gene can specify a dorsal tube or a four-chambered heart. The toolkit is ancient and shared; the developmental instructions for deploying it are what evolve.1, 10, 11
Conserved signaling pathways
Deep homology extends beyond individual master regulatory genes to entire signaling cascades that coordinate tissue growth and patterning. The Hedgehog signaling pathway, which plays a central role in limb development, neural patterning, and organ formation in vertebrates, is conserved in arthropods, where it regulates segment polarity and appendage growth. Goodrich and colleagues demonstrated that the vertebrate Patched receptor, which mediates Hedgehog signaling in mammals, is a direct ortholog of the Drosophila Patched protein, and that the vertebrate and insect versions of the pathway share the same ligand-receptor logic and downstream transcriptional effectors.15 Similarly, the Wnt, Notch, and TGF-β signaling pathways are conserved across the animal kingdom, performing analogous functions in cell fate determination, tissue boundary formation, and stem cell maintenance in organisms as different as sea anemones, flatworms, and mammals.11, 16
The conservation of these signaling systems has practical implications beyond evolutionary theory. Because the core molecular components of developmental signaling are shared between invertebrate model organisms and humans, discoveries made in Drosophila or C. elegans can directly inform the understanding of human genetic diseases. Mutations in human Hedgehog pathway components cause conditions such as holoprosencephaly and basal cell carcinoma; mutations in Notch signaling cause Alagille syndrome and certain leukemias. In each case, the fundamental biology was first elucidated in invertebrate systems precisely because the pathways are deeply homologous.10, 15
The Wnt/β-catenin pathway further illustrates the breadth of deep homology. This pathway controls axis formation in vertebrate embryos and regulates regeneration in planarian flatworms, where inhibition of Wnt signaling causes the formation of heads instead of tails at posterior amputation sites. Bely and Nyberg showed that the molecular pathways controlling regeneration are shared across diverse animal phyla, suggesting that the capacity for regeneration was present in the bilaterian ancestor and has been independently reduced or lost in lineages such as mammals, rather than independently evolved in organisms like planarians and salamanders.16, 17
References
Homologous genetic mechanisms underlie the formation of vastly different structures in vertebrates and invertebrates
Homology of the eyeless gene of Drosophila to the Small eye gene in mice and Aniridia in humans
Decapentaplegic and short gastrulation-like genes are components of a conserved dorsal-ventral patterning pathway
Conservation of the Hedgehog/Patched signaling pathway from flies to mice: induction of a mouse patched gene by Hedgehog
The evolution of the bilaterian gut: insights from the fossil record and molecular phylogeny