Overview
- A genetic bottleneck is a severe, temporary reduction in population size that eliminates most allelic diversity; the surviving gene pool reflects chance rather than fitness, and much genetic variation is permanently lost.
- Multiple independent analytical methods — including pairwise sequentially Markovian coalescent (PSMC) modeling, heterozygosity analysis, and HLA/MHC allele counting — converge on the conclusion that the ancestral human population never dropped below roughly 10,000 breeding individuals at any point in the last several hundred thousand years.
- The genetic diversity present in modern humans is mathematically incompatible with descent from a single couple or a family of eight; HLA allele diversity alone requires thousands of simultaneously breeding lineages, making a literal Adam and Eve or a global Noachian flood genetically impossible.
A genetic bottleneck is a sharp, temporary reduction in the size of a population that leaves a permanent imprint on its genome. When a population crashes — whether from disease, environmental catastrophe, geographic isolation, or random chance — the survivors carry only a fraction of the genetic diversity that existed before the event. Because the survivors are a small, non-representative sample of the original gene pool, many alleles are lost permanently, and those that survive may have done so purely by chance rather than by conferring any selective advantage. The downstream consequences are measurable for thousands of generations: reduced genetic drift-buffering capacity, elevated inbreeding coefficients, impaired immune response, and distinctive patterns in the frequency spectrum of genetic variants across the population’s genome.15 The study of bottlenecks sits at the intersection of population genetics, paleoclimatology, and — increasingly — the assessment of claims made by religious traditions about human origins.
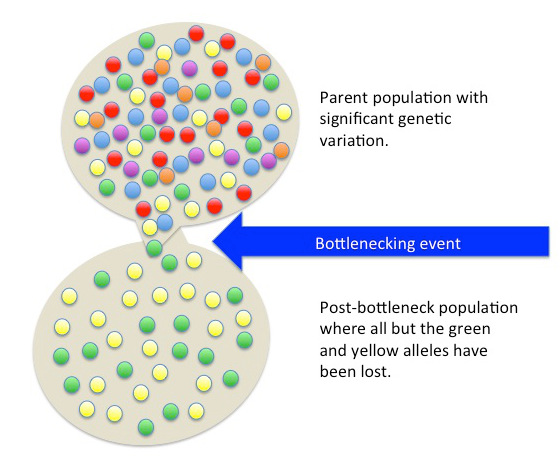
What a bottleneck is
Population geneticists distinguish between the census population size (the total number of living individuals) and the effective population size (Ne), which is the size of an idealized, randomly mating population that would exhibit the same rate of genetic drift as the real population. Effective population size is nearly always smaller than census size, because real populations have unequal sex ratios, overlapping generations, variance in reproductive success, and substructure that prevents fully random mating. A bottleneck reduces both quantities, but its genetic signature is captured through Ne.15
During a bottleneck, three processes accelerate simultaneously. First, rare alleles — those present in only a few copies in the pre-bottleneck population — are disproportionately likely to be lost by sampling chance, because there are simply fewer individuals to carry them through. Second, alleles that do survive become elevated in frequency relative to their pre-bottleneck prevalence; this produces the characteristic signature of a bottleneck in the site-frequency spectrum, where there is an excess of common variants and a deficit of rare ones. Third, the overall level of heterozygosity — the probability that two randomly drawn copies of a gene differ from each other — falls, and recovers only slowly through the accumulation of new mutations over subsequent generations.15, 14 The severity and duration of the bottleneck determine how much diversity is lost and how long recovery takes: a brief crash from 10,000 to 1,000 individuals leaves a modest signature, while a prolonged reduction to fewer than a few hundred individuals can permanently impoverish the genome.
Detecting bottlenecks in genomic data
Modern population genetics has developed several independent methods for detecting past bottlenecks from genome sequences, and their convergence on similar estimates gives population geneticists considerable confidence in their conclusions.
Heterozygosity and allele frequency spectra. The simplest signature of a bottleneck is reduced genome-wide heterozygosity and a skewed distribution of allele frequencies. Statistical tests such as Tajima’s D detect departures from the neutral expectation by comparing two estimators of the mutation rate — one sensitive to the number of segregating sites and one to average pairwise differences. A bottleneck produces a positive Tajima’s D because it eliminates rare alleles while retaining common ones, distorting the frequency spectrum in a predictable direction.14
Coalescent analysis. Coalescent theory describes how gene lineages trace back to common ancestors as one looks backward in time. In a large, stable population, coalescence times are long and uniformly distributed; in a population that recently passed through a bottleneck, many lineages coalesce rapidly near the time of the crash, leaving a distinctive star-shaped genealogy with short internal branches and long terminal ones. By fitting coalescent models to large samples of genetic loci, researchers can estimate past population sizes and identify periods of severe constriction.11
PSMC and MSMC methods. The most powerful approach for reconstructing population size through time from individual genomes is the pairwise sequentially Markovian coalescent (PSMC) method developed by Heng Li and Richard Durbin.1 PSMC treats the pattern of heterozygous sites along a single diploid genome as a hidden Markov model encoding the local time to most recent common ancestor (TMRCA) at each genomic position. Because TMRCA is proportional to effective population size at the time of coalescence, the distribution of heterozygosity across the genome encodes a continuous record of Ne through time, typically recoverable for the period between roughly 10,000 and several million years ago. The subsequent MSMC method extended this framework to multiple genomes, allowing inference of population divergence and migration as well as size history.1 Both methods have been applied to hundreds of human genomes from populations worldwide and consistently reconstruct an ancestral human effective population size that never drops below approximately 10,000 breeding individuals during the last several hundred thousand years.1, 12
Linkage disequilibrium decay. A bottleneck extends the length scale over which adjacent genomic variants are correlated (linkage disequilibrium, or LD), because it reduces the number of recombination events separating the crash from the present. By measuring the rate at which LD decays with physical distance, researchers can estimate both the timing and severity of past bottlenecks independent of allele frequency data.9
Human effective population size: the evidence
The human effective population size has been estimated by multiple independent research groups using the distinct methods described above, and the estimates are remarkably consistent. Li and Durbin’s 2011 PSMC analysis of whole-genome sequences from individuals representing six continental populations found that the ancestral human Ne ranged between approximately 10,000 and 20,000 breeding individuals across the last 100,000 years, with a well-defined African expansion beginning roughly 100,000–60,000 years ago and a non-African bottleneck associated with the out-of-Africa migration reducing Ne to around 1,200–2,500 — severe by modern-population standards, but nowhere near a pair of individuals.1
Tenesa and colleagues, analyzing linkage disequilibrium patterns across the human genome, independently estimated a long-term ancestral effective population size of approximately 10,000 individuals, with evidence that this figure had been relatively stable over deep timescales.2 Bayesian demographic inference applied to genome-wide SNP data by Sheehan and colleagues confirmed that human Ne has fluctuated but never approached the single-digit values that would be required by a literal Adam and Eve scenario.12 Scally and Durbin’s comprehensive review of mutation rate and demographic inference methods reinforced this picture: across a wide range of methodological assumptions, human Ne during the Pleistocene is consistently estimated at 10,000–30,000 individuals.11
The out-of-Africa bottleneck that accompanied the dispersal of modern humans into Eurasia beginning approximately 50,000–70,000 years ago is detectable in non-African populations as reduced nucleotide diversity, extended haplotype blocks, and altered allele frequency spectra.10 Genome-wide analyses place the founding effective size of the non-African expansion at roughly 1,000–3,000 individuals — a genuine and significant bottleneck — but still orders of magnitude larger than two or eight.10, 14 The African populations that did not pass through this founder event retain substantially greater genetic diversity, consistent with the bottleneck model but incompatible with a scenario in which all humans descend from a single couple at any point in the past few hundred thousand years.7
The Toba catastrophe hypothesis and documented bottlenecks
The most widely discussed candidate bottleneck in human prehistory is associated with the eruption of the Toba supervolcano in Sumatra approximately 74,000 years ago. Rampino and Self originally proposed that the eruption — the largest known volcanic event of the past two million years — triggered a volcanic winter lasting several years and a prolonged cooling episode, potentially reducing the global human population to a few thousand individuals.6 Ambrose elaborated this into the Toba catastrophe hypothesis, arguing that the resulting bottleneck explains the relatively low genetic diversity of non-African humans compared to African populations, and may have contributed to the subsequent rapid cultural and demographic expansion of modern humans.5
The Toba hypothesis remains contested. Archaeological evidence from southern Africa and India shows continuous human occupation through the Toba ash layer, suggesting that at least some populations survived without catastrophic disruption.7 Mitochondrial DNA analyses by Behar and colleagues found no unambiguous genetic signature of a severe population crash approximately 74,000 years ago; the coalescent patterns in global mtDNA diversity are consistent with a modest reduction followed by expansion, but not with a near-extinction event.7 PSMC analyses of whole genomes likewise do not show a sharp Ne minimum at 74,000 years ago that would be expected from a catastrophic bottleneck, though the time resolution of PSMC is insufficient to detect very brief crashes.1 The current consensus is that Toba may have imposed regional population stress on some human groups, but a global near-extinction bottleneck is not supported by the genomic evidence.
Other species provide well-documented cases of severe historical bottlenecks that serve as informative comparisons. The cheetah (Acinonyx jubatus) is perhaps the most famous example: population genomic analyses by O’Brien and colleagues demonstrated that cheetahs have extremely low genetic diversity across all molecular markers, with heterozygosity values approximately ten-fold below those of other felids.9 Whole-genome sequencing by Dobrynin and colleagues confirmed that cheetahs passed through at least two severe bottlenecks — one approximately 100,000 years ago at the end-Pleistocene megafaunal crisis and one roughly 10,000–12,000 years ago — reducing Ne to a few hundred individuals.8 The genomic consequences are stark: most cheetahs are nearly immunologically identical, such that skin grafts can be exchanged between unrelated individuals without rejection, and captive populations suffer elevated mortality from infectious disease due to homogeneous immune-gene repertoires.9 Northern elephant seals provide another example: hunted to fewer than 30 individuals in the late nineteenth century, they have since recovered numerically but retain almost no allozyme variation — a living record of the genomic impoverishment a genuine near-extinction bottleneck produces.
HLA diversity and the impossibility of two founders
The human leukocyte antigen (HLA) system, encoded by genes of the major histocompatibility complex (MHC) on chromosome 6, provides some of the most direct and intuitive evidence against a two-person founding population. HLA genes are the most polymorphic loci in the human genome: HLA-A, HLA-B, and HLA-C together have thousands of known alleles in the global human population, with any given individual carrying at most two alleles at each locus.8
A founding couple could have contributed at most four alleles per HLA locus to all subsequent humanity (two from each parent). If modern humans descended from such a pair, we would expect to find at most four alleles per HLA locus in the global population — in practice somewhat fewer, since some alleles would have been lost to genetic drift in a small founding lineage. Instead, HLA-A has more than 3,000 known alleles, HLA-B more than 4,000, and HLA-C more than 2,500, with new rare alleles still being discovered.8 The coalescence times of these alleles extend back well over one million years — predating the origin of anatomically modern Homo sapiens — and some allelic lineages are shared with chimpanzees and other great apes, demonstrating that they are maintained by balancing selection operating across deep evolutionary time.13
Bergström and colleagues’ analysis of HLA-A diversity found that allelic lineages at this single locus require a minimum ancestral population of several thousands of individuals to explain the observed diversity under any plausible model of demographic history and selection.8 Leffler and colleagues’ cross-species analysis of balancing selection found hundreds of genomic regions in both humans and chimpanzees sharing ancient polymorphisms that predate the human–chimpanzee split, requiring continuously maintained diversity across millions of years of evolution — not a pattern consistent with any recent founding bottleneck.13 Even setting aside HLA, the breadth of autosomal genetic diversity in modern humans — the number of distinct haplotype blocks, the number of rare variants private to individual populations, the variance in coalescence times across the genome — is collectively incompatible with derivation from two founders within the last 10,000 years or even the last 200,000 years.2, 1
Implications for Adam, Eve, and Noah’s Ark
The genetic evidence directly challenges two central claims of literalist interpretations of the Genesis narratives: that all humans descend from a single couple (Adam and Eve), and that a global flood approximately 4,000–5,000 years ago reduced the entire human population to eight survivors (Noah’s family).
The Adam and Eve claim, taken literally to mean that all humans share common ancestors who were the only two humans alive at any point in the past, is ruled out by the genomic evidence described above. The effective population size of humans has never dropped below approximately 10,000 breeding individuals within the evolutionary history of the species, and the diversity of HLA alleles alone requires a minimum effective population size orders of magnitude larger than two.1, 8 A weaker reading — that Adam and Eve are the genealogical ancestors of all currently living humans, even if they were not the only humans at their time — is technically compatible with genetics, since all humans do share common genealogical ancestors within the last few thousand years through the mathematics of genealogical coalescence. However, this interpretation requires abandoning the claim that Adam and Eve were the sole progenitors of humanity, and it does not restore the theological premise that all humans inherited a singular moral or biological nature from a single couple.
The Noah’s Ark scenario is more sharply falsified. A global flood approximately 4,000–5,000 years ago that reduced the human population to eight individuals would have produced a catastrophic bottleneck with a signature detectable in modern genomes: near-complete homogeneity at most loci, a single haplotype block spanning entire chromosomes, absence of allelic diversity older than approximately 4,000–5,000 years, and coalescence times for virtually all lineages converging on the same recent date. None of these patterns are observed. Human populations from Africa, the Americas, Oceania, and Eurasia carry genetic diversity whose coalescence times span from tens of thousands to over a million years, the distribution of heterozygosity is consistent with ancient and continuous population structure, and population-specific allele frequencies reflect deep divergence histories that precede any plausible flood date by hundreds of thousands of years.10, 7, 1 The genetic evidence is not merely inconsistent with eight founders at 4,000 years ago — it is inconsistent with any founding event of fewer than several thousand individuals during the entire span of modern human existence.2, 12
Some theologians and scientists within faith traditions have proposed reconciling genetic evidence with scripture through frameworks such as "Adam and Eve as representatives rather than sole progenitors," or through allegorical or literary readings of Genesis that do not require literal historical events. These interpretive moves are outside the scope of genetic analysis. What the genetic evidence establishes is the empirical boundary: no scenario in which all living humans descend from two individuals within the last several hundred thousand years is consistent with observed genomic diversity.
Conservation implications
Understanding genetic bottlenecks has direct practical consequences for conservation biology. Populations that have passed through recent bottlenecks — whether from hunting, habitat destruction, or natural catastrophe — carry reduced allelic diversity at immune-system loci, elevated frequencies of deleterious recessive alleles (whose effects are normally masked in heterozygotes), and impaired capacity to respond to novel pathogens or environmental stressors.8 The cheetah serves as a cautionary case: its near-extinction bottlenecks have produced a population so immunologically uniform that a single novel feline pathogen could sweep through individuals with little variation in resistance.9
Population managers now routinely estimate Ne and monitor heterozygosity as conservation metrics, aiming to maintain effective population sizes above the threshold of approximately 500 breeding individuals generally considered necessary to prevent ongoing genetic erosion, and above approximately 5,000 to retain long-term evolutionary potential. These thresholds are derived directly from the same population genetic theory that establishes minimum viable population sizes and informs captive breeding programs for critically endangered species. The genetics of bottlenecks is thus not merely an academic exercise in prehistory but an applied discipline with immediate relevance to the survival of species pushed to the brink by human activity.
Summary
Genetic bottlenecks leave legible marks in genomic data: reduced heterozygosity, skewed allele frequency spectra, extended linkage disequilibrium, and coalescent genealogies that compress toward the period of the crash. The toolkit of modern population genetics — PSMC modeling, coalescent inference, LD decay analysis, and allelic diversity counting — has been applied to human genomes from populations worldwide, and the results converge with high consistency: the ancestral human effective population size never fell below approximately 10,000 breeding individuals, the out-of-Africa migration imposed a real but modest founding bottleneck of perhaps 1,000–3,000 individuals, and the HLA system alone preserves allelic lineages millions of years old that require thousands of continuously breeding ancestors to maintain.1, 2, 8, 11 The Toba supervolcano may have stressed regional human populations 74,000 years ago, but the genomic evidence does not support a global near-extinction event. The cheetah and the northern elephant seal illustrate what a genuine severe bottleneck looks like in genomic terms — and modern humans show none of those signatures. The implication for literalist interpretations of Genesis is straightforward: the genetic architecture of living humans is incompatible with descent from a single couple or from eight survivors of a global flood at any point within the history of the species.1, 8, 12
References
No evidence of a severe bottleneck in human evolution from analysis of extant mtDNA diversity
Extensive polymorphism of the HLA-A locus in humans: an ancient origin and transspecies evolution
Multiple instances of ancient balancing selection shared between humans and chimpanzees
Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data