Overview
- Mitochondrial Eve and Y-chromosomal Adam are the most recent common ancestors of all living humans through the strictly maternal (mtDNA) and strictly paternal (Y-chromosome) lineages, respectively, and both trace back to Africa, with current estimates placing mitochondrial Eve at roughly 150,000 to 200,000 years ago and Y-chromosomal Adam at approximately 200,000 to 300,000 years ago.
- These two individuals were not contemporaries, were not a couple, and were not the only humans alive at their respective times; they simply represent the inevitable coalescence points of uniparental genetic lineages, a mathematical consequence of population genetics rather than a literal first man and first woman.
- Deep-rooting lineages such as the Y-chromosome haplogroup A00, found among Mbo men in Cameroon and dating to approximately 338,000 years ago, and the mitochondrial haplogroup L0 lineages of southern African Khoisan populations have pushed the estimated coalescence times deeper into the past and revealed the extraordinary time depth of African genetic diversity.
Mitochondrial Eve and Y-chromosomal Adam are scientific designations for the most recent common ancestors of all living humans through the strictly maternal and strictly paternal genetic lineages, respectively. Mitochondrial DNA (mtDNA), a small circular genome located in the cell's energy-producing organelles, is inherited exclusively from the mother and does not recombine with paternal DNA, preserving a clean record of maternal descent that can be traced backward through time to a single ancestral woman. The Y chromosome, passed intact from father to son with only gradual mutational change, similarly allows reconstruction of a patrilineal genealogy converging on a single ancestral man. The landmark 1987 study by Rebecca Cann, Mark Stoneking, and Allan Wilson established that all human mtDNA lineages coalesce in Africa approximately 200,000 years ago, providing the first molecular evidence for the Recent African Origin of modern humans and introducing the concept of "mitochondrial Eve" to both science and popular culture.1
Despite the evocative biblical names, these two ancestors were not the only humans alive at their respective times, were not a couple, and almost certainly did not live in the same millennium. They represent coalescence points in uniparental lineages — a mathematical inevitability predicted by population genetics — rather than a literal first man and first woman from whom all genetic diversity descends. Current estimates place mitochondrial Eve at roughly 150,000 to 200,000 years ago and Y-chromosomal Adam at approximately 200,000 to 300,000 years ago, though these dates have shifted substantially as sequencing technology, molecular clock calibrations, and the discovery of deep-rooting lineages have refined the underlying phylogenies.5, 9, 12
The original mtDNA study
The concept of mitochondrial Eve emerged from a 1987 paper in Nature by Rebecca Cann, Mark Stoneking, and Allan Wilson at the University of California, Berkeley. The researchers analysed mitochondrial DNA from 147 individuals drawn from five geographic populations — Africa, Asia, Australia, Europe, and New Guinea — using restriction enzyme mapping to characterise variation across the mitochondrial genome. Because mtDNA is inherited only through the maternal line and does not undergo recombination, differences between individuals accumulate solely through mutation, making it possible to reconstruct a phylogenetic tree of maternal lineages.1
The resulting tree had two primary branches: one composed exclusively of African lineages and a second containing both African and non-African lineages. All non-African mtDNA types were nested within the broader African radiation, indicating that the deepest divergences in human maternal genealogy occurred within Africa and that non-African populations represented a subset of African diversity. By calibrating the rate of mtDNA divergence against the fossil record of human colonisation of Australia and New Guinea, Cann, Stoneking, and Wilson estimated that the most recent common ancestor of all sampled mtDNA lineages — a woman they refrained from naming but whom the press quickly dubbed "mitochondrial Eve" — lived approximately 200,000 years ago, most probably in Africa.1
The study was immediately controversial. Critics questioned the statistical methods used to root the tree, the calibration of the molecular clock, and the small sample size of African populations. In 1991, Vigilant, Stoneking, Harpending, Hawkes, and Wilson addressed several of these criticisms by sequencing two hypervariable segments of mtDNA from 189 individuals, including 121 native Africans, and applying more rigorous statistical tests. The expanded analysis confirmed an African root for the human mtDNA tree and supported the approximate 200,000-year date for the most recent common ancestor.2 Despite continuing methodological refinements, the core findings of the original study have withstood more than three decades of subsequent analysis: all human mtDNA lineages coalesce in Africa within the past 100,000 to 200,000 years.3, 5
Coalescent theory and uniparental lineages
The concept of a most recent common ancestor for mtDNA or the Y chromosome is not a biological anomaly but a mathematical certainty. In any finite population, genetic lineages are lost by chance in every generation: a woman who has no daughters, or whose daughters themselves have no daughters, sees her mtDNA lineage go extinct regardless of her other genetic contributions to the population. Over thousands of generations, this random process — called genetic drift — eliminates all but one maternal lineage, which becomes the ancestor of all surviving mtDNA types. The same logic applies to the Y chromosome through the paternal line.14
The formal mathematical framework for understanding this process is coalescent theory, developed by the mathematician John Kingman in 1982.14 The coalescent describes the genealogy of a sample of gene copies by tracing their ancestry backward in time. At each step into the past, two lineages may "coalesce" — that is, trace back to the same ancestral copy. For a non-recombining locus such as mtDNA or the Y chromosome, the entire genealogy is a single tree whose root is the most recent common ancestor (MRCA). The expected time to the MRCA in a randomly mating population of constant effective size N is proportional to N for a haploid locus (mtDNA, Y chromosome) and to 2N for a diploid autosomal locus. Because the effective population size for uniparental markers is approximately one-quarter that of autosomal markers — only females transmit mtDNA, and only males transmit the Y chromosome — the MRCA for these loci is expected to be more recent than the MRCA for any autosomal gene.14
A critical implication of coalescent theory is that the time to the most recent common ancestor (TMRCA) depends on the effective population size and demographic history, not on any special biological event. Mitochondrial Eve is not the woman from whom all humans descend; she is the woman from whom all mitochondrial DNA descends. Every other woman alive at the same time could have contributed autosomal DNA to the present-day gene pool through her sons or through daughters whose own maternal lineages later went extinct. The coalescence of mtDNA to a single ancestor is simply the inevitable outcome of drift operating on a haploid, non-recombining genome over sufficient time.1, 14
Refining the date of mitochondrial Eve
The original 200,000-year estimate for mitochondrial Eve relied on a molecular clock calibrated by the divergence between humans and chimpanzees and by the archaeological dating of human colonisation events. Subsequent studies using complete mitochondrial genome sequences and improved clock calibrations have refined this estimate, generally producing dates in the range of 150,000 to 200,000 years ago, though the precise figure depends on the calibration method and the correction applied for purifying selection — the removal of slightly deleterious mutations that inflates apparent divergence rates over short timescales relative to long ones.1, 5
In 2000, Ingman, Kaessmann, Pääbo, and Gyllensten published the first study based on complete mitochondrial genome sequences, analysing 53 individuals from diverse global populations. The resulting phylogeny confirmed the African root and placed the TMRCA at approximately 171,500 years ago, with African populations displaying more than twice the mtDNA diversity of non-African populations.3 In 2008, Behar and colleagues constructed a matrilineal tree from 624 complete mtDNA genomes belonging to the deepest African haplogroup L lineages, estimating that the Khoisan matrilineal ancestry diverged from the rest of the human mtDNA pool between 90,000 and 150,000 years ago and that at least five additional maternal lineages existed in parallel during this period.4
A major methodological advance came in 2009 when Soares and colleagues demonstrated that purifying selection acts differently on synonymous and nonsynonymous sites within the mitochondrial coding region, causing the apparent mutation rate to be time-dependent: higher over short timescales (when mildly deleterious mutations have not yet been removed by selection) and lower over longer timescales. By correcting for this effect using a worldwide phylogeny of more than 2,000 complete mtDNA genomes, they produced an improved molecular clock that placed the human mtDNA TMRCA at approximately 200,000 years ago when calibrated against the human-chimpanzee divergence.5 In 2013, Poznik and colleagues applied equivalent sequencing and analytical methods to both mtDNA and the Y chromosome in the same set of individuals, estimating the mtDNA TMRCA at 99,000 to 148,000 years ago using a mutation rate calibrated within the human lineage rather than against the chimpanzee divergence.10 The range of estimates across studies reflects genuine uncertainty in molecular clock calibration rather than fundamental disagreement about the African origin or approximate antiquity of the common ancestor.
Estimated dates for mitochondrial Eve across major studies1, 3, 5, 10
Y-chromosomal Adam and the patrilineal tree
The Y chromosome provides the patrilineal counterpart to mtDNA. Like mtDNA, the male-specific region of the Y chromosome (MSY) does not recombine during meiosis — except at the small pseudoautosomal regions at the tips of the chromosome — and is transmitted as a single haploid block from father to son. Mutations accumulate along the Y chromosome at a roughly steady rate, producing a branching phylogenetic tree whose root represents the most recent common ancestor of all living male Y-chromosome lineages: Y-chromosomal Adam.6, 10
Early estimates of the Y-chromosome TMRCA were substantially more recent than those for mtDNA, initially suggesting a patrilineal coalescence only 50,000 to 100,000 years ago. A 2000 study by Thomson, Pritchard, and Feldman, using DNA sequence data from three Y-chromosome genes in a worldwide sample, estimated the TMRCA at roughly 59,000 years ago.7 This surprisingly recent date led to speculation that the Y-chromosome genealogy had been shaped by strong selective sweeps or by a smaller male effective population size driven by high variance in male reproductive success. In the same year, Underhill and colleagues published a comprehensive survey of 166 biallelic polymorphisms across the MSY in over 1,000 globally representative males, constructing a parsimonious genealogy of 116 haplotypes that confirmed the African origin of Y-chromosome diversity but did not resolve the TMRCA with precision.6
The picture changed dramatically in the early 2010s as whole Y-chromosome sequencing became feasible. In 2011, Cruciani and colleagues sequenced approximately 200 kilobases of the MSY from seven Y chromosomes representing the deepest branches of the tree and redefined the root of the Y-chromosome phylogeny, placing the origin of patrilineal diversity in central-northwest Africa with a TMRCA of approximately 142,000 years ago.8 In 2013, Poznik and colleagues applied next-generation sequencing to 69 male genomes from nine globally diverse populations and, by using equivalent analytical methods for both the Y chromosome and mtDNA, estimated the Y-chromosome TMRCA at 120,000 to 156,000 years ago — largely overlapping with their mtDNA estimate of 99,000 to 148,000 years ago. This finding resolved the earlier apparent discrepancy between male and female coalescence times, demonstrating that the gap had been an artefact of limited data and inconsistent methodologies rather than a genuine biological difference.10
Deep-rooting lineages and haplogroup A00
The estimated date of Y-chromosomal Adam was pushed dramatically deeper in 2013 by the discovery of an extraordinarily divergent Y-chromosome lineage. Mendez and colleagues reported that an African American man from South Carolina carried a Y chromosome that fell outside all previously known branches of the human Y-chromosome phylogenetic tree. Sequencing approximately 240 kilobases of this chromosome revealed that it carried the ancestral state at every single-nucleotide polymorphism (SNP) that had previously defined the basal portion of the tree. The researchers designated this new lineage haplogroup A00 and estimated the TMRCA for the revised Y-chromosome tree at approximately 338,000 years ago, with a 95 percent confidence interval of 237,000 to 581,000 years ago.9
This date was remarkable because it exceeded not only the previous estimates for the mtDNA TMRCA but also the age of the oldest known anatomically modern human fossils at the time. Subsequent genealogical and genetic investigation traced the A00 lineage to the Mbo people of western Cameroon, where it persists at low frequency. The A00 lineage does not imply that its carriers are more "primitive" or that they represent a separate species; it simply means that the patrilineal split between A00 and all other Y-chromosome lineages occurred very deep in time, and that the A00 line has been maintained in a small population within Africa ever since.9
In 2014, Scozzari and colleagues published a comprehensive reanalysis of the deepest branches of the Y-chromosome tree using high-coverage sequencing of 68 male genomes, identifying thousands of novel SNPs and confirming the deep phylogenetic structure within Africa. Their analysis placed the root of the Y-chromosome tree at approximately 190,000 to 260,000 years ago when A00 was included, and revealed multiple distinct deep-branching lineages (A0, A1a, A1b) that are found exclusively in sub-Saharan Africa, consistent with a long period of population substructure on the continent before the out-of-Africa dispersal.11
On the mitochondrial side, the deepest-rooting lineages also point to ancient substructure within Africa. The macrohaplogroup L0, found at highest frequencies among the Khoisan-speaking hunter-gatherer populations of southern Africa, represents the earliest divergence in the human mtDNA tree. Behar and colleagues estimated that the L0 lineages diverged from the rest of the human mtDNA pool 90,000 to 150,000 years ago, and identified at least five additional maternal lineages that existed in parallel during this period.4 The geographic distribution of these deep-rooting lineages — A00 in western Cameroon, L0 in southern Africa — provides evidence for a complex, geographically structured origin of our species across the African continent, rather than a simple expansion from a single localised population.
Why they did not live at the same time
One of the most common points of confusion about mitochondrial Eve and Y-chromosomal Adam is the assumption that they must have been contemporaries. In fact, there is no biological or mathematical reason why the matrilineal and patrilineal coalescence points should coincide in time. The TMRCA for any uniparental lineage depends on the effective population size for that lineage's mode of inheritance, the demographic history of the population, and the variance in reproductive success within each sex. Because these factors differ for males and females, the two coalescence times are expected to differ, and in practice they have been estimated at different dates by every major study.10, 14
The effective population size for the Y chromosome is particularly sensitive to variance in male reproductive success. In polygynous mating systems, a small number of males may father a disproportionately large share of offspring, reducing the effective number of Y-chromosome lineages transmitted to the next generation and driving faster coalescence. Conversely, the effective population size for mtDNA is influenced by the variance in female reproductive success, which in most human societies is lower than for males. Karmin and colleagues demonstrated this effect powerfully in 2015 by analysing 456 whole Y-chromosome sequences from 110 populations. They found that while the Y-chromosome TMRCA was approximately 254,000 years ago (95 percent confidence interval: 192,000 to 307,000 years ago), the tree showed a dramatic reduction in Y-chromosome lineage diversity within the past 10,000 years — a bottleneck not observed in mtDNA data — which they attributed to cultural changes during the Neolithic transition that increased the variance in male reproductive success.12
Poznik and colleagues' 2016 analysis of 1,244 Y-chromosome sequences from the 1000 Genomes Project revealed "punctuated bursts" of male lineage expansion that occurred independently in each of five continental superpopulations at times corresponding to known migrations and technological innovations.13 These bursts — in which a small number of patrilineal lineages expanded rapidly, presumably because their bearers achieved disproportionate reproductive success — have no parallel in the mtDNA record, illustrating how sex-specific demographic processes can decouple the matrilineal and patrilineal genealogies.
In addition to demographic factors, the two coalescence times are stochastic quantities: even in a population with identical effective sizes for males and females, random drift would produce different TMRCAs for mtDNA and the Y chromosome in any given simulation. The observation that the two dates differ by tens of thousands of years is therefore entirely expected and carries no implication that one sex experienced a more recent origin than the other.14
Common misconceptions
The names "mitochondrial Eve" and "Y-chromosomal Adam" have generated persistent public misconceptions that merit explicit correction. The most fundamental misunderstanding is the belief that these individuals were the only humans alive at their respective times — that the human species passed through a bottleneck of a single breeding pair. This is emphatically not the case. Mitochondrial Eve lived within a population that may have numbered tens of thousands of individuals, all of whom were fully human and many of whom contributed autosomal DNA to living people. What distinguishes Eve is solely that her mitochondrial lineage, transmitted through an unbroken chain of mothers and daughters, is the only one that has survived to the present. Every other woman alive at the same time also had mtDNA, but her particular maternal line eventually terminated — not because she had no descendants, but because at some point in the intervening millennia, one of her matrilineal descendants had only sons, or had no surviving children at all.1, 4
A second misconception is that mitochondrial Eve was the first human woman or the "mother of all humanity" in a total genetic sense. In reality, the vast majority of any individual's genome — the 22 pairs of autosomes and the X chromosome — is inherited from both parents and recombines every generation, producing a genealogical network vastly more complex than any single-locus tree. The autosomal most recent common ancestor of all living humans almost certainly lived far more recently than mitochondrial Eve, perhaps within the past few tens of thousands of years, because recombination allows autosomal lineages to coalesce through any genealogical path, not only through unbroken chains of mothers.14
A third misconception, particularly common in popular media, is that the discovery of mitochondrial Eve confirms or supports the biblical account of Adam and Eve. The scientific concepts are fundamentally different: mitochondrial Eve and Y-chromosomal Adam are statistical constructs derived from population genetics, not literal first humans created de novo. They were members of existing populations, separated by potentially hundreds of thousands of years, and their identification depends entirely on the current distribution of surviving genetic lineages. If a present-day maternal lineage were to go extinct — through the chance failure of all its carriers to produce daughters — the identity of mitochondrial Eve would shift to a more recent individual. The coalescence point is a property of the surviving genealogy, not a fixed point in biological history.14
Haplogroup diversity and African deep structure
The phylogenetic trees of both mtDNA and the Y chromosome reveal a pattern of deep, extensive branching within Africa and comparatively shallow, recent branching outside the continent. This geographic asymmetry in genetic diversity is among the strongest lines of molecular evidence for the African origin of Homo sapiens and for the Out of Africa dispersal as the primary source of non-African populations.1, 6
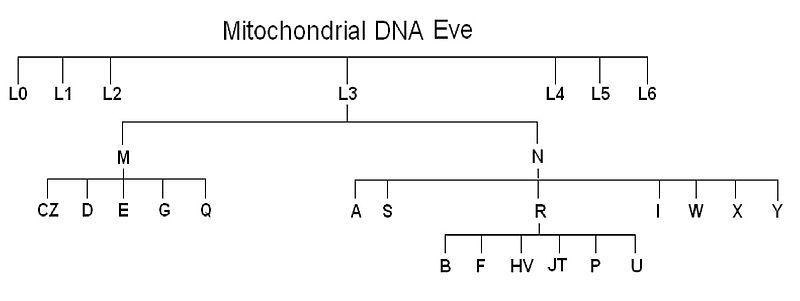
Mitochondrial haplogroups are designated by letters of the alphabet, with the deepest-branching African lineages assigned to macrohaplogroup L. The basal split in the human mtDNA tree separates L0 (found predominantly among Khoisan and some East African populations) from L1 through L6 (the remaining African lineages), with all non-African haplogroups (M, N, and their descendants, including R, U, H, V, J, T, and others) nested within the L3 branch. This nesting means that the entire non-African mtDNA diversity represents a single offshoot of African diversity — a subset that left the continent roughly 50,000 to 70,000 years ago and subsequently diversified into the haplogroups found today across Europe, Asia, Oceania, and the Americas.4, 5
The Y-chromosome haplogroup tree displays a similar architecture. The deepest branches — haplogroups A00, A0, A1a, and A1b — are found exclusively in sub-Saharan Africa, while the vast majority of non-African Y-chromosome diversity falls within haplogroups descended from a single node (formerly called "CT" or "BT") that diverged from the African-only lineages relatively recently. Scozzari and colleagues' 2014 reanalysis of the tree's basal structure confirmed that at least four deeply divergent lineages have persisted independently within Africa for more than 100,000 years, reflecting ancient population substructure on the continent.11
Major deep-rooting haplogroups and their estimated coalescence dates4, 9, 11, 12
| Lineage | Type | Estimated TMRCA | Geographic distribution |
|---|---|---|---|
| Y-chromosome A00 | Y-DNA | ~338 kya (237–581 kya) | Mbo, western Cameroon |
| Y-chromosome A0 | Y-DNA | ~200–260 kya | Central-west Africa |
| Y-chromosome A1a | Y-DNA | ~140–200 kya | West & central Africa |
| mtDNA L0 | mtDNA | ~90–150 kya | Khoisan, southern Africa |
| mtDNA L1 | mtDNA | ~80–130 kya | Central & West Africa |
| mtDNA L3 | mtDNA | ~60–70 kya | East Africa (source of non-African lineages) |
The persistence of these deep-rooting lineages at low frequencies within geographically restricted African populations suggests that the ancestral human population was not a single, homogeneous group but rather a collection of partially isolated subpopulations that exchanged migrants at low rates over hundreds of thousands of years. This model of African population structure — sometimes described as a "structured metapopulation" — is consistent with the pan-African distribution of early Homo sapiens fossils and the archaeological evidence for regionally distinct Middle Stone Age technologies across the continent.4, 15
Molecular clocks and the fossil record
The dates derived from mtDNA and Y-chromosome phylogenies do not stand in isolation; they must be reconciled with the fossil and archaeological record of modern human origins. The concordance between these independent lines of evidence has strengthened considerably over the past two decades, as both the molecular estimates and the fossil record have converged on a picture of Homo sapiens emerging in Africa between roughly 200,000 and 300,000 years ago.15, 16
The oldest fossils currently attributed to Homo sapiens come from Jebel Irhoud in Morocco, where Hublin and colleagues reported in 2017 a series of cranial, mandibular, and dental remains dated by thermoluminescence to approximately 315,000 years ago. These specimens display a mosaic of modern and archaic features — a modern-looking face combined with a more elongated, archaic braincase — and were associated with Middle Stone Age stone tools and evidence of fire use. The Jebel Irhoud fossils pushed the earliest appearance of Homo sapiens morphology back by more than 100,000 years from the previously accepted oldest fossils and supported a pan-African rather than East African origin for the species.15 The Herto fossils from Ethiopia, described by White and colleagues in 2003 and dated to approximately 160,000 years ago, represent a more anatomically modern form classified as the subspecies Homo sapiens idaltu.16
The Y-chromosome TMRCA estimates of 190,000 to 338,000 years ago, depending on whether the A00 lineage is included and which mutation rate is applied, bracket the age of the Jebel Irhoud fossils remarkably well.9, 11, 12 The mtDNA TMRCA of approximately 150,000 to 200,000 years ago is somewhat more recent than the oldest fossils, but this is expected: the coalescence of any single genetic locus necessarily postdates the origin of the species, because the ancestral population from which the species emerged would have carried multiple lineages, only one of which survives to the present. The TMRCA of mtDNA provides a minimum bound on the age of the population, not a date for the origin of anatomically modern morphology.
The molecular and fossil evidence together support a model in which the anatomical features of Homo sapiens assembled gradually across Africa over a period spanning roughly 300,000 to 200,000 years ago, with the earliest populations exhibiting mosaic morphologies and the fully modern form emerging later. The deep-rooting genetic lineages preserved in living African populations record this period of protracted evolution and geographic structuring, providing a temporal framework that complements and corroborates the fragmentary fossil record.4, 15, 16
Demographic insights from uniparental markers
Beyond providing coalescence dates, the phylogenetic trees of mtDNA and the Y chromosome encode information about past population sizes, migration events, and demographic expansions. The density and distribution of branches in a phylogenetic tree reflect the demographic history of the population that generated them: rapid population growth produces a characteristic "star-like" phylogeny with many lineages diverging in a short interval, while a population bottleneck leaves a tree with few surviving lineages separated by long branches.12, 13
Poznik and colleagues' 2016 analysis of 1,244 Y-chromosome sequences from the 1000 Genomes Project identified multiple episodes of rapid patrilineal expansion, or "punctuated bursts," that occurred independently on different continents at times corresponding to major cultural and technological transitions. In Africa, a burst of Y-chromosome lineage expansion coincided with the spread of agriculture and pastoralism. In Europe, a pronounced expansion corresponded to the Bronze Age migrations associated with the Yamnaya culture. In East Asia, a dramatic expansion occurred within the past 5,000 years, possibly linked to the spread of particular ruling lineages.13
Karmin and colleagues' 2015 study revealed a striking asymmetry between the mtDNA and Y-chromosome records of the past 10,000 years. While female effective population size, inferred from mtDNA, increased steadily with the adoption of agriculture, male effective population size underwent a dramatic contraction beginning around 8,000 to 4,000 years ago — a "Neolithic Y-chromosome bottleneck" that reduced the effective number of male lineages to as few as one-twentieth of the female effective population size in some regions. The most parsimonious explanation is that the transition to agricultural societies, with their increased social stratification, patrilineal inheritance of land and livestock, and potential for polygyny among high-status males, concentrated reproductive success among a smaller number of patrilines.12
These findings illustrate a fundamental point: the matrilineal and patrilineal genealogies are not merely passive records of species history but active reflections of the social, cultural, and ecological forces that have shaped human populations over hundreds of thousands of years. The different demographic signals encoded in mtDNA and the Y chromosome provide complementary and sometimes contrasting perspectives on the same population history, together yielding a richer picture than either could provide alone.12, 13
Significance for human origins research
The concepts of mitochondrial Eve and Y-chromosomal Adam have had a transformative impact on the study of human origins, establishing molecular genetics as a central discipline in paleoanthropology and providing the empirical foundation for the Out of Africa model of modern human evolution. The original Cann, Stoneking, and Wilson study was among the first to demonstrate that questions about human evolution could be answered with DNA from living people rather than from fossils alone, inaugurating the field of molecular anthropology.1
The subsequent refinement of uniparental phylogenies through whole-genome sequencing has produced an increasingly detailed picture of human demographic history. The discovery of deep-rooting lineages such as Y-chromosome haplogroup A00 has revealed that the genetic history of our species extends deeper in time than previously appreciated, while the identification of sex-specific demographic signals — such as the Neolithic Y-chromosome bottleneck — has opened new avenues for understanding how cultural practices shape genetic diversity.9, 12
At the same time, the study of uniparental markers has inherent limitations that must be acknowledged. Both mtDNA and the Y chromosome represent single genetic loci that trace only one of the many genealogical paths connecting present-day individuals to their ancestors. The autosomal genome, which recombines every generation and is inherited from both parents, contains far more information about population history, admixture, and selection, and the development of whole-genome sequencing and ancient DNA analysis has increasingly shifted the focus of human evolutionary genetics from uniparental markers to the autosomal genome. Nevertheless, mtDNA and the Y chromosome remain invaluable tools precisely because their lack of recombination makes them uniquely amenable to phylogenetic analysis, providing clear, unambiguous trees of descent that serve as the scaffolding on which the more complex history of the autosomal genome can be interpreted.
The legacy of the mitochondrial Eve and Y-chromosomal Adam concepts extends beyond genetics into the public understanding of human origins. By demonstrating that all living humans share recent common ancestors traced through both the maternal and paternal lines, and that these ancestors lived in Africa, the molecular evidence has powerfully reinforced the fundamental unity of the human species. The genetic differences among the world's populations are shallow and recent, superimposed on a shared African ancestry that connects every living person to a common heritage stretching back hundreds of thousands of years.1, 12, 13
References
A revised root for the human Y chromosomal phylogenetic tree: the origin of patrilineal diversity in Africa
An African American paternal lineage adds an extremely ancient root to the human Y chromosome phylogenetic tree
Sequencing Y chromosomes resolves discrepancy in time to common ancestor of males versus females
An unbiased resource of novel SNP markers provides a new chronology for the human Y chromosome and reveals a deep phylogenetic structure in Africa
A recent bottleneck of Y chromosome diversity coincides with a global change in culture
Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences